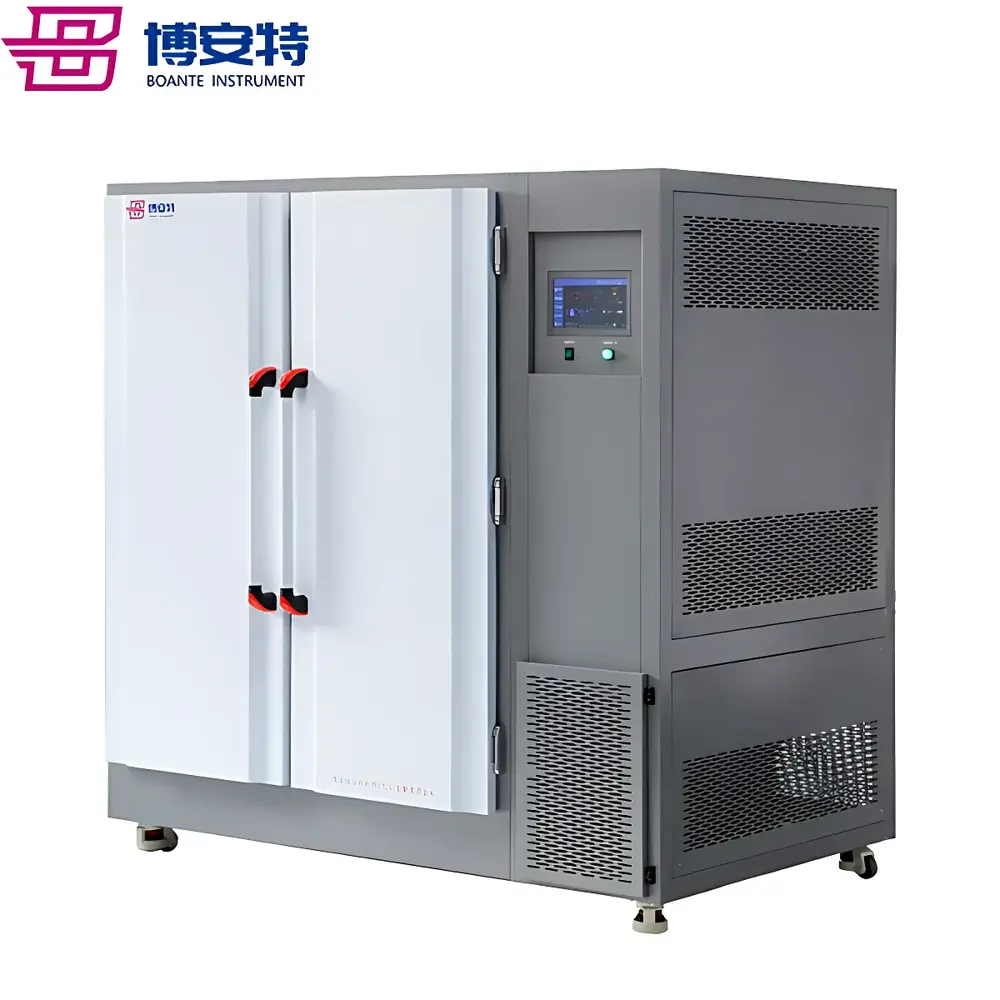
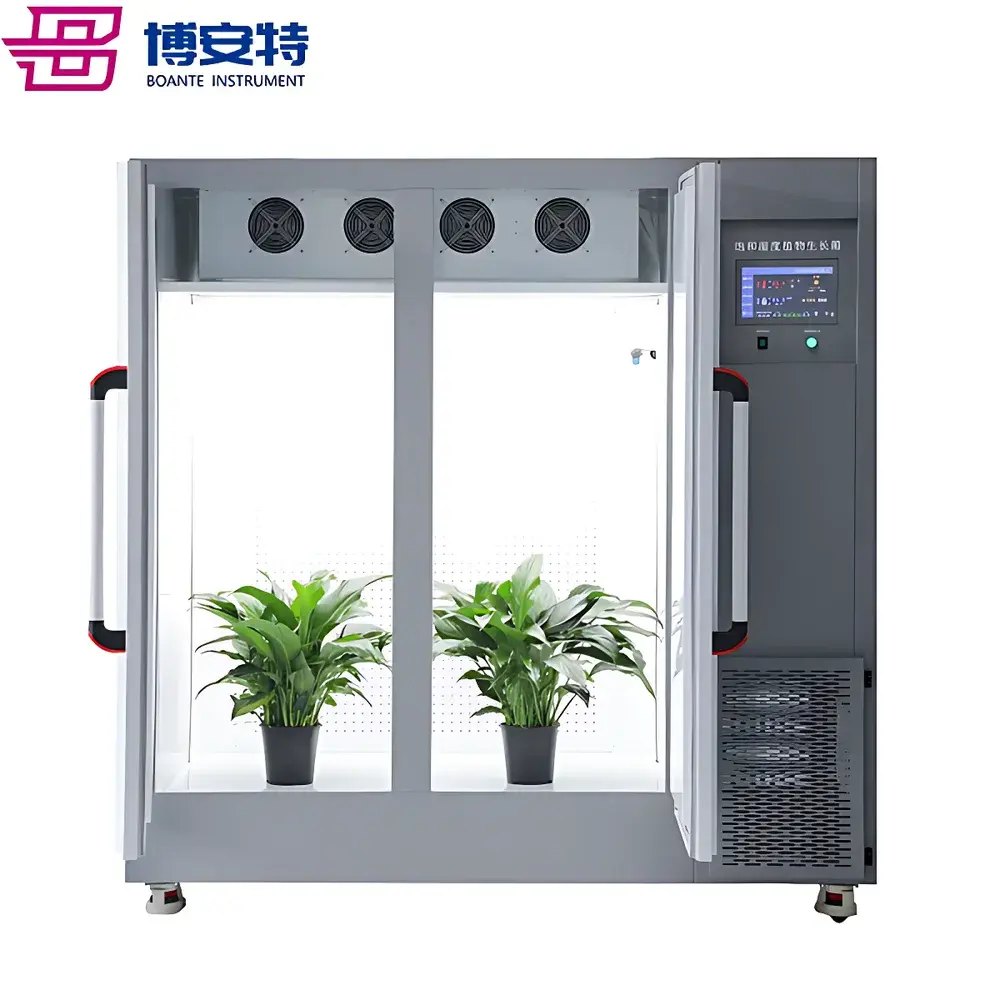
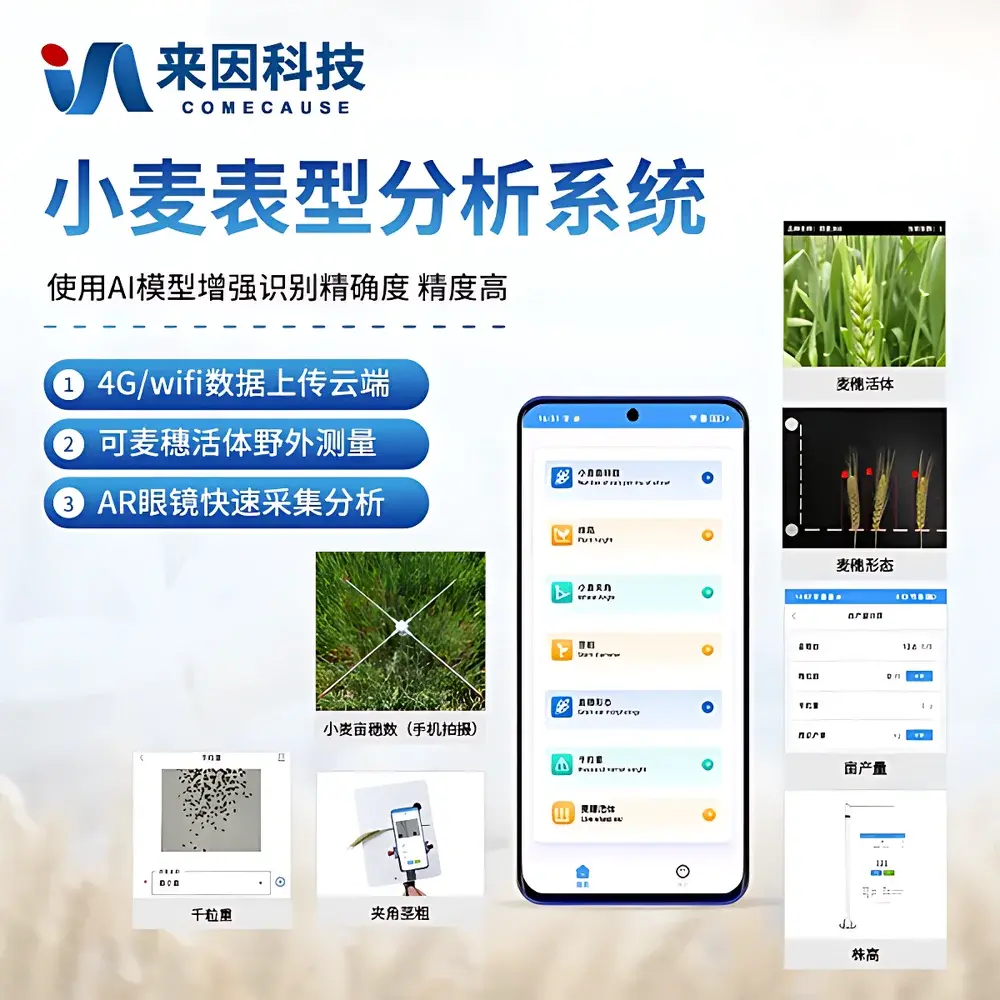
COMECAUSE IN~XM04 Wheat Phenotyping System
| Brand | COMECAUSE |
|---|---|
| Origin | Shandong, China |
| Manufacturer Type | Original Equipment Manufacturer (OEM) |
| Country of Origin | China |
| Model | IN~XM04 |
| Price | USD 4,200 (FOB) |
Overview
The COMECAUSE IN~XM04 Wheat Phenotyping System is a field-deployable, modular digital phenotyping platform engineered for high-throughput, non-destructive, and standardized acquisition of morphological, structural, and functional traits across the entire wheat growth cycle—from seedling emergence through tillering, heading, grain filling, to physiological maturity. Unlike traditional manual phenotyping methods constrained by observer bias, low throughput, and temporal resolution limits, the IN~XM04 integrates calibrated optical sensing, AI-driven computer vision, and embedded Android-based edge computing to convert dynamic field observations into structured, traceable, and interoperable phenotypic data. Its core measurement principles include active optical triangulation (for plant height), multi-angle image-based segmentation and feature extraction (for spike morphology, tiller count, and canopy architecture), geometric calibration with reference markers (for inter-node angle and stem diameter), and spectral-informed regression modeling (for indirect estimation of chlorophyll content, nitrogen status, and water-use efficiency). Designed in alignment with FAO’s Global Strategy for Crop Phenotyping and the MIAPPE (Minimum Information About a Plant Phenotyping Experiment) v1.1 metadata standards, the system enables reproducible, audit-ready data collection suitable for QTL mapping, genomic selection training sets, and G×E interaction modeling.
