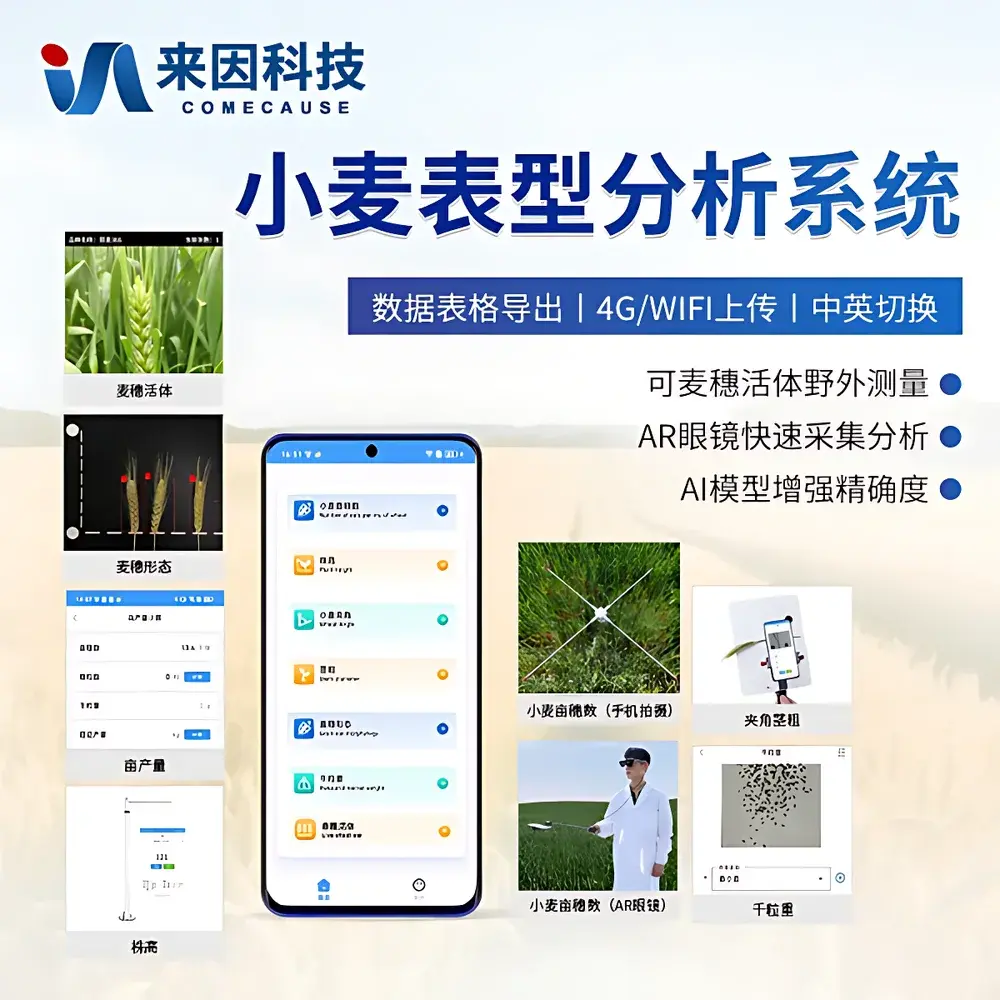
COMECAUSE IN~XM05 Wheat Phenotyping System
| Brand | COMECAUSE |
|---|---|
| Origin | Shandong, China |
| Manufacturer Type | Direct Manufacturer |
| Model | IN~XM05 |
| Measurement Accuracy | ±1–3% (varies by module) |
| Imaging Resolution | 4000×3000 |
| Camera System | Dual-sensor (50 MP + 12 MP) |
| Operating System | Android-based embedded UI |
| Data Export | Excel (.xlsx), GPS-tagged metadata |
| Cloud Integration | Wi-Fi/4G-enabled automatic upload |
| Language Support | Switchable English/Chinese UI |
| Storage Capacity | Up to 256 GB internal |
| Battery Life | >10 hours continuous operation |
| Field Deployment | Portable, handheld, and tripod-mountable modules |
Overview
The COMECAUSE IN~XM05 Wheat Phenotyping System is a modular, field-deployable platform engineered for high-throughput, non-destructive, and standardized acquisition of morphological, structural, and yield-related phenotypic traits across the wheat growth cycle—from germination through grain filling and maturity. Unlike conventional single-parameter instruments, this system implements a multi-sensor fusion architecture grounded in computer vision, optical metrology, and AI-driven image analysis. It operates on the principle of geometric feature extraction from calibrated digital imagery, combined with embedded optical distance measurement and angular kinematics analysis. Each module applies validated algorithms trained on diverse wheat germplasm under variable environmental conditions to ensure robustness across genotypes and growth stages. Designed for integration into breeding pipelines and precision agronomy workflows, the IN~XM05 bridges genotype–phenotype–environment relationships by transforming field observations into structured, auditable, and interoperable datasets compliant with FAO Crop Ontology and MIAPPE (Minimum Information About a Plant Phenotyping Experiment) standards.
