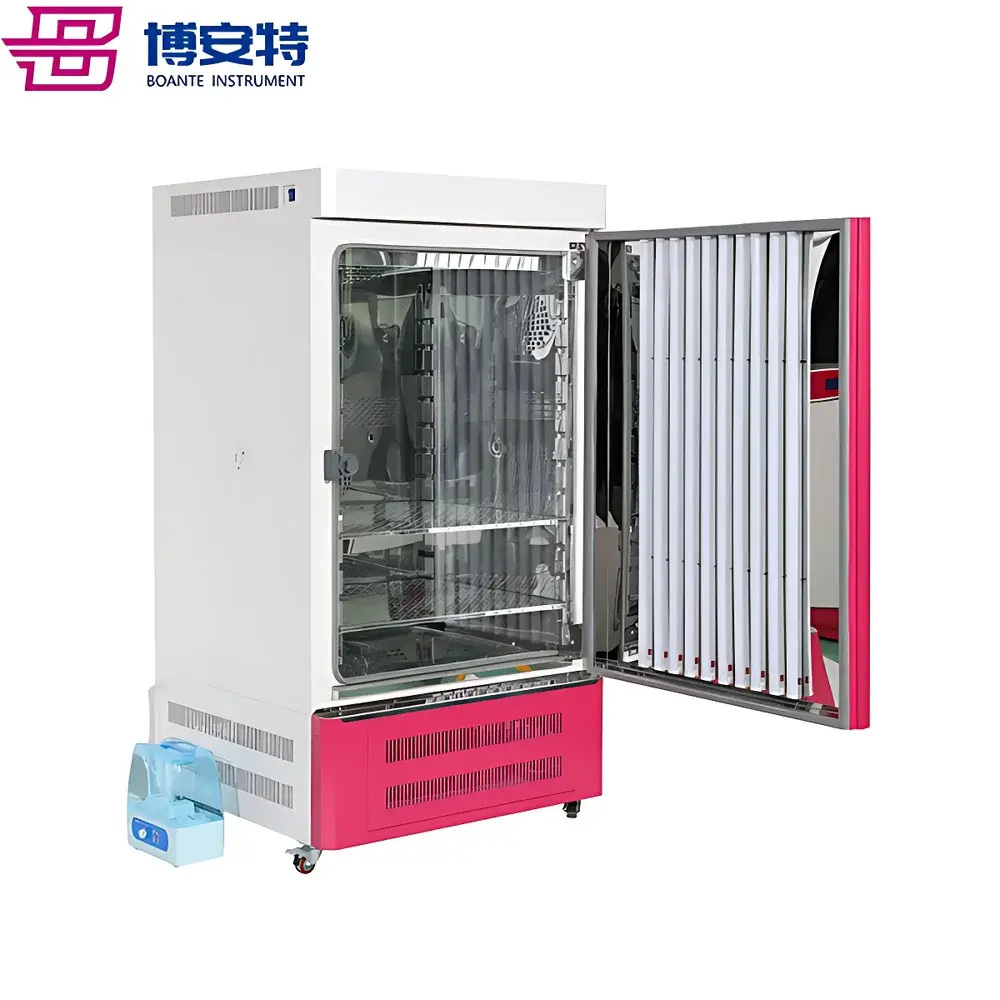
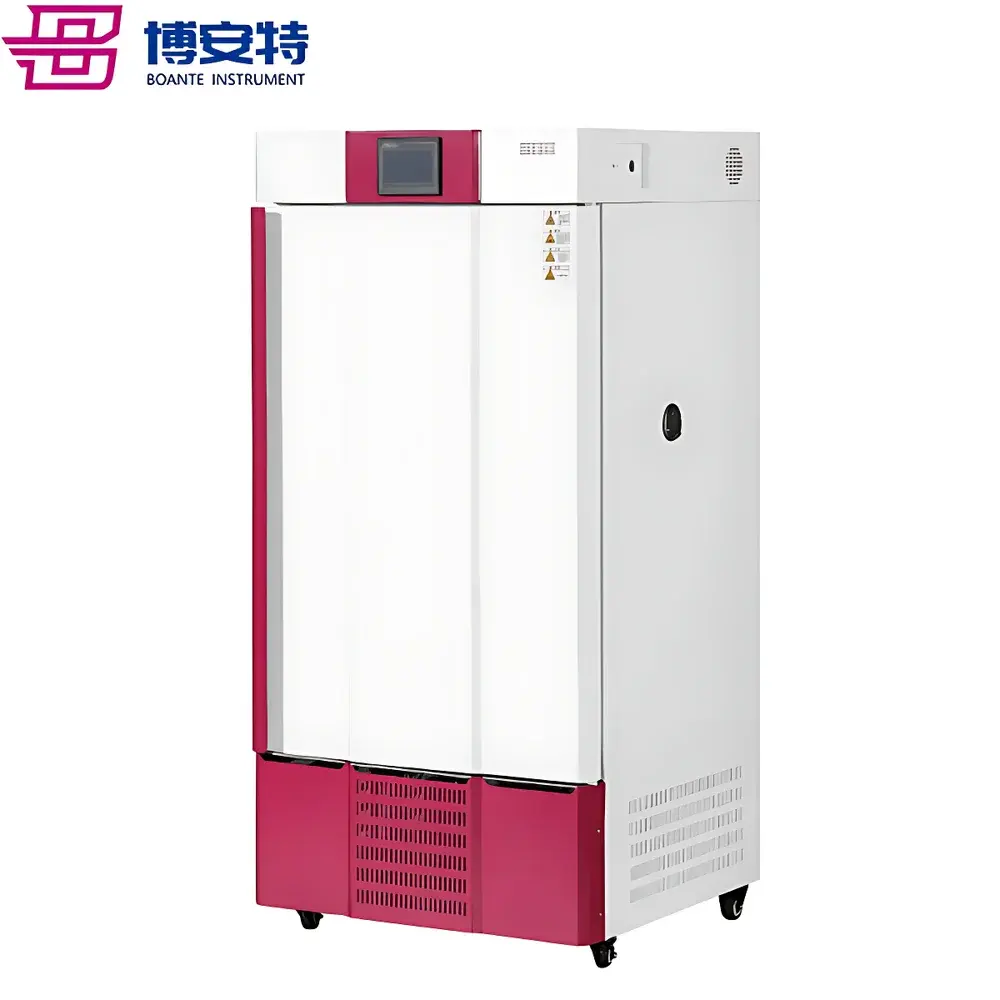
COMECAUSE IN-Pheno50 Plant Phenotyping Imaging System
| Brand | COMECAUSE |
|---|---|
| Origin | Shandong, China |
| Manufacturer Type | Direct Manufacturer |
| Model | IN-Pheno50 |
| Max Plant Height | 300–1000 mm |
| Max Plant Width | 100–500 mm |
| Max Fresh Weight | 100–75,000 g |
| Enclosure Dimensions | 1400 × 830 × 2140 mm (L×W×H) |
| Camera Resolution | Up to 12 MP |
| Imaging Modalities | Top, Side-Above, Side-Below Visible-Light Multi-View Acquisition |
| 3D Reconstruction Method | AI-Enhanced Multi-View Stereo (MVS) with Dynamic Pose Estimation |
| Environmental Sensors | Integrated Temperature & Relative Humidity Monitoring |
