COMECAUSE IN.Pheno200 High-Spectral Plant Phenotyping Imaging System
| Brand | COMECAUSE |
|---|---|
| Origin | Shandong, China |
| Manufacturer Type | Direct Manufacturer |
| Country of Origin | China |
| Model | IN.Pheno200 |
| Price | USD 138,000 (FOB Qingdao) |
| Plant Height Range | 300–1000 mm |
| Plant Width Range | 100–500 mm |
| Fresh Weight Capacity | 100–75,000 g |
| Hyperspectral Range | 400–1000 nm |
| Spectral Resolution | <2.5 nm |
| Spectral Bands | 1200 channels |
| Illumination Source | Low-flicker, high-CRI halogen lamp array |
| Enclosure Dimensions | 1500 × 830 × 2140 mm (L×W×H) |
| Camera Resolution | Up to 12 MP (RGB), hyperspectral sensor native resolution: 1392 × 1040 px |
| Data Acquisition Time | 40 s per plant |
| 3D Reconstruction Time | ≤30 s per plant |
| Operating System | Windows 10/11 (64-bit), Intel Core i5 or higher, ≥8 GB RAM, NVIDIA GPU with ≥8 GB VRAM |
Overview
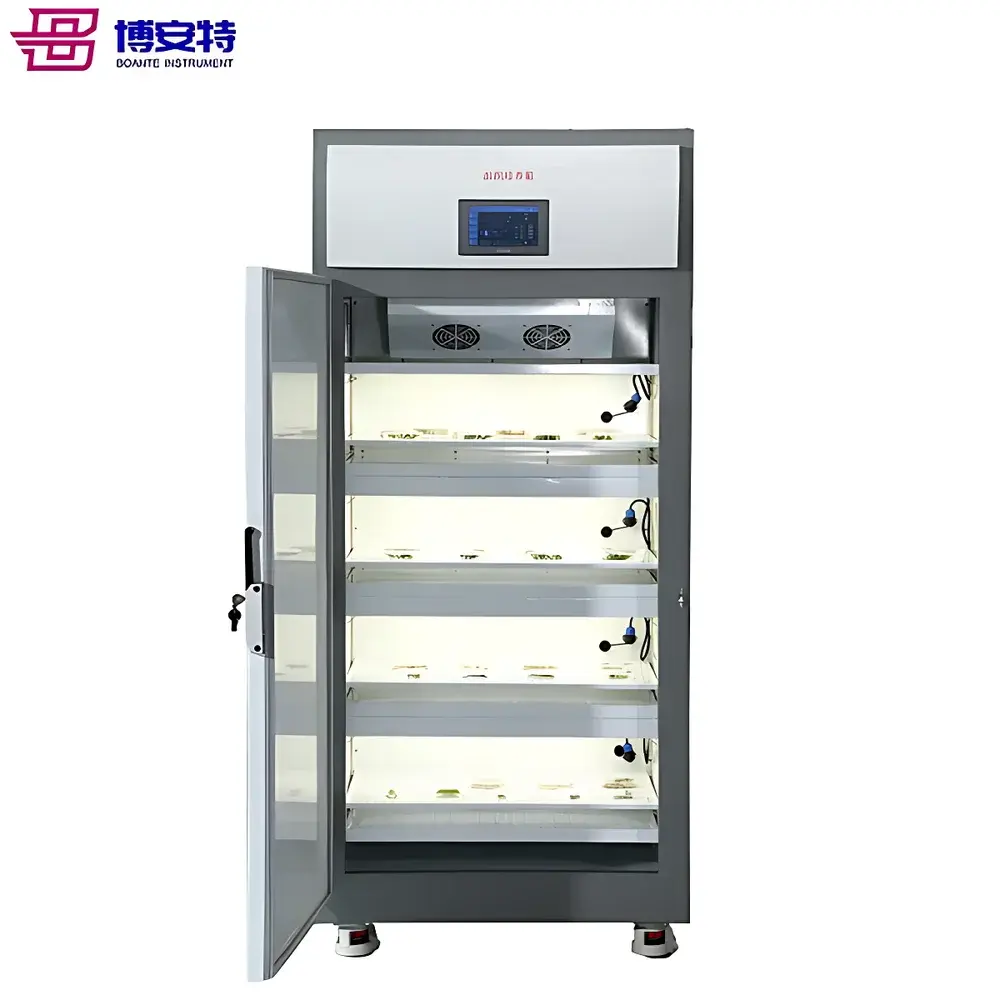
The COMECAUSE IN.Pheno200 High-Spectral Plant Phenotyping Imaging System is a fully integrated, multi-modal phenotyping platform engineered for non-destructive, quantitative assessment of plant morphological, chromatic, textural, and biochemical traits under controlled environmental conditions. It operates on the principle of multi-angle visible-light photogrammetry combined with push-broom hyperspectral imaging in the 400–1000 nm range, enabling simultaneous acquisition of spatially registered RGB, Lab, HSV, and reflectance spectra across the full visible–NIR spectrum. The system employs a motorized rotating turntable and synchronized vertical/horizontal positioning stages to acquire top-, side-top-, and side-bottom-view images—ensuring complete 360° surface coverage without occlusion. Its AI-accelerated 3D reconstruction pipeline uses structure-from-motion (SfM) and multi-view stereo (MVS) algorithms to generate geometrically accurate, texture-mapped point clouds and mesh models, from which volumetric, angular, and topological descriptors—including canopy volume, branch angle distribution, skeleton topology, and fractal dimension—are extracted with sub-millimeter reproducibility. Designed for longitudinal studies, the IN.Pheno200 supports repeatable, operator-independent measurements compliant with FAIR (Findable, Accessible, Interoperable, Reusable) data principles and GLP-aligned experimental workflows.
