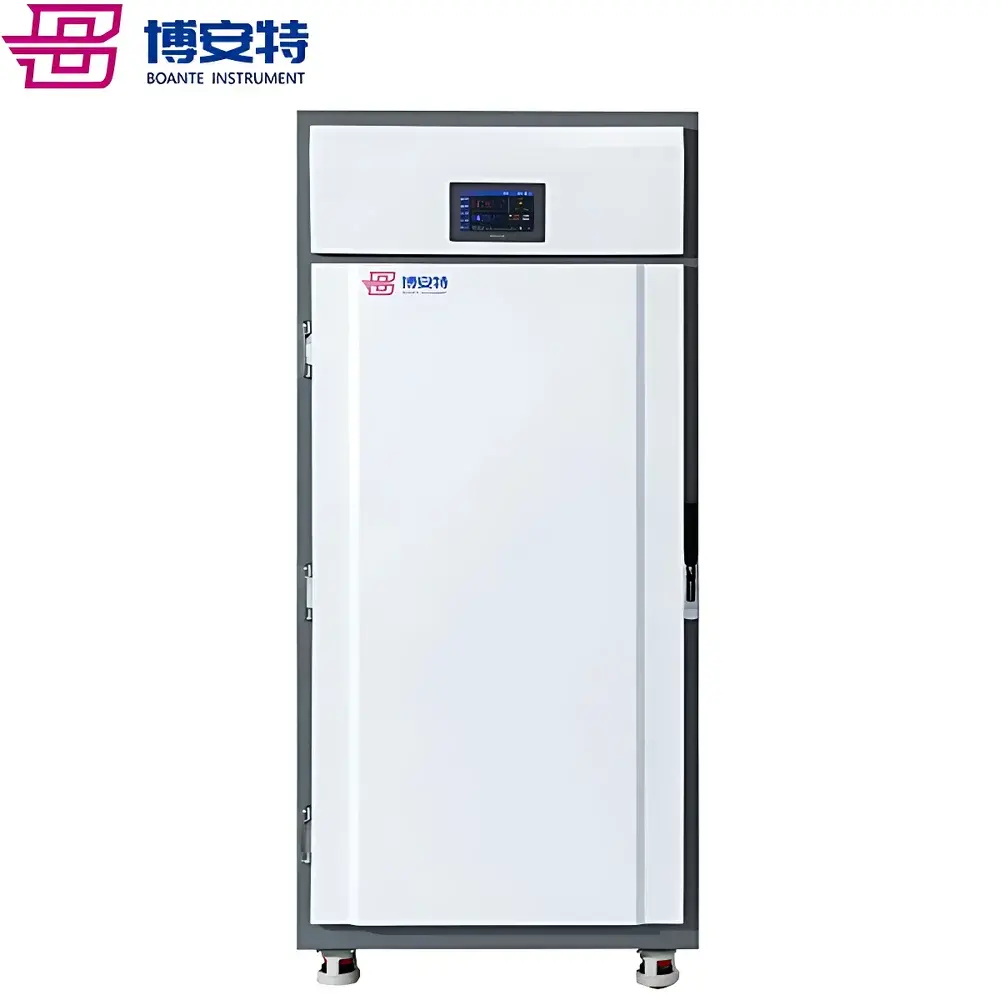
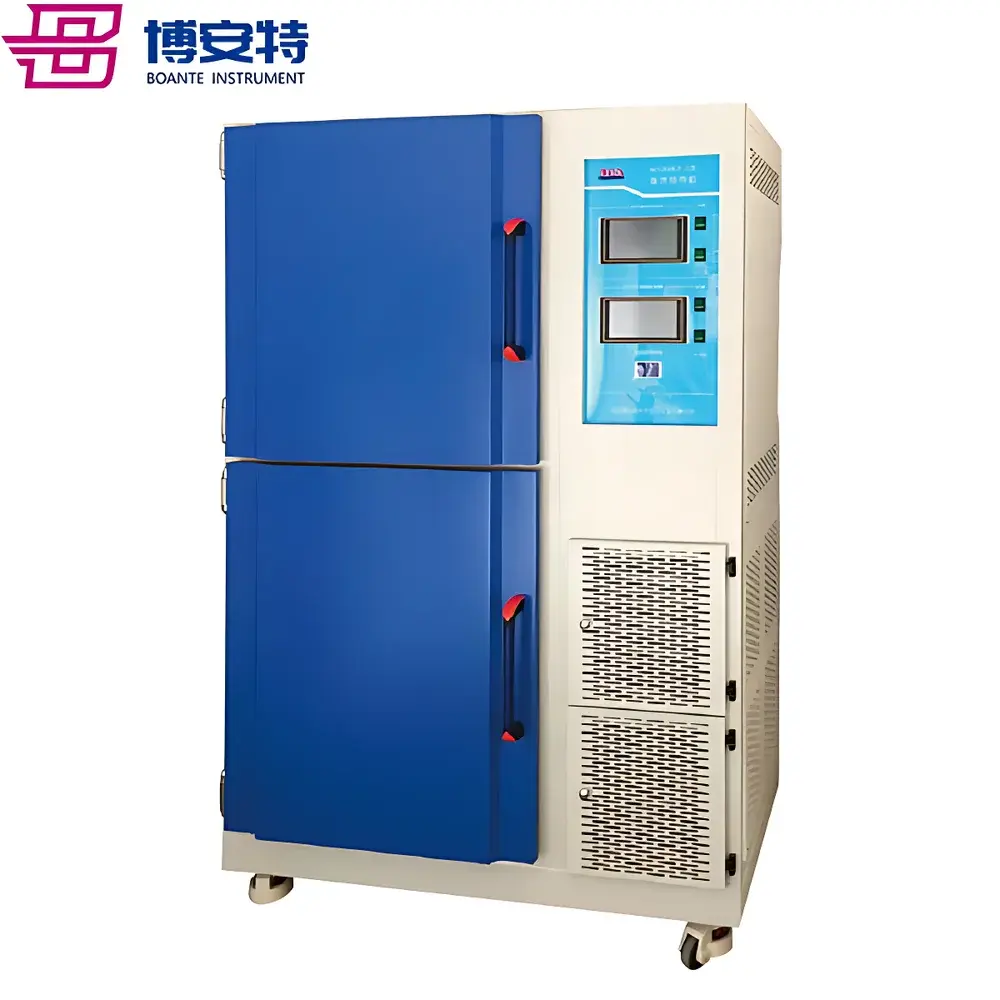
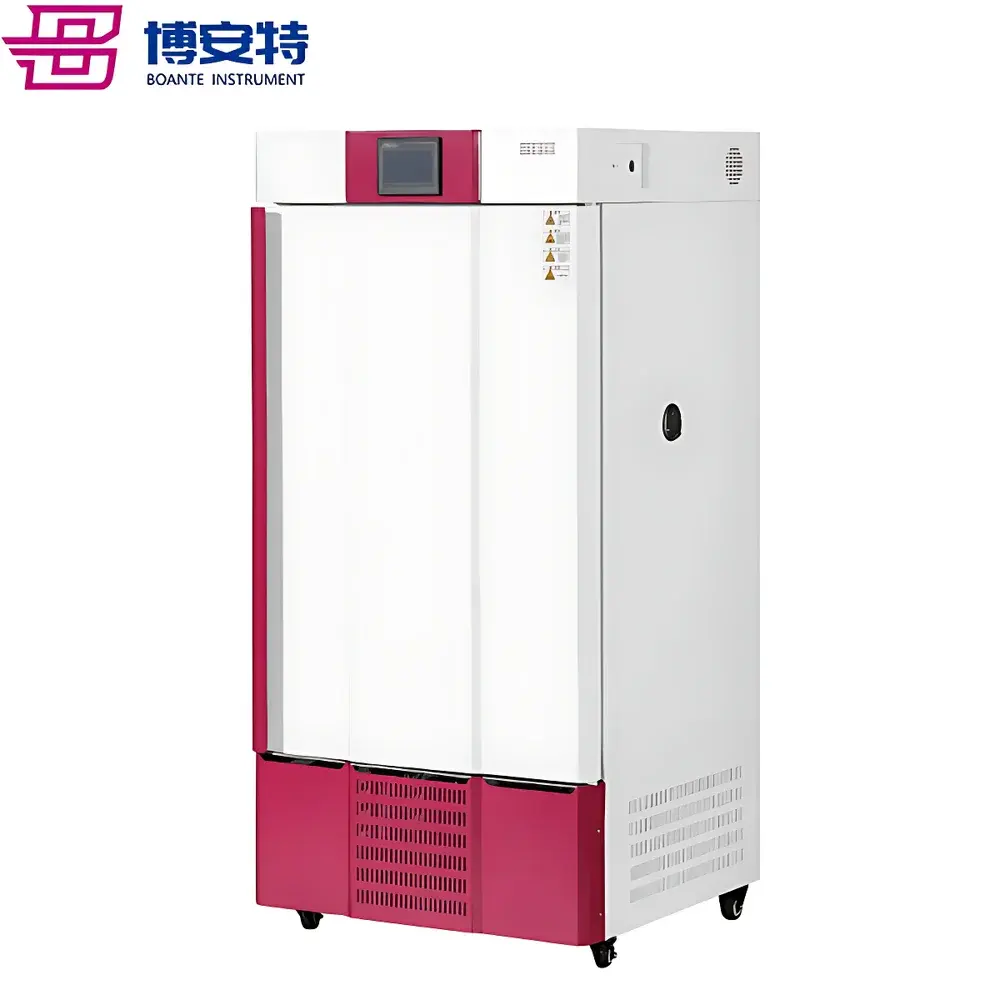
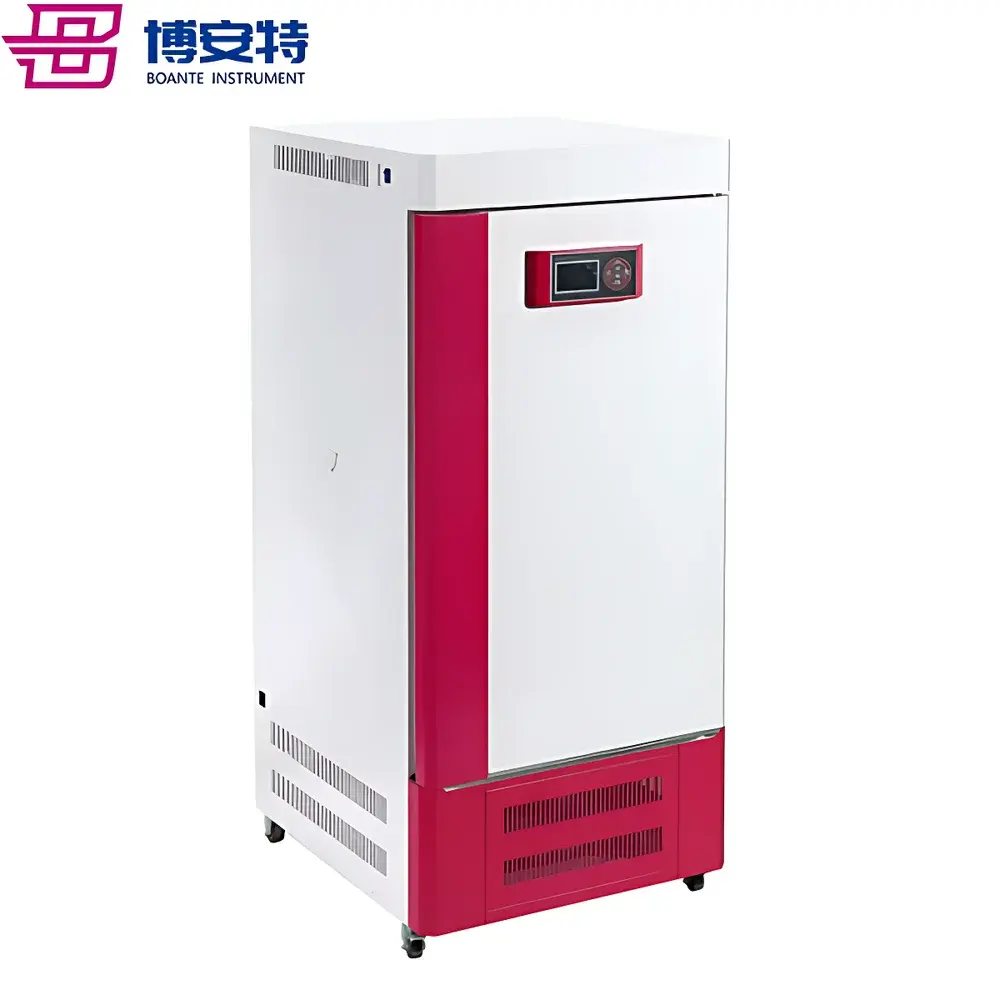
COMECAUSE IN-Pheno50 Plant Phenotyping Imaging System
| Brand | COMECAUSE |
|---|---|
| Origin | Shandong, China |
| Manufacturer Type | Original Equipment Manufacturer (OEM) |
| Country of Origin | China |
| Model | IN-Pheno50 |
| Price | USD 76,500 (FOB Qingdao, ex-works pricing |
Overview
The COMECAUSE IN-Pheno50 Plant Phenotyping Imaging System is a fully integrated, non-destructive, multi-modal phenotyping platform engineered for high-throughput, quantitative assessment of plant morphological, chromatic, and textural traits across controlled-environment conditions. It operates on the principle of multi-angle structured-light-assisted photogrammetry combined with calibrated visible-spectrum imaging (400–700 nm), enabling robust 3D reconstruction from synchronized top, upper-side, and lower-side image acquisition sequences. The system integrates a motorized rotating turntable (±0.1° angular resolution), programmable LED illumination arrays (full-spectrum white, 5000 K CCT, D65-compliant), and environmental monitoring sensors to ensure measurement repeatability under standardized physiological conditions. Designed for longitudinal studies in controlled growth chambers or greenhouse modules, the IN-Pheno50 delivers spatially resolved trait quantification aligned with FAO/CGIAR phenotyping standards and supports traceable data acquisition per GLP-aligned workflows.
Key Features
- Triple-axis visible-light imaging architecture: Top-view (vertical), upper-lateral (45° oblique), and lower-lateral (−45° oblique) camera units synchronized with turntable rotation for complete 360° surface coverage.
- AI-accelerated 3D reconstruction pipeline: Real-time mesh generation using structure-from-motion (SfM) and multi-view stereo (MVS) algorithms; output includes watertight polygonal meshes with vertex-level color mapping (RGB + CIELAB).
- Automated adaptive imaging protocol: System autonomously evaluates plant dimensions via preliminary scan, selects optimal focal plane and magnification, and performs dynamic image stitching for specimens exceeding field-of-view limits.
- Integrated environmental telemetry: Onboard temperature (±0.3°C) and relative humidity (±2% RH) sensors log metadata with each acquisition; data embedded in EXIF headers and exported CSV reports.
- Dual-mode operation: Concurrent 2D high-throughput analysis (≤60 s/sample) and background 3D model generation—enabling uninterrupted batch processing without workflow interruption.
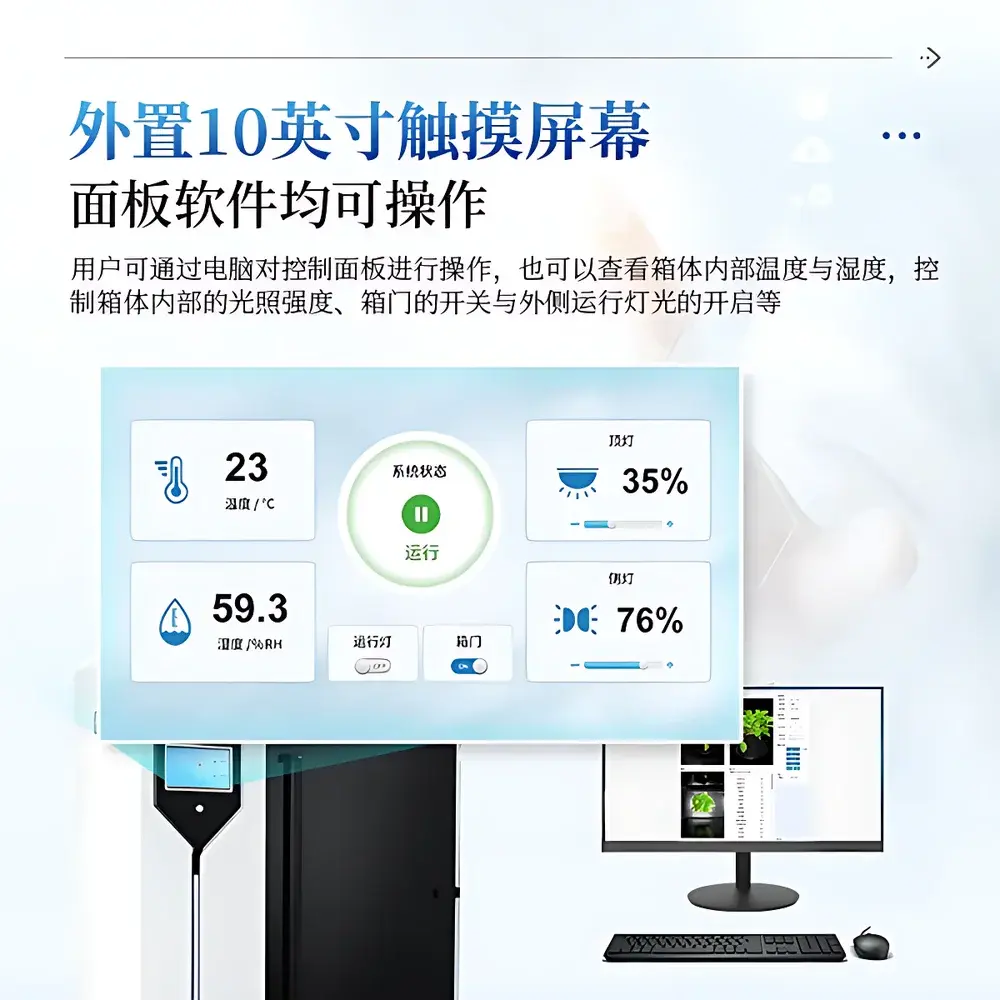
- Full hardware controllability via 10-inch industrial touchscreen HMI or remote Windows client: Includes motorized Z-axis lens positioning (±1 mm precision), LED intensity modulation (0–100% PWM), chamber door interlock management, and real-time weight synchronization (0.1 g resolution).