| Control Interface | 10-inch External Touchscreen + PC Software (Windows 10/11, Intel i5+, 8 GB RAM, NVIDIA GPU ≥ 8 GB VRAM) |
| Data Output | 2D/3D Morphological, Colorimetric (RGB, CIELab D65/2°, HSV, Grayscale), and Textural Parameters (GLCM-based: ASM, Contrast, Correlation, Entropy, Homogeneity, etc.), SSIM, Skeleton Metrics, Convex Hull & Projection Area Ratios |
Overview
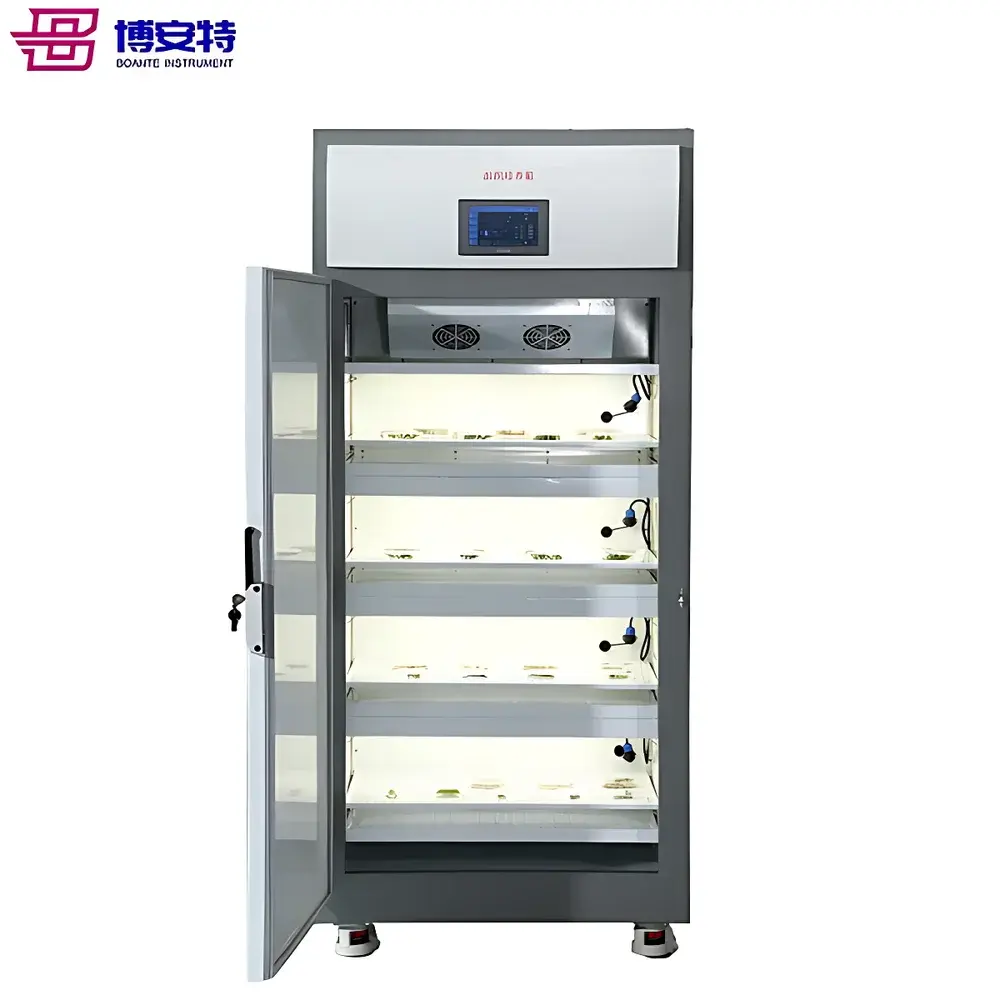
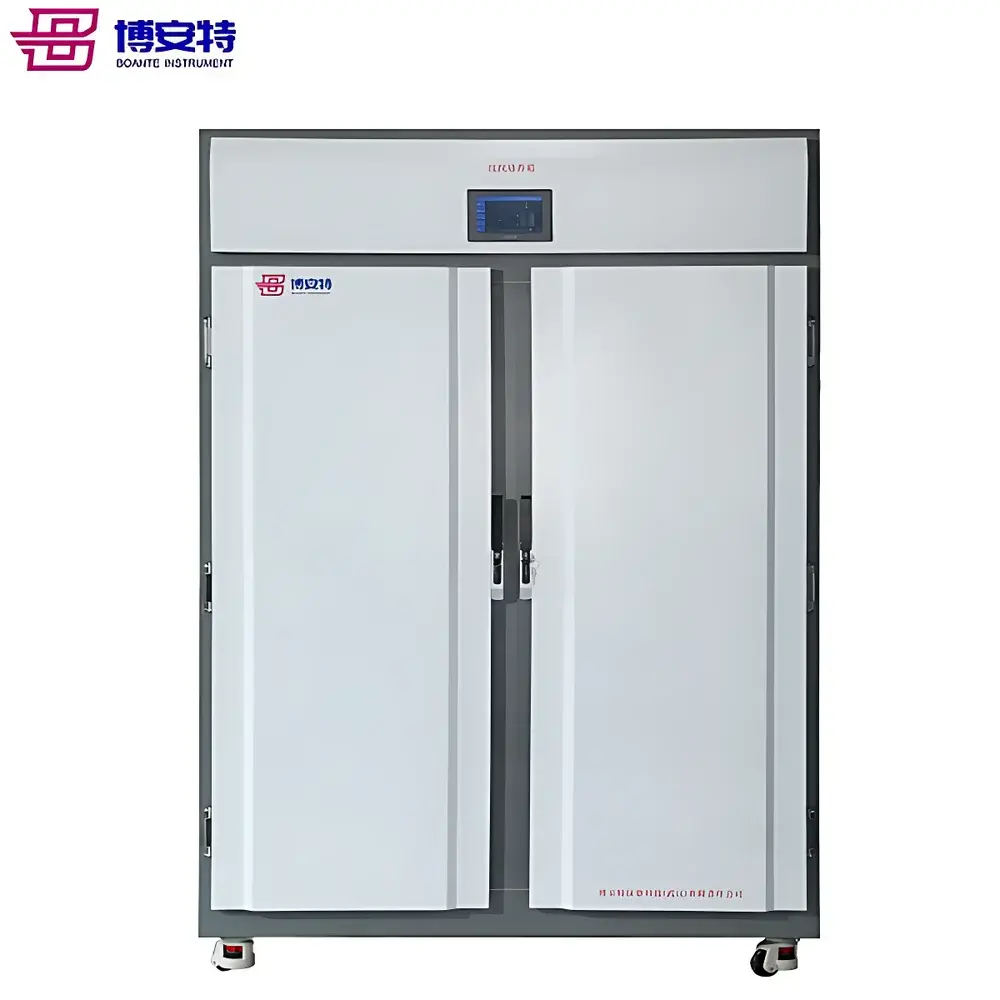
The COMECAUSE IN-Pheno50 Plant Phenotyping Imaging System is a modular, high-throughput phenotyping platform engineered for non-destructive, quantitative analysis of plant morphology, color, and surface texture in controlled environments. It operates on the principle of multi-view visible-light imaging combined with computer vision–driven 3D reconstruction. Three synchronized optical