Key Features
- Fully automated imaging protocol: Real-time plant size detection triggers adaptive image capture; automatic mosaic stitching for specimens exceeding field-of-view dimensions.
- Integrated environmental monitoring: Onboard temperature and humidity sensors provide continuous logging with timestamped metadata for environmental correlation analysis.
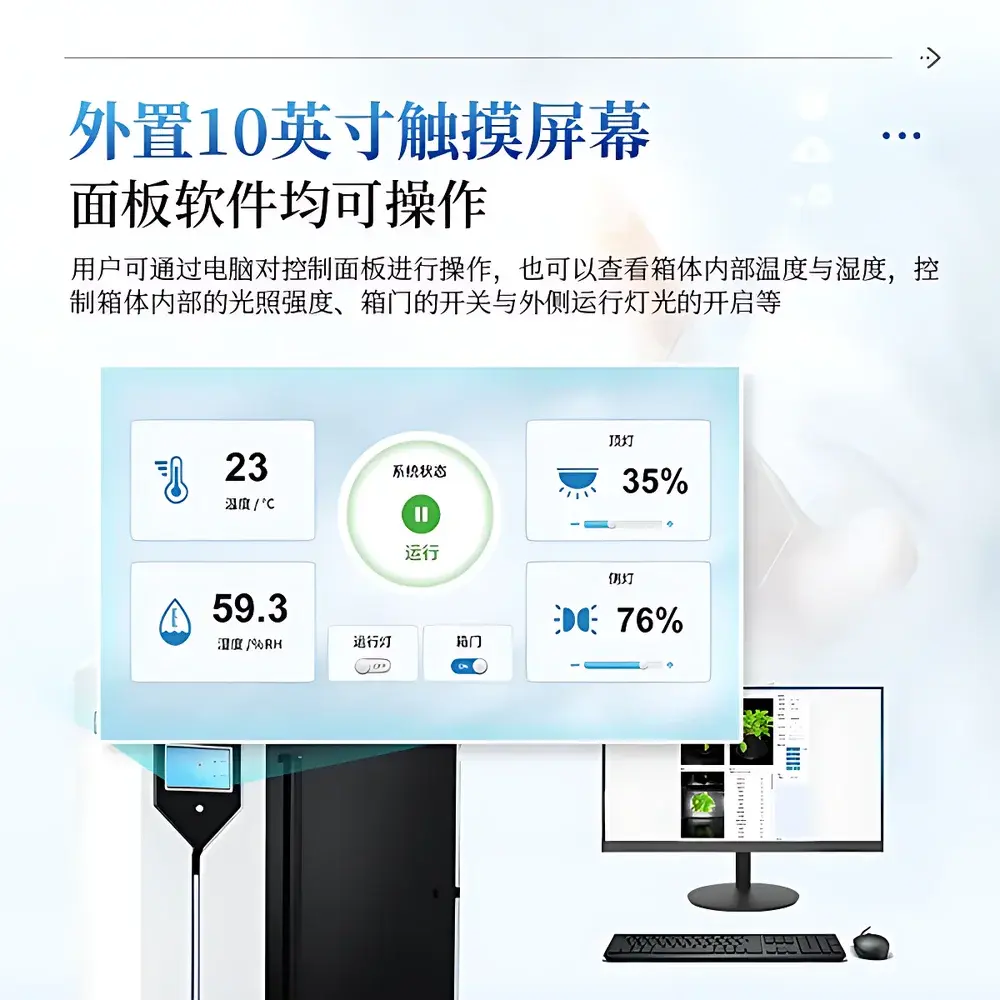
- Modular hardware control: 10-inch industrial touchscreen interface enables direct adjustment of illumination intensity, door actuation, and chamber lighting—complemented by remote PC-based control via Ethernet-connected software suite.
- Motorized optical positioning: Software-controlled Z-axis lift mechanism adjusts camera height and stage elevation to maintain optimal working distance across variable plant sizes (300–1000 mm).
- Dual-mode analysis architecture: Parallel 2D high-throughput mode (≤60 s/sample) and on-demand 3D modeling module operate independently—enabling continuous sample throughput while background 3D reconstruction proceeds without interrupting acquisition.
- Cloud-synced data management: Device-registered datasets—including raw hyperspectral cubes (.spe), processed parameter tables (CSV/JSON), and annotated 3D model exports (OBJ/GLTF)—are encrypted and uploaded to secure cloud storage with role-based access control.