Key Features
- Modular hardware architecture: Four independent but interoperable units—tiller density imager, spike morphology analyzer, leaf-stem angle & stem diameter sensor, and optical height profiler—each optimized for specific trait domains.
- Dual-spectrum imaging capability: 50 MP primary sensor + 12 MP auxiliary sensor enables high-fidelity texture and spatial resolution for segmentation-sensitive tasks such as spikelet counting and internode boundary detection.
- Real-time AI inference engine: On-device deep learning models (YOLOv5-based detectors and U-Net segmentation architectures) perform inference without cloud dependency; model weights are periodically updated via secure OTA patches.
- AR-assisted field acquisition: Integrated 2K-resolution augmented reality eyewear enables hands-free framing, geo-referenced sampling, and live overlay of measurement guidance—critical for rapid plot-level surveys under variable lighting or wind conditions.
- Multi-stage calibration framework: Includes physical reference bars (adjustable 610–1920 mm), auto-correcting perspective distortion algorithms, and dynamic white balance compensation to maintain measurement fidelity across diurnal cycles and weather transitions.
- Comprehensive metadata tagging: All outputs include timestamp, GPS coordinates (±3 m CEP), device orientation, ambient illuminance (lux), and operator ID—enabling traceability required for GLP-compliant trials.
Sample Compatibility & Compliance
The IN~XM05 supports phenotyping of Triticum aestivum L. across all major growth stages defined by the BBCH scale (BBCH 10–99), including seedling emergence (BBCH 11), tillering (BBCH 21–29), stem elongation (BBCH 31–39), booting (BBCH 41–49), heading (BBCH 51–59), flowering (BBCH 61–69), milk ripening (BBCH 71–79), dough ripening (BBCH 81–89), and full maturity (BBCH 91–99). Its modular design extends compatibility to related Poaceae species—including Oryza sativa (rice) and Brassica napus (oilseed rape)—for comparative phenomics studies. The system adheres to ISO 20607:2019 (Agricultural machinery — Requirements for data exchange between farm management information systems), and its data export structure conforms to ISA-Tab v1.1 for reproducible plant science reporting. All software components meet FDA 21 CFR Part 11 requirements for electronic records and signatures, including audit trails, role-based access control, and immutable log archiving.
Software & Data Management
The IN~XM05 runs on a hardened Android 12 LTS platform with proprietary firmware (v4.3.x) that isolates instrument control logic from user applications. Data acquisition, preprocessing, and trait quantification occur locally; no raw images are transmitted unless explicitly authorized. Processed outputs—including trait values, confidence scores, segmented masks, and annotated thumbnails—are stored in SQLite databases encrypted at rest (AES-256). Users export datasets directly to Excel (.xlsx) with embedded metadata schemas aligned with BreedBase and Gramene ontologies. Cloud synchronization (via AWS IoT Core endpoint) is optional and configurable per trial—supporting both private VPC deployment and federated access for multi-site breeding programs. The companion web portal provides RESTful API access for integration with genomic selection platforms (e.g., AlphaSimR, rrBLUP) and statistical analysis environments (R, Python).
Applications
- High-resolution QTL mapping: Enables precise correlation of spike architecture traits (e.g., spike length, spikelet count, rachis internode length) with SNP markers in biparental or MAGIC populations.
- Climate-resilience screening: Quantifies dynamic canopy architecture changes (leaf angle, stem lodging resistance index) under drought or heat stress treatments—feeding into predictive models of photosynthetic efficiency and water-use efficiency.
- Yield component decomposition: Integrates tiller density, fertile spike count, grain number per spike, and thousand-kernel weight to compute theoretical yield potential and identify physiological bottlenecks.
- Pre-breeding acceleration: Supports early-generation selection (F₂–F₄) using non-destructive proxies for harvest index, biomass partitioning, and grain fill duration—reducing cycle time by up to 30% versus traditional threshing-based assays.
