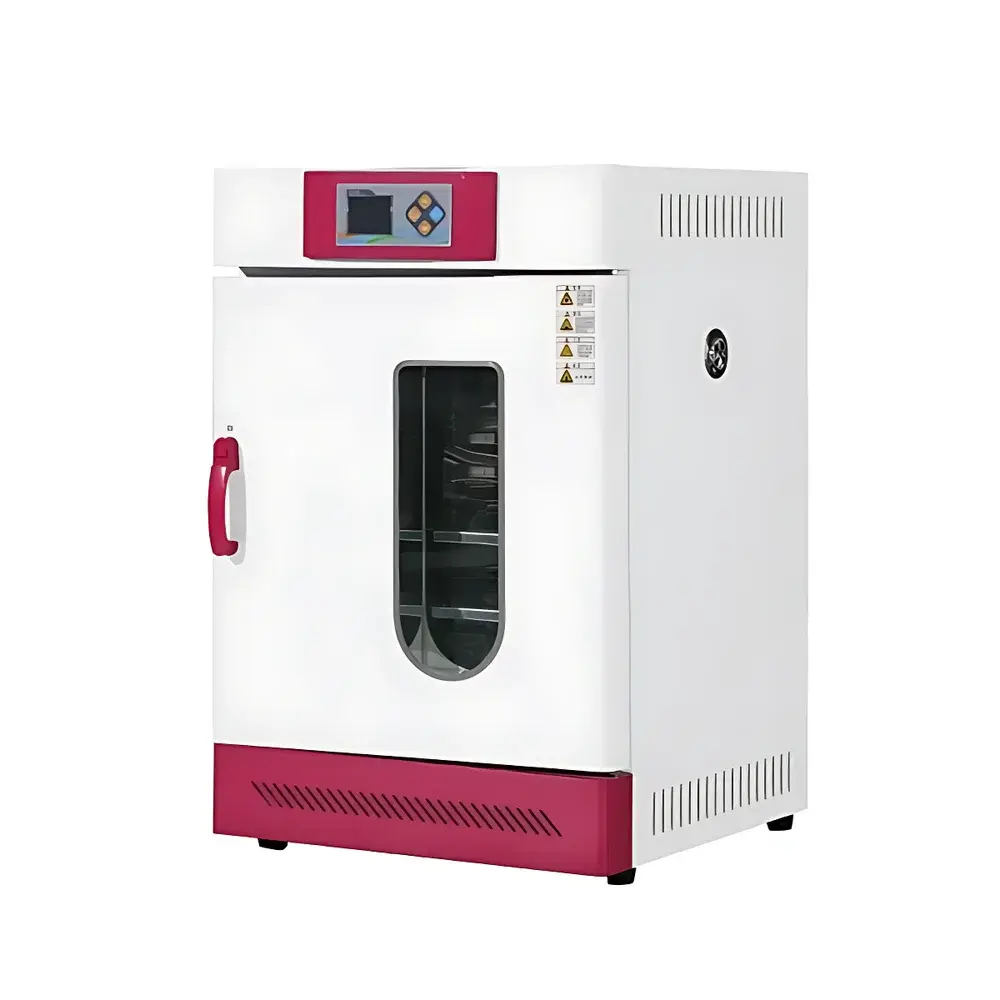
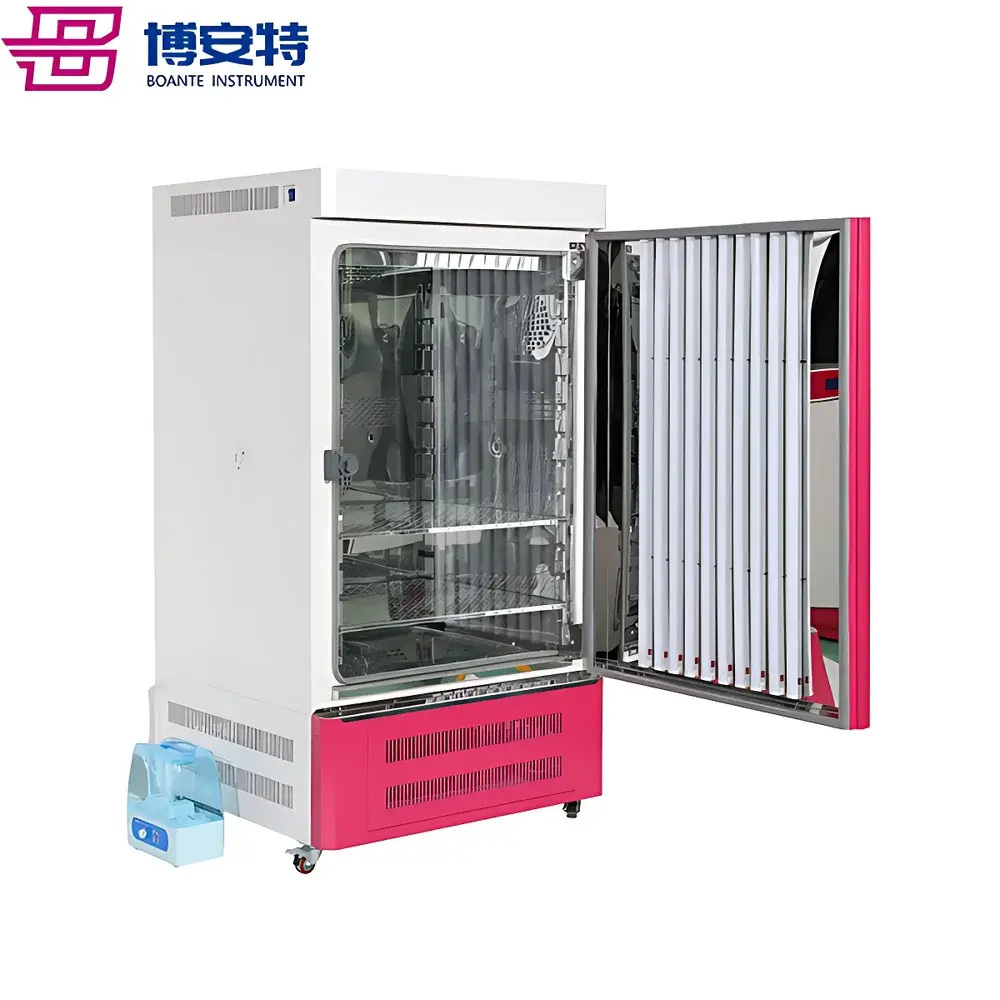
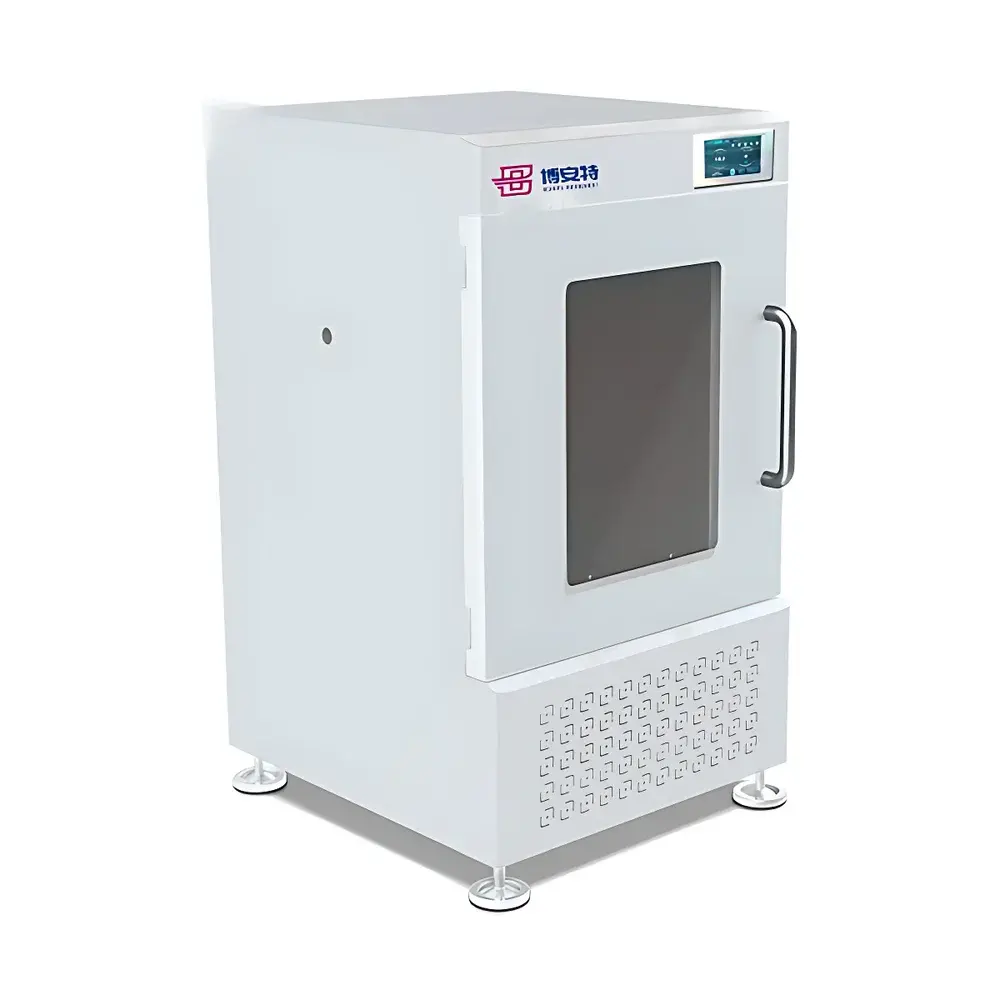
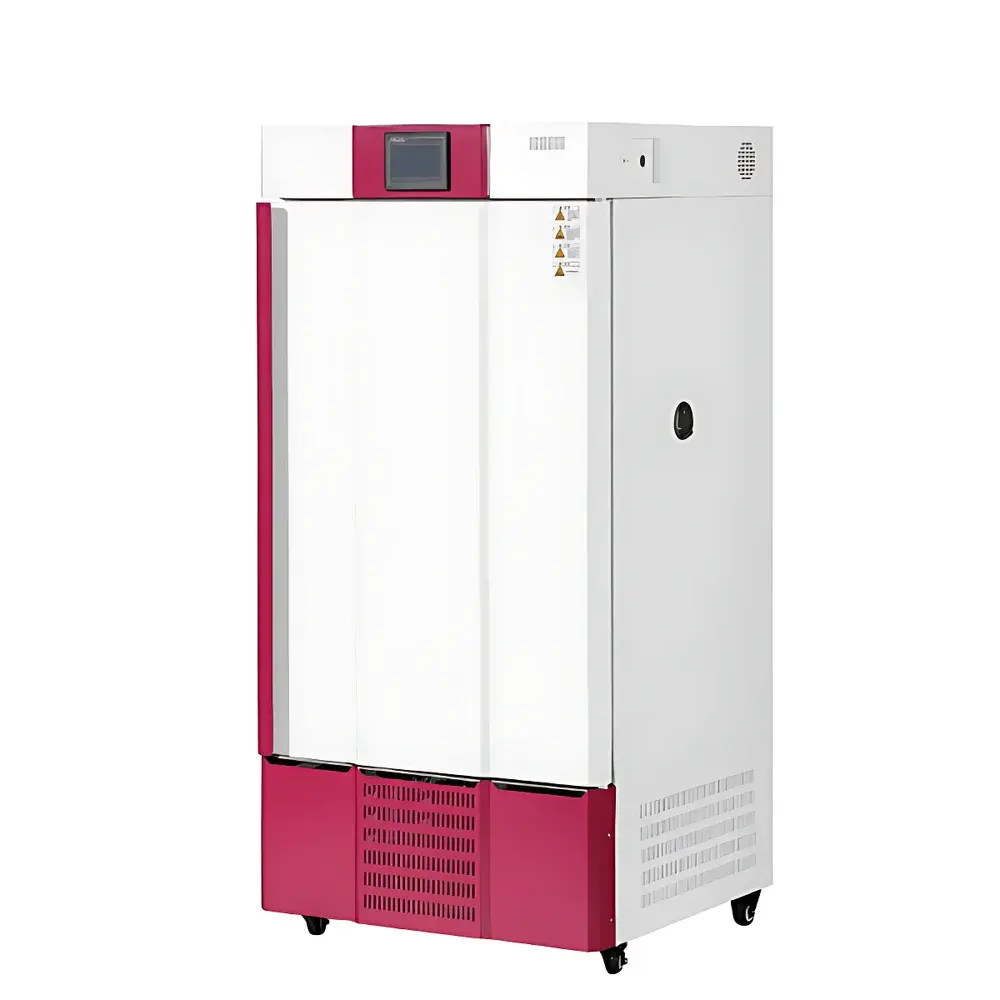
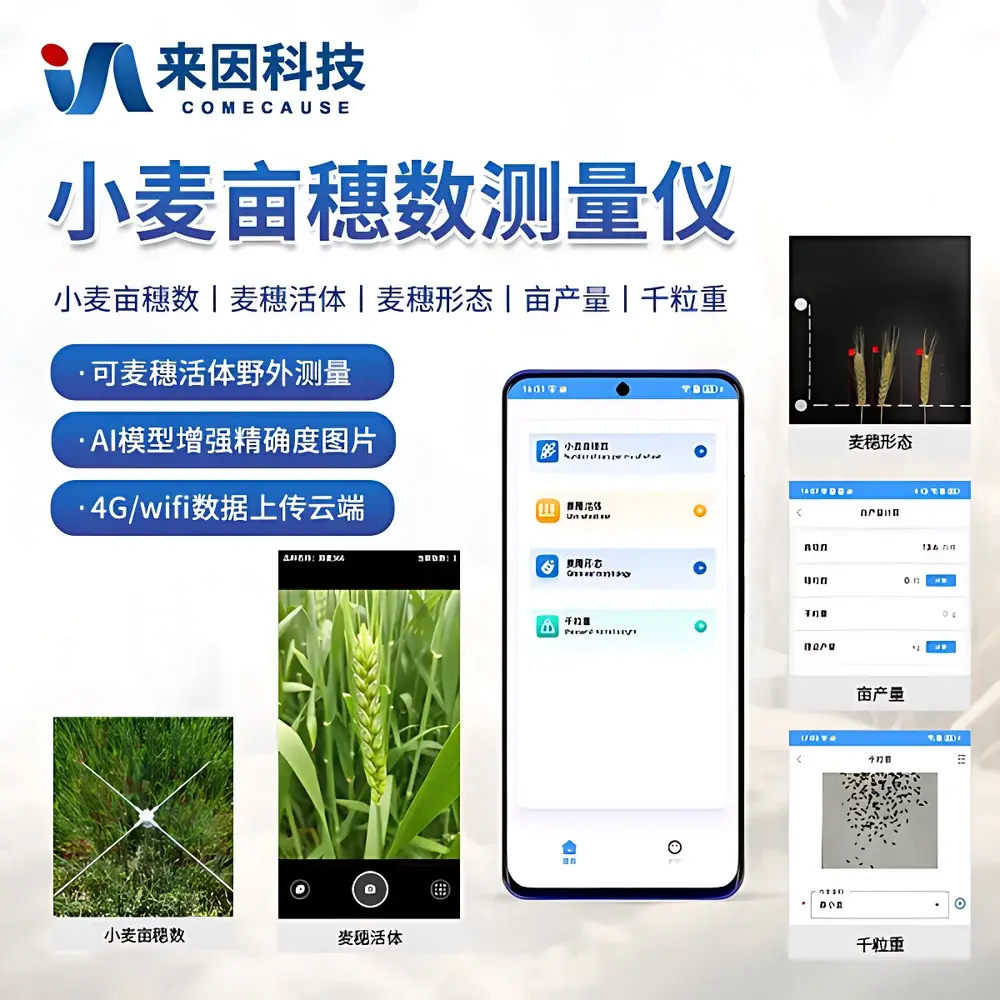
COMECAUSE IN-XM01 Intelligent Wheat Phenotyping System
| Brand | COMECAUSE |
|---|---|
| Origin | Shandong, China |
| Manufacturer Type | OEM Manufacturer |
| Product Origin | Domestic (China) |
| Model | IN-XM01 |
| Price | USD 3,500 (FOB) |
| Camera Resolution | 4000 × 3000 |
| Dual-Camera Setup | 50 MP + 12 MP |
| Measurement Accuracy (Spike Count) | ≤ ±3% |
| Live-Field Spike Detection Error | ±1% |
| Thousand-Kernel Weight (TKW) Accuracy | ±2% (calibrated to ≤0.5% residual error) |
| Adjustable Calibration Rod Range | 1000 mm × 1000 mm × (610–1120) mm & 1000 mm × 1000 mm × (1010–1920) mm |
| On-Device Storage | 256 GB |
| Operating System | Android-based UI with touch interface |
| Connectivity | Wi-Fi / 4G LTE |
| Data Export Format | Excel (.xlsx) with embedded GPS timestamps |
| Language Support | English & Chinese (toggleable) |
Overview