units—mounted at top, side-above, and side-below positions—acquire calibrated images as the specimen rotates on a motorized turntable. This geometry enables full circumferential coverage without occlusion, supporting robust 3D point cloud generation via structure-from-motion (SfM) and multi-view stereo (MVS) algorithms enhanced by deep learning–based depth refinement. Unlike single-angle or fixed-position systems, the IN-Pheno50 captures spatially registered data across orthogonal planes, permitting true volumetric quantification—including canopy volume, branch angle distribution in 3D space, and stem-to-leaf orientation metrics—not inferable from 2D projections alone. The system is designed for reproducible acquisition under standardized illumination (D65-spectrum white LED arrays with adjustable intensity), enabling longitudinal studies compliant with FAIR (Findable, Accessible, Interoperable, Reusable) data principles.
Key Features
- Automated adaptive imaging: Real-time size estimation triggers dynamic camera positioning (vertical lift/lower, horizontal translation) and auto-stitching for specimens exceeding field-of-view limits.
- Integrated environmental monitoring: Onboard temperature and relative humidity sensors log chamber conditions synchronously with image capture, supporting environmental covariate correlation in statistical modeling.
- Dual-mode analysis pipeline: Concurrent 2D high-throughput processing (≤60 s/sample) and background-queued 3D reconstruction ensure uninterrupted throughput—no workflow blocking during model generation.