- Hardware-level safety compliance: Mechanical limit switches, infrared proximity sensors, and emergency stop interlocks conform to IEC 61000-6-2/6-4 EMC standards and ISO 13857 safety distances for enclosed laboratory equipment.
Sample Compatibility & Compliance
The IN.Pheno200 accommodates intact, living plants from major angiosperm families—including Poaceae (e.g., rice, wheat, maize), Solanaceae (tomato, tobacco), Brassicaceae (Arabidopsis, rapeseed), and Fabaceae (soybean, pea)—across vegetative and early reproductive stages. Specimen handling requires no tissue excision or staining; root systems remain undisturbed in standard growth pots (diameter ≤500 mm). All optical components meet ISO 9022-3:2015 specifications for spectral transmittance stability under thermal cycling. Hyperspectral calibration follows NIST-traceable procedures using certified reflectance standards (Labsphere Spectralon® SRM-990) and is validated prior to each acquisition session. Data output formats comply with MIAPPE v1.1 metadata standards and support export to Crop Ontology (CO) and Plant Trait Ontology (TO) identifiers. The system meets requirements for ISO/IEC 17025-accredited laboratories when operated within documented SOPs and maintained with annual radiometric recalibration.
Software & Data Management
The proprietary IN.PhenoSuite™ software (v4.2+) provides a unified interface for instrument control, real-time visualization, and parameter computation. It implements FDA 21 CFR Part 11-compliant audit trails—including user login history, parameter modification logs, and export timestamps—with optional digital signature integration. All hyperspectral indices—including NDVI, EVI, TCARI, MCARI, PRI, ARI, WI, and PSRI—are calculated using peer-reviewed, publication-standard equations (e.g., Gitelson et al. 2003; Barnes et al. 2000; Gamon et al. 1992) with wavelength-specific bandpass definitions embedded in the spectral response function database. Texture metrics (ASM, Contrast, Homogeneity, etc.) are computed via Gray-Level Co-occurrence Matrix (GLCM) analysis following Haralick et al. (1973), with configurable window size, offset, and quantization levels. 3D-derived morphometrics—including convex hull volume, branch angle histograms, and skeletonized path length—are generated using Open3D and VTK libraries with voxel-based smoothing and outlier rejection. Export options include FAIR-aligned JSON-LD metadata bundles, time-series CSV matrices, and video-rendered 3D model rotations (MP4/H.264) compatible with downstream platforms such as R Shiny, Python Dash, or commercial LIMS systems.
Applications
The IN.Pheno200 serves as a core infrastructure tool in academic and industrial plant science programs focused on quantitative trait locus (QTL) mapping, genome-wide association studies (GWAS), abiotic stress phenotyping (drought, heat, salinity), biotic stress screening (pathogen resistance), nutrient use efficiency evaluation, and breeding program acceleration. Its capacity to resolve subtle spectral shifts associated with chlorophyll degradation, anthocyanin accumulation, carotenoid dynamics, and water content variation enables early detection of physiological perturbations—often preceding visible symptom onset by 2–5 days. In ecological research, it supports comparative analyses of functional trait variation across genotypes, populations, or treatment gradients. Within regulatory contexts, its traceable measurement chain and standardized index computation support submissions to EFSA, USDA APHIS, and OECD for variety registration and biosafety assessments. Integration with greenhouse climate control systems (via Modbus TCP) further enables closed-loop phenotyping experiments where environmental variables are dynamically adjusted based on real-time trait feedback.
FAQ
What spectral calibration standards are used during factory setup and field recalibration?
NIST-traceable Spectralon® reflectance standards (SRM-990, 99% reflectance) and calibrated tungsten-halogen lamps (NIST SRM-2032) are employed for absolute radiometric calibration; wavelength accuracy is verified using mercury-argon emission line sources.
Is the system compatible with third-party statistical or machine learning pipelines?
Yes—raw hyperspectral data (.spe), processed feature tables (CSV/TSV), and 3D model files (OBJ/GLTF) are natively supported by Python (scikit-learn, PyTorch), R (hyperspec, phenofit), and MATLAB; API documentation and sample notebooks are provided with each installation.
How is data integrity ensured during long-term multi-experiment deployments?
Each dataset includes embedded SHA-256 checksums, instrument configuration snapshots, environmental logs, and version-stamped software build IDs; cloud backups retain immutable copies with retention policies aligned to institutional data governance policies.
Can the system be deployed in growth chambers or walk-in growth rooms?
Yes—the compact footprint (1.5 m × 0.83 m base) and castor-mounted enclosure allow flexible placement; optional vibration-dampening feet and EMI-shielded cabling are available for integration into controlled-environment facilities.
Does the software support batch processing of historical datasets acquired on earlier versions?
IN.PhenoSuite™ v4.2+ includes backward-compatible import modules for legacy .pheno, .hsp, and .3dm files generated on IN.Pheno100 and IN.Pheno150 systems, with automated metadata migration and index recalculation using current reference equations.