- Smart field scouting: Generates georeferenced trait heatmaps across experimental plots, enabling zone-specific irrigation or nitrogen top-dressing decisions aligned with EU Farm to Fork Strategy indicators.
FAQ
What growth stages are supported for tiller density measurement?
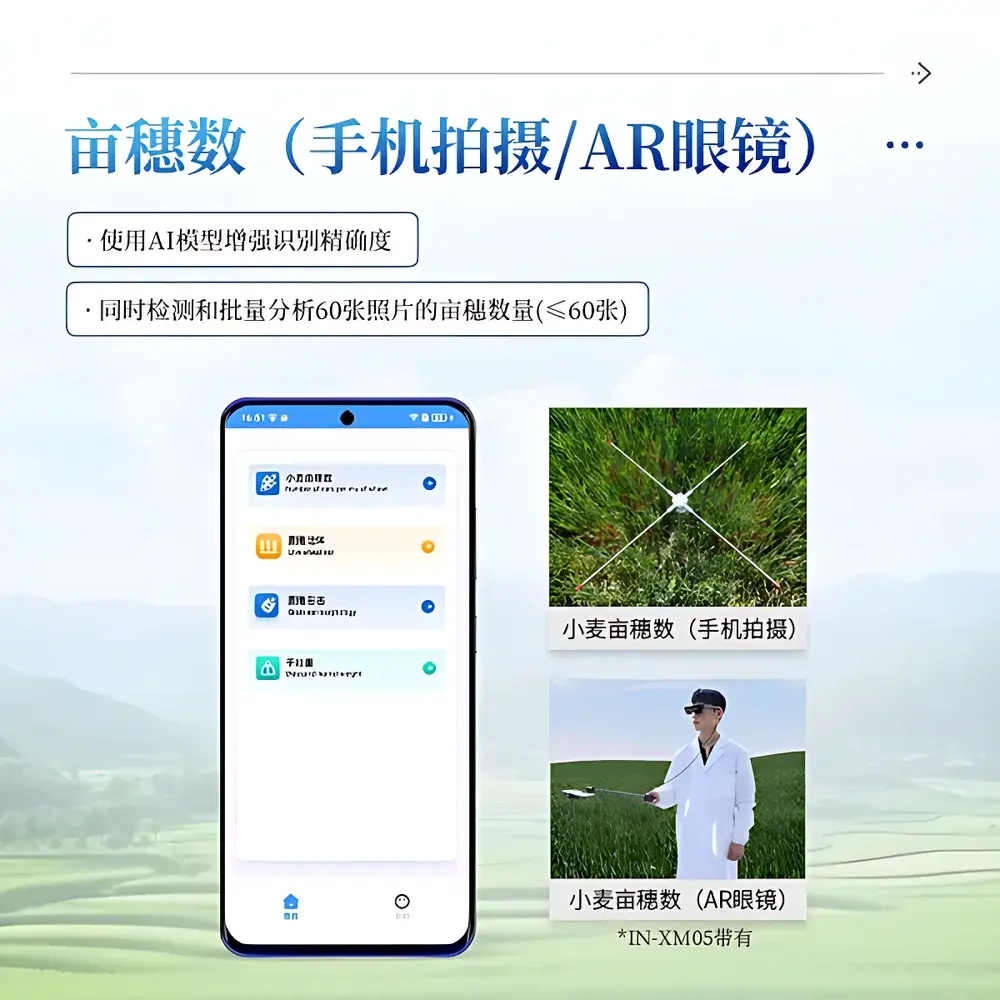
Tiller density (per unit area) is optimized for BBCH 51–85—covering heading through late grain filling—when spike emergence is complete and canopy closure allows reliable top-down imaging.
Can the system operate offline in remote field locations?
Yes. All core image processing, trait extraction, and quality control checks occur on-device. Cloud upload is optional and only activates when network connectivity is detected.
Is calibration required before each use?
Physical calibration using included reference bars is recommended before first use in a new environment or after significant temperature shifts (>15°C); software auto-calibration handles minor perspective and illumination drift during operation.
How does the system handle mixed-genotype stands or weed interference?
AI models are trained on heterogeneous field backgrounds and include background subtraction layers trained on common weed morphologies (e.g., Avena fatua, Bromus tectorum); manual ROI masking is also supported via touchscreen interface.
What data security protocols are implemented?
All local storage uses full-disk encryption (FDE); cloud transfers employ TLS 1.3 with mutual authentication; audit logs record every data export, deletion, or parameter modification event with cryptographic hashing for forensic verification.