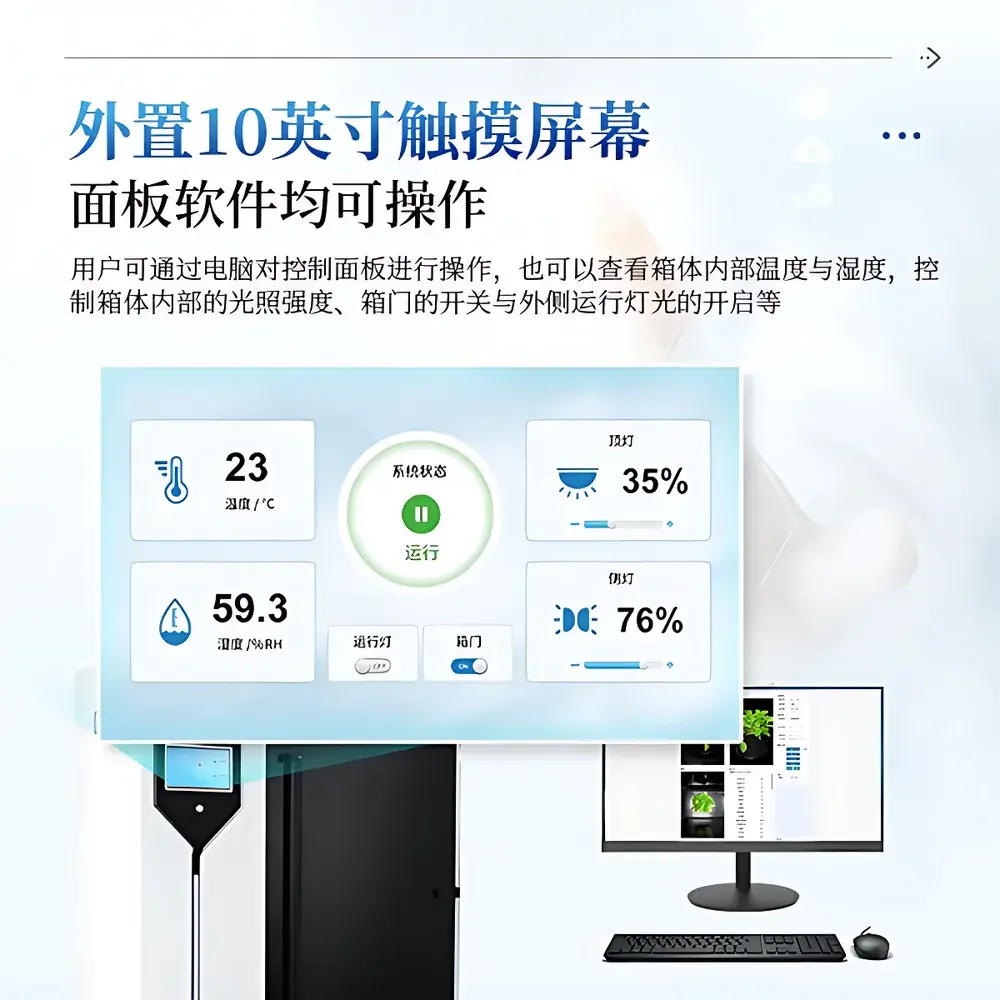
- Full hardware control via dual interface: Local 10-inch touchscreen for rapid parameter adjustment (light intensity, door actuation, ambient lighting) and remote PC software for advanced protocol scripting, batch scheduling, and calibration management.
- Multi-scale resolution flexibility: Configurable camera resolution (up to 12 megapixels) balances detail fidelity against storage bandwidth and processing latency per experimental requirement.
- Weight-integrated turntable: Precision load cell embedded in rotation stage automatically records fresh biomass upon placement and time-synchronizes mass data with imaging timestamps.
- Movable enclosure design: Self-contained, caster-mounted chassis allows repositioning within growth rooms or climate-controlled facilities without structural modification or permanent installation.
- Hardware safety architecture: Redundant limit switches, proximity sensors, and mechanical end-stops prevent overtravel, collision, or operator exposure during automated motion sequences.
Sample Compatibility & Compliance
The IN-Pheno50 accommodates intact, living plants across major botanical families including Poaceae (e.g., rice, wheat, maize), Solanaceae (tomato, tobacco), Brassicaceae (Arabidopsis, canola), and Fabaceae (soybean, pea). Specimen dimensions are supported within 300–1000 mm height and 100–500 mm width; fresh weight range spans 100 g to 75 kg—covering seedlings through mature vegetative or early reproductive stages. All optical components comply with IEC 62471 (Photobiological Safety) for LED lighting systems. Data acquisition workflows support GLP-aligned metadata tagging (operator ID, timestamp, chamber ID, protocol version), while encrypted cloud upload (via device-unique serial binding) ensures traceability. Though not FDA-cleared, the system’s audit-trail logging, user-access controls, and exportable raw image stacks align with ISO/IEC 17025 documentation requirements for accredited testing laboratories.
Software & Data Management
The proprietary IN-Pheno Suite (v3.x) provides unified control, analysis, and archival. It supports real-time preview, manual/auto focus selection, optical zoom control, and live environmental telemetry overlay. Analysis modules compute over 120 standardized parameters: morphological (plant height, width ratio, fractal dimension, skeleton length, convex hull area/perimeter), colorimetric (CIELab D65/2°, HSV, grayscