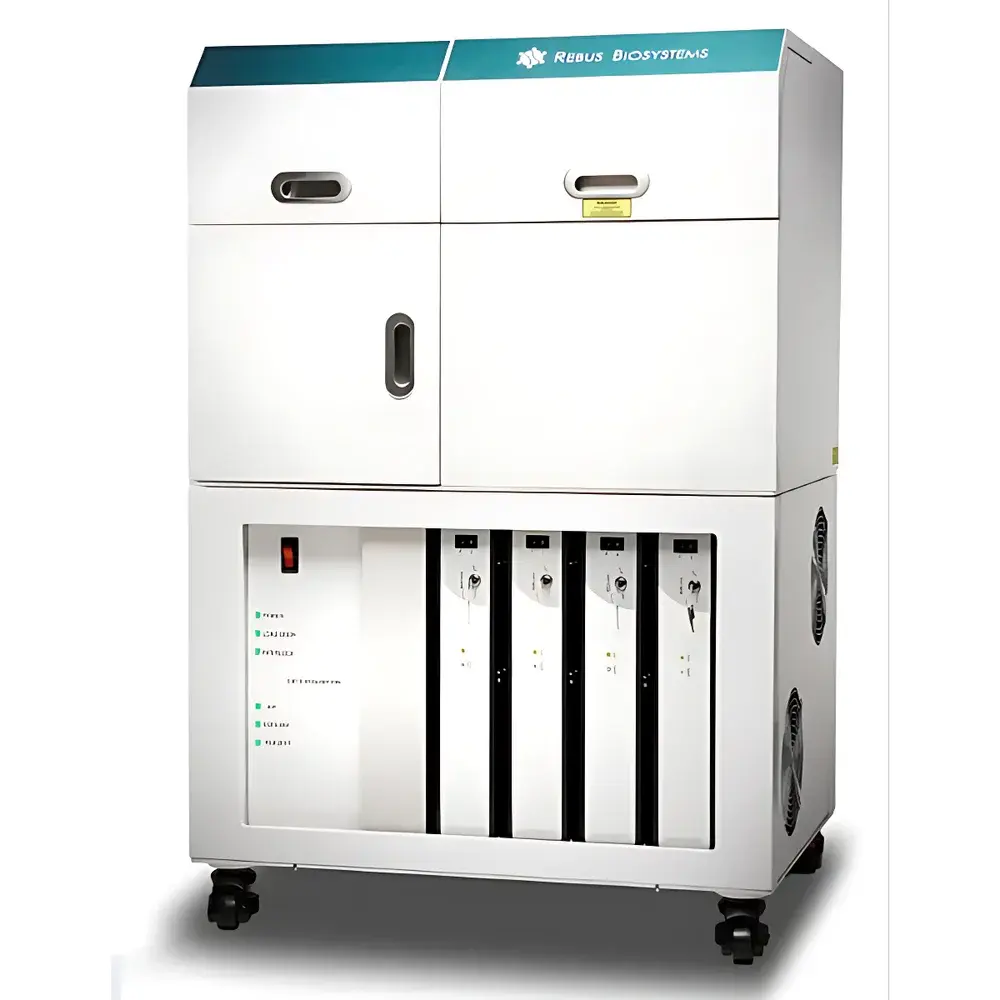
Artinis Rebus ESPER Spatial Transcriptomics Platform
| Brand | Artinis |
|---|---|
| Origin | USA |
| Manufacturer Type | Authorized Distributor |
| Origin Category | Imported |
| Model | Rebus ESPER |
| Price Range | USD 270,000 – 540,000 |
Overview
The Artinis Rebus ESPER Spatial Transcriptomics Platform is a fully integrated, automated end-to-end system engineered for quantitative, subcellular-resolution spatial transcriptomics in intact tissue sections. It leverages synthetic aperture optics (SAO)—a patented illumination strategy based on interferometric laser beam patterning—to acquire high-fidelity, diffraction-limited images without the physical constraints of high-NA oil-immersion objectives. Unlike conventional fluorescence in situ hybridization (FISH)-based or sequencing-derived spatial methods, Rebus ESPER combines single-molecule sensitivity with true spatial context preservation, enabling unbiased detection and localization of RNA species at single-cell and subcellular levels within native tissue architecture. The platform is purpose-built for translational and discovery research in neuroscience, oncology, immunology, infectious disease, and developmental biology—fields where cellular identity, functional state, and microenvironmental interactions must be resolved simultaneously.