Key Features
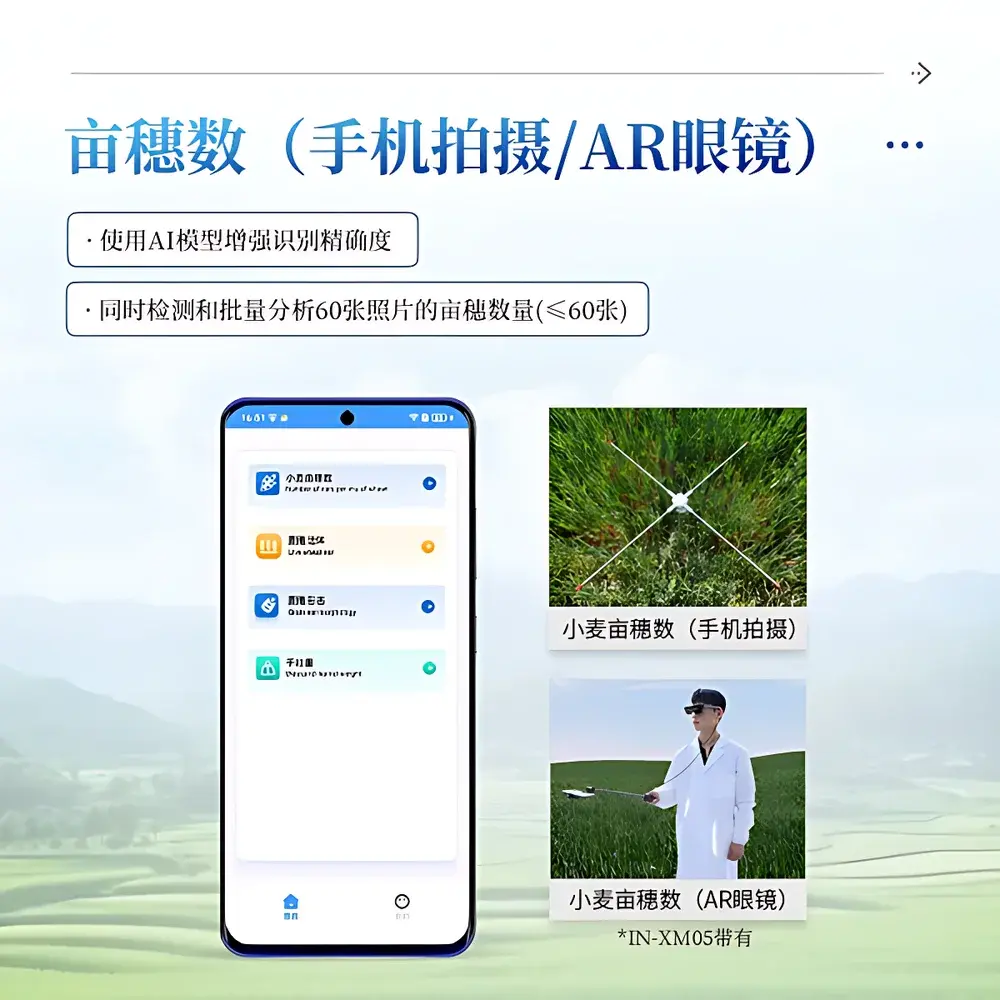
- Modular hardware architecture: Four interoperable, battery-powered modules—Spike Count & Yield Estimation Unit, Spike Morphology Analyzer, Leaf-Stem Angle & Stem Diameter Sensor, and Optical Height Profiler—each optimized for specific trait classes and deployable independently or in synchronized workflows.
- Edge-AI processing: On-device inference using lightweight CNN models trained on >50,000 annotated wheat images across diverse genotypes and environments; supports real-time spike detection (F1-score ≥0.93), length measurement (±1% error), and angle quantification (±1°).
- Robust field operation: Dual-camera imaging system (50 MP main + 12 MP auxiliary) with auto-white balance, tilt-compensated perspective correction, and ambient-light-invariant exposure control; no external shading or controlled lighting required.
- Standardized metrology: All modules incorporate NIST-traceable calibration references—including adjustable-height ground truth frames (610–1920 mm vertical range), precision-machined acrylic backboards (500 × 400 mm), and stainless-steel articulating fixtures—to ensure cross-site and cross-operator measurement consistency.
- Integrated data pipeline: Local storage (128–256 GB microSD), GPS-tagged metadata embedding (time, location, operator ID, module ID), Excel export (CSV/ XLSX), and optional encrypted cloud sync via Wi-Fi or LTE for centralized database ingestion (e.g., BreedBase, BrAPI-compliant repositories).
- Bilingual UI & regulatory compliance: Android-based touchscreen interface with one-tap EN/CN language switching; software architecture supports audit trails, user role management, and electronic signature logging per FDA 21 CFR Part 11 Annex 11 requirements for GLP/GMP-aligned trials.
Sample Compatibility & Compliance
The IN~XM04 is validated for use with Triticum aestivum L. (bread wheat) and exhibits cross-species utility for Oryza sativa (rice) and Brassica napus (oilseed rape) in leaf angle and stem diameter measurements. It complies with ISO 20672-1:2021 (Plant phenotyping — Vocabulary — Part 1: General terms) and adheres to the experimental design conventions outlined in ASTM E3274-22 (Standard Guide for Field Phenotyping of Cereal Crops). All measurement protocols align with the CIMMYT Wheat Program’s Standard Evaluation Protocols (SEPs) for yield component analysis and the IWG-Phenotyping’s Recommended Practices for Field-Based High-Throughput Phenotyping. Data outputs conform to BrAPI v2.1 schema specifications for trait, observation, and germplasm metadata exchange.
Software & Data Management
The system runs COMECAUSE PhenotypeOS v3.2—a purpose-built Android application supporting offline-first operation, version-controlled firmware updates, and modular plugin architecture. Core capabilities include batch image processing (up to 60 frames/session), automated trait vector generation (e.g., “spike_length_cm”, “tiller_angle_deg”, “stem_diameter_mm”), and integrated yield modeling (theoretical yield =亩穗数 × 穗粒数 × 千粒重 / 10⁶). All raw images, processed masks, and metadata are stored with SHA-256 checksums to ensure data integrity. Export formats include MIAPPE-compliant JSON-LD, Excel (.xlsx) with embedded validation rules, and BrAPI-compatible POST payloads. Cloud synchronization employs TLS 1.3 encryption and OAuth 2.0 authentication; audit logs record all user actions (login, measurement, export, delete) with timestamps and device fingerprints—fully compliant with ISO/IEC 27001:2022 information security controls.
Applications
- Genetic resource characterization: High-resolution phenotyping of core collections (e.g., ICARDA wheat germplasm bank) to build trait ontologies linked to SNP arrays and whole-genome sequencing data for GWAS and genomic prediction.
- Early-generation selection: Non-destructive screening of F₂–F₄ populations for architectural traits (e.g., lodging resistance index derived from stem diameter-to-height ratio) to reduce field nursery size by ≥40% without sacrificing genetic gain.
- Multi-environment trial (MET) analysis: Standardized acquisition of canopy temperature, normalized difference vegetation index (NDVI), and spike density across ≥10 locations/year to quantify genotype-by-environment stability for drought tolerance and heat resilience.
- Precision agronomy research: Quantifying dynamic responses of biomass accumulation rate, leaf area index (LAI), and senescence timing under variable nitrogen rates and planting densities—feeding crop simulation models (e.g., APSIM-Wheat) with empirically derived parameters.
- Regulatory phenotyping: Supporting OECD Test Guidelines 522 (Seed Dormancy) and 523 (Seed Vigor) through objective, repeatable quantification of germination uniformity, coleoptile length, and early vigor indices under controlled and field conditions.
FAQ
What growth stages are supported for each trait module?
Spike counting: Heading to late grain-filling stage (Zadoks GS 55–85); spike morphology: post-harvest lab analysis only; leaf-stem angle & stem diameter: GS 31–77 (jointing to milk ripeness); plant height: all vegetative and reproductive stages (GS 10–92); thousand-kernel weight: lab-based threshed grain analysis.
Is calibration required before each field session?
Yes—each module includes built-in calibration routines: height profiler uses internal inclinometer and laser distance sensor self-check; spike analyzer performs automatic scale-bar detection from embedded fiducials; angle/stem module validates mechanical zero via reference goniometer and caliper check.
Can data be integrated with third-party breeding management software?
Yes—via BrAPI v2.1 RESTful endpoints or direct CSV/Excel import into platforms including BreedBase, FieldBook, and AgriWebb; custom API connectors available upon request.
What environmental conditions affect measurement accuracy?
No significant degradation observed under full sunlight, light rain, or wind ≤5 m/s; performance validated at ambient temperatures from –5°C to 45°C and relative humidity 20–95% RH (non-condensing).
Does the system support longitudinal monitoring of individual plants?
Yes—GPS geotagging, unique plant ID assignment, and time-series trait tracking enable cohort-level growth curve modeling; recommended minimum revisit interval: 3–5 days depending on growth stage.