The COMECAUSE IN-XM01 Intelligent Wheat Phenotyping System is a field-deployable, AI-powered phenotyping platform engineered for high-throughput, non-destructive quantification of key agronomic traits in wheat (Triticum aestivum). It operates on the principle of computer vision–driven morphometric analysis, combining calibrated multi-spectral imaging, geometric normalization, and convolutional neural network (CNN)-based object detection to extract biologically meaningful phenotypic parameters—including spike count per unit area (spikes/m²), spike length, grain number per spike, thousand-kernel weight (TKW), and dynamic growth-stage indexing. Unlike conventional manual or semi-automated methods, the IN-XM01 eliminates observer bias through standardized image acquisition protocols, automatic perspective correction, and environment-robust white balance adaptation. Its architecture supports both live-field measurement (no background required) and controlled-lab sampling modes, enabling longitudinal monitoring across critical developmental stages—from late booting through grain filling to physiological maturity. The system is designed for integration into breeding pipelines compliant with FAO crop evaluation guidelines and aligned with ISO 20671 (phenotyping data quality standards) and OECD Seed Sowing and Harvest Protocols.
Key Features
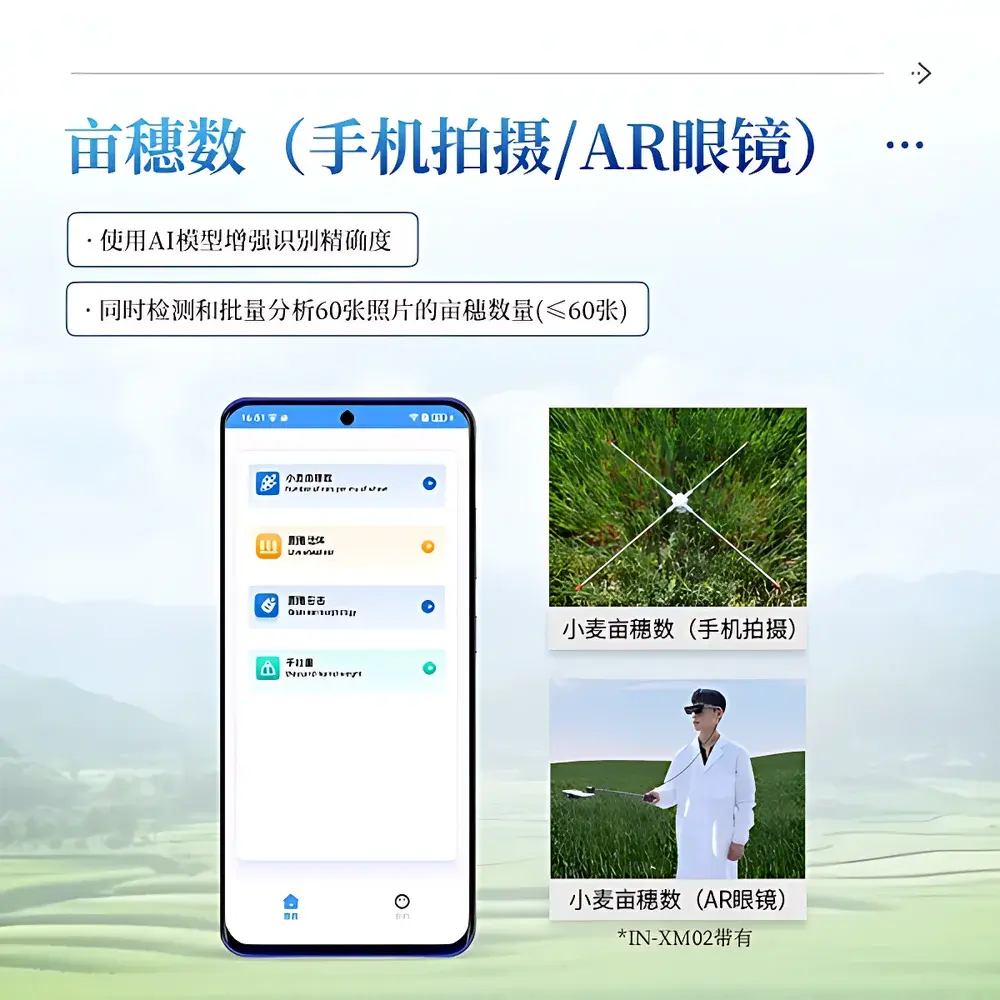
- AI-enhanced spike detection and counting using a dual-camera array (50 MP primary sensor + 12 MP auxiliary sensor) with real-time inference latency <3 seconds per image
- Calibration rod–guided spatial referencing for absolute metric scaling under variable field conditions; height-adjustable dual-range rod (610–1120 mm and 1010–1920 mm)
- Live-field mode: spike segmentation without physical backboards; adaptive illumination compensation for variable solar irradiance and cloud cover
- Laboratory sampling mode: automated image rectification for tilted or oblique captures; black matte acrylic reference board (500 × 400 × 5 mm) enabling simultaneous measurement of up to 10 spikes
