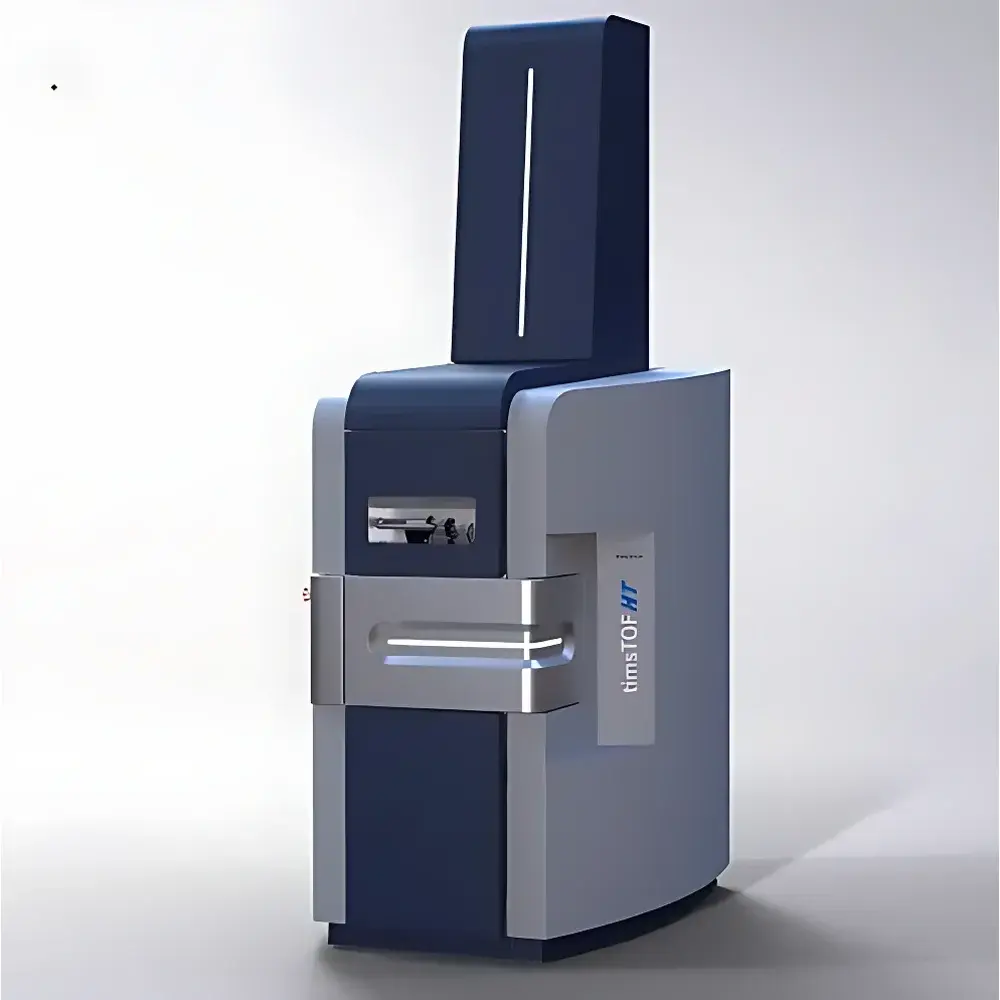
Bruker timsTOF HT Trapped Ion Mobility Time-of-Flight Mass Spectrometer
| Brand | Bruker |
|---|---|
| Origin | Germany |
| Manufacturer Type | Authorized Distributor |
| Origin Category | Imported |
| Model | timsTOF HT |
| Instrument Type | Trapped Ion Mobility Spectrometry (TIMS) Mass Spectrometer |
| Technology | Parallel Accumulation–Serial Fragmentation (PASEF®), CCS-enabled 4D-Proteomics |
Overview
The Bruker timsTOF HT is a high-performance trapped ion mobility spectrometry coupled with time-of-flight mass spectrometry (TIMS-TOF) platform engineered for deep, quantitative, and reproducible 4D-proteomics. Unlike conventional drift-tube or traveling-wave ion mobility systems, the timsTOF HT employs a dual-stage, radiofrequency-driven TIMS analyzer that traps and elutes ions based on their collisional cross-section (CCS) values—providing an orthogonal, gas-phase separation dimension prior to quadrupole selection and TOF mass analysis. This architecture enables true multidimensional data acquisition where retention time, m/z, intensity, and CCS jointly define each peptide feature—hence the “4D” designation. The system integrates fourth-generation TIMS-XR technology with enhanced ion capacity, digital signal processing, and optimized ion optics to deliver extended dynamic range, high duty cycle, and robust performance across thousands of consecutive LC-MS runs without routine maintenance.
Key Features
- PASEF® Acquisition Engine: Enables ultra-high-speed, duty-cycle-optimized MS/MS scanning at >150 Hz without sacrificing resolution or sensitivity—ideal for short-gradient LC separations and high-throughput workflows.