Key Features
- Fully automated workflow—from tissue loading to high-resolution image reconstruction—with <1 hour hands-on time per run
- Synthetic Aperture Optics (SAO) technology delivering effective resolution equivalent to 100× oil-immersion imaging using a robust 20× air objective
- Onboard fluidic automation handling all reagent delivery, wash cycles, and hybridization steps with temperature-controlled flow cell (±0.1°C stability)
- Integrated refrigeration and rapid thermal regulation in the imaging chamber to maintain optimal reaction kinetics and minimize RNA degradation
- End-to-end software suite (Esper Control, Esper Process, Esper Explore) validated for reproducible spot detection, nuclear segmentation (DAPI-based), and feature-to-cell assignment
- Output format compliant with standard single-cell analysis pipelines: tissue-wide Cell × Feature matrices containing >100,000 cells and >1 million molecular features
Sample Compatibility & Compliance
Rebus ESPER supports formalin-fixed paraffin-embedded (FFPE) and fresh-frozen tissue sections (4–20 µm thickness) mounted on standard glass slides. The platform accommodates standard histological staining workflows—including H&E and IHC—prior to spatial profiling, ensuring compatibility with routine pathology practice. All onboard protocols adhere to GLP-aligned documentation standards; audit trails, user access logs, and parameter versioning are embedded in Esper Control and Esper Process software. While not certified under FDA 21 CFR Part 11 out-of-the-box, the system architecture supports configuration for regulated environments—including electronic signature enforcement, data integrity validation, and raw data immutability—upon customer-specific qualification (IQ/OQ/PQ).
Software & Data Management
The Esper software ecosystem comprises three modular applications: Esper Control manages hardware orchestration, ROI selection, and protocol execution; Esper Process performs GPU-accelerated image reconstruction, RNA spot detection via deep learning–enhanced computer vision, DAPI-based nuclear segmentation, and spatial feature assignment using deterministic graph-based clustering; Esper Explore—a Napari-based open-source visualization engine—enables interactive 3D rendering, multi-layer overlay (e.g., RNA density + protein expression + morphology), and manual curation of segmentation boundaries. All processed data exports follow HDF5 and Loom standards, ensuring seamless integration with Seurat, Scanpy, and Space Ranger. Raw image stacks and metadata are stored with embedded MIAME-compliant annotations.
Applications
- Neuroscience: Mapping neuronal subtype distributions and synaptic gene expression gradients across laminar cortical structures
- Oncology: Resolving tumor–stroma–immune spatial niches, identifying therapy-resistant clones, and correlating transcriptomic heterogeneity with histopathological grade
- Immunology: Quantifying immune cell infiltration patterns, checkpoint ligand co-expression domains, and antigen-presenting cell–T cell interface dynamics
- Developmental Biology: Tracking lineage-restricted gene expression trajectories across embryonic tissue compartments with micron-scale precision
- Infectious Disease: Localizing pathogen RNA alongside host response signatures in infected tissue microenvironments
FAQ
What tissue preparation methods are compatible with Rebus ESPER?
FFPE and fresh-frozen sections (4–20 µm) on standard charged or silanized glass slides are supported. Prior H&E or IHC staining does not interfere with downstream spatial transcriptomic profiling.
Does the system require specialized training to operate?
No. The platform is designed for bench scientists with basic histology and molecular biology experience. All operational steps—from slide loading to data export—are guided via intuitive touchscreen interface with real-time feedback.
Can Rebus ESPER data be integrated into existing single-cell RNA-seq analysis pipelines?
Yes. Output matrices conform to standard formats (Loom, HDF5) and include UMAP coordinates, cluster labels, and spatial coordinates—enabling direct import into Seurat, Scanpy, or custom Python/R workflows.
Is the SAO optical system serviceable in the field?
The SAO module is a sealed, alignment-free subsystem with no user-serviceable components. Preventive maintenance is performed remotely via diagnostic telemetry and scheduled on-site calibration every 12 months.
How is data integrity ensured during long-duration runs (e.g., >48 hours)?
The system logs all hardware states, reagent consumption timestamps, and thermal profiles in real time. Raw image frames are checksum-verified before reconstruction; any corrupted frame triggers automatic reacquisition without user intervention.