- Integrated GPS tagging and timestamping for georeferenced phenotypic records; metadata export includes location, acquisition time, device orientation, and environmental context flags
- On-device Android OS with intuitive touchscreen interface; bilingual (English/Chinese) UI with one-touch language switching
- Local storage capacity: 256 GB SSD for offline operation in low-connectivity environments; batch processing of up to 60 images per session
Sample Compatibility & Compliance
The IN-XM01 is validated for use with common bread wheat (T. aestivum) cultivars grown under temperate and semi-arid agroecological zones. Optimal performance is achieved during the grain-filling to pre-maturity window (Zadoks growth stages 73–87), when spike morphology is fully expressed but before shattering or lodging occurs. The system accommodates variations in canopy density, spike orientation, and light scattering due to dew or dust via spectral preprocessing and contrast normalization algorithms. All measurement outputs conform to FAO’s Wheat Phenotyping Minimum Data Set (v2.1) and support traceability requirements under GLP-aligned field trial documentation. While not certified for regulatory submission under FDA 21 CFR Part 11, the audit trail functionality—including immutable image hashes, user login logs, and versioned AI model metadata—enables internal validation for GxP-compliant research institutions.
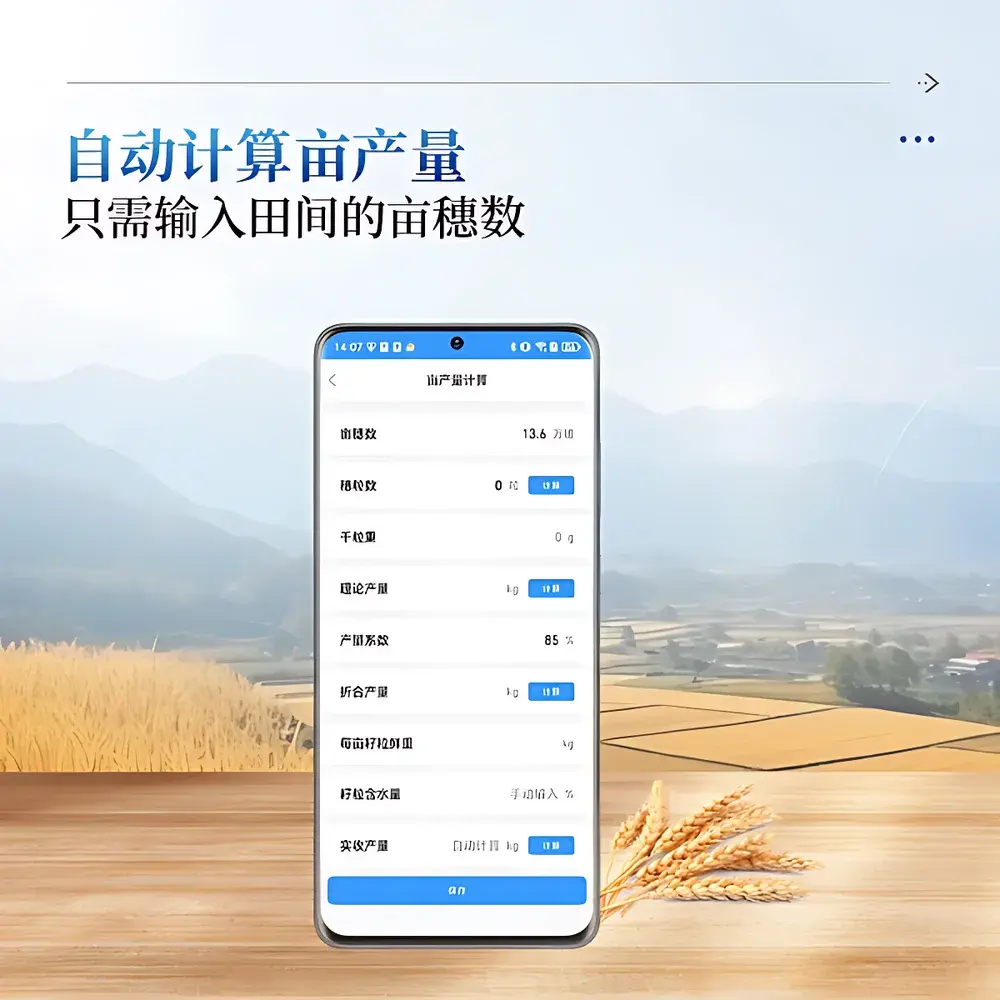
Software & Data Management
Data acquisition, annotation, and export are managed through the proprietary COMECAUSE Phenotype Manager (CPM) software suite. CPM implements role-based access control (RBAC) and encrypted local database storage. Field-collected images and derived metrics synchronize automatically to secure cloud infrastructure (AWS-hosted S3 buckets with TLS 1.3 encryption) when Wi-Fi or 4G connectivity is available. All exports include structured Excel (.xlsx) files containing raw pixel coordinates, normalized trait values, confidence scores per detection, and QC flags (e.g., occlusion index, lighting uniformity score). API endpoints support integration with third-party platforms including BreedBase, T3 (Trait Ontology), and BrAPI-compliant databases. Software updates are delivered over-the-air (OTA) with SHA-256 signature verification.