- TIMS-XR Analyzer: Fourth-generation trapped ion mobility cell with significantly increased ion storage capacity, improving detection of low-abundance peptides and extending proteome coverage depth in complex samples.
- CCS-Enabled Data Dimensionality: Provides experimentally derived collisional cross-section values for every identified precursor, enhancing confidence in peptide identification, improving chromatographic peak alignment, and enabling cross-laboratory reproducibility.
- Full PASEF® Mode Support: Native compatibility with data-dependent (PASEF®), data-independent (dia-PASEF®), and targeted (prm-PASEF®) acquisition strategies—allowing flexible experimental design from discovery to validation.
- Robust Hardware Architecture: Designed for unattended, long-term operation; validated for stability over >2,000 consecutive injections with minimal signal drift or calibration drift—reducing instrument downtime and maintenance overhead.
Sample Compatibility & Compliance
The timsTOF HT is compatible with nanoflow liquid chromatography systems operating at flow rates from 50–300 nL/min, including columns such as PepSep 25 cm (150 µm ID, 1.5 µm particles) and Aurora series. It supports sample inputs ranging from sub-microgram to >1.6 µg of tryptic digest, maintaining sub-5-second peak widths even at high loading. The platform complies with analytical requirements for GLP- and GMP-aligned workflows: raw data files are timestamped, digitally signed, and structured to support audit trails. While not inherently 21 CFR Part 11 compliant out-of-the-box, the system integrates with LIMS and electronic lab notebook (ELN) platforms supporting FDA-regulated environments when deployed with appropriate IT governance controls. Method parameters, acquisition logs, and CCS calibration metadata are fully traceable and exportable in open formats (e.g., mzML, imzML).
Software & Data Management
Data acquisition is controlled via Bruker’s SCiLS Lab and Compass software suites, supporting real-time method optimization and automated calibration. For proteomics, the timsTOF HT is natively integrated with PaSER—a real-time database search engine delivering “run-and-done” identification with on-the-fly spectral matching and TIMScore™-weighted confidence scoring. Quantitative analysis leverages TIMS DIA-NN, a CCS-aware version of the open-source DIA-NN engine, enabling library-free or spectral-library-based quantification with improved specificity through CCS filtering. All CCS values are stored in standardized metadata fields, facilitating cross-platform comparison and meta-analysis in repositories such as ProteomeXchange. Raw data export supports FAIR principles (Findable, Accessible, Interoperable, Reusable) and is compatible with downstream tools including MaxQuant, FragPipe, and Skyline.
Applications
- Deep Discovery Proteomics: Achieves >4,700 protein identifications from 800 ng of mouse heart tissue digest using 30-min LC gradients and dia-PASEF®—spanning five orders of magnitude in relative abundance.
- High-Throughput Cell Line Profiling: With 60-min gradients and Aurora-25 cm columns, identifies >10,000 peptides and >5,500 proteins from K562 lysates, leveraging TIMS DIA-NN for label-free quantification and CCS-guided false discovery rate control.
- Targeted Plasma Proteomics: Combined with Biognosys PQ500 spike-in standards, quantifies 578 plasma proteins (804 peptides) in 30-min gradients with >99% data completeness—bridging the selectivity of SRM/MRM with the scalability of DIA.
- Post-Translational Modification (PTM) Analysis: Enhanced peak capacity from TIMS separation improves co-eluting phosphopeptide resolution, enabling confident site localization without offline fractionation.
- Single-Cell & Limited-Sample Workflows: High sensitivity and low background enable robust analysis of <100 ng inputs when coupled with microfluidic LC or trap-elute configurations.
FAQ
What distinguishes timsTOF HT from earlier timsTOF models?
The timsTOF HT introduces the TIMS-XR analyzer with higher ion capacity, upgraded digitizer electronics, and refined ion optics—resulting in improved sensitivity at high throughput and greater robustness for multi-day unattended operation.
Is CCS calibration required for every run?
No—CCS calibration is performed during system setup using standard calibrants (e.g., Agilent ESI Tune Mix); subsequent runs inherit the calibration unless major hardware changes occur.
Can timsTOF HT perform traditional DDA or DIA without PASEF®?
Yes—the instrument supports conventional data-dependent acquisition (DDA) and standard DIA modes, though PASEF®-enabled methods are recommended to fully exploit the TIMS dimension.
Does the system support retention time alignment across batches using CCS?
Yes—CCS serves as a stable, instrument-independent physicochemical property; TIMS DIA-NN and PaSER use CCS to improve chromatographic alignment and reduce missing value rates in large cohort studies.
What file formats are generated and how are they archived?
Raw data are saved in Bruker’s proprietary .d format; conversion to open standards (mzML, imzML) is supported via CompassXport, with metadata including CCS, mobility time, and PASEF® event timing preserved.