- Cloud-connected architecture: Encrypted HTTPS-based upload of processed datasets (including 3D model binaries, trait matrices, and time-series metadata) to user-configurable AWS S3 or private cloud endpoints.
Sample Compatibility & Compliance
The IN-Pheno50 accommodates intact, living plants within height (300–1000 mm), width (100–500 mm), and fresh mass (100–75,000 g) operational envelopes—covering vegetative and early reproductive stages of Poaceae (e.g., rice, wheat), Solanaceae (e.g., tomato, tobacco), Brassicaceae (e.g., Arabidopsis, canola), and Fabaceae (e.g., soybean, pea). All optical calibrations adhere to ISO 17025 traceability requirements via NIST-traceable reference targets. Software complies with FDA 21 CFR Part 11 for electronic records and signatures, including audit trails for user actions, parameter modifications, and result exports. Data structures conform to MIAPPE v1.1 (Minimum Information About a Plant Phenotyping Experiment) metadata schema, facilitating interoperability with BreedBase, Gramene, and ELIXIR Plant Sciences resources.
Software & Data Management
The IN-Pheno50 Control & Analysis Suite (v4.2+) runs natively on Windows 10/11 (64-bit) with NVIDIA GPU acceleration (minimum GeForce RTX 3060, 8 GB VRAM). It provides dual-language UI (English/Chinese), configurable export paths for raw images, trait tables (CSV/Excel), annotated 3D models (OBJ/GLB), and time-lapse video exports (MP4/H.265). All analytical outputs include uncertainty propagation estimates derived from pixel-level noise modeling and geometric reprojection error metrics. Exported datasets embed ISO 8601 timestamps, instrument serial numbers, environmental context tags, and version-stamped algorithm identifiers—ensuring full provenance for peer-reviewed publication or regulatory submission.
Applications
- Quantitative genetics: High-resolution QTL mapping of architectural traits (e.g., canopy volume, branching angle distribution, fractal dimension) under abiotic stress gradients (drought, salinity, elevated CO₂).
- Breeding program support: Early-stage selection for ideotype traits—including compactness index, leaf area index (LAI) proxies, and senescence progression—across F₂, RIL, or DH populations.
- Ecophysiology research: Correlation of spectral texture features (e.g., ASM, SSIM, entropy) with photosynthetic efficiency parameters measured concurrently via PAM fluorometry or gas exchange systems.
- Phytosanitary monitoring: Detection of subtle biotic stress signatures (e.g., chlorosis heterogeneity, lesion edge irregularity) prior to symptom visibility under standard lighting.
- Controlled-environment phenomics: Integration with automated watering/nutrient delivery systems for closed-loop phenotyping experiments compliant with OECD GLP Annex III guidelines.
FAQ
What is the minimum and maximum plant size supported by the IN-Pheno50?
Plants must be between 300 mm and 1000 mm in height and 100 mm to 500 mm in maximum lateral span. Specimens exceeding these dimensions require manual segmentation or external mounting fixtures.
Does the system support calibration against international color standards?
Yes—CIE D65 illuminant and 2° standard observer are embedded in all Lab-space calculations; optional NIST SRM 2039 color checker validation kits are available for routine verification.
Can 3D models be exported for use in third-party simulation software?
OBJ and GLB formats are natively supported, including per-vertex RGB and normal vector data; models retain metric scale (mm) and coordinate origin alignment with the turntable center.
Is offline operation possible without cloud connectivity?
All core imaging, reconstruction, and analysis functions operate fully offline; cloud upload is an optional post-processing step requiring no internet access during acquisition.
What computing resources are required to run the analysis suite at full capacity?
A dedicated workstation with Intel Core i5-11400 or higher, 16 GB RAM, NVIDIA RTX 3060 (8 GB VRAM), and ≥512 GB NVMe SSD is recommended for concurrent 2D/3D processing of >20 samples/hour.