Applications
- High-throughput yield component analysis in early-generation breeding nurseries (e.g., F₂–F₄ populations)
- Quantitative trait locus (QTL) mapping studies requiring precise spike architecture descriptors
- Abiotic stress phenotyping (drought, heat, salinity) via temporal tracking of spikelet fertility and grain set stability
- Pathogen resistance screening by correlating spike morphology deviations (e.g., shriveled grains, necrotic rachis segments) with disease severity indices
- Seed certification workflows requiring TKW verification against national seed quality standards (e.g., GB/T 3543.3–1995)
- Extension service deployment for regional yield forecasting models incorporating real-time spike density inputs
FAQ
What growth stages are optimal for spike counting with the IN-XM01?
The system delivers highest accuracy between Zadoks stages 73 (mid-grain filling) and 85 (hard dough), when spike architecture is stable and grain boundaries are optically distinct.
Can the IN-XM01 be used for other small-grain cereals such as barley or rye?
While trained primarily on wheat, the underlying CNN architecture supports transfer learning; users may retrain the detector on custom barley or rye datasets using the CPM annotation toolkit.
Is calibration required before each field session?
Yes—each session requires placement of the adjustable calibration rod within the frame to establish scale invariance; the system validates rod detection confidence prior to analysis.
How does the device handle overlapping or partially occluded spikes?
The AI model incorporates instance segmentation with mask refinement heuristics; occlusion-aware scoring reduces false negatives in dense canopies while flagging low-confidence detections for manual review.
Does the system support integration with genomic data pipelines?
Yes—CPM exports trait vectors compatible with TASSEL, GAPIT, and R/qtl input formats; genotype-phenotype association modules are available as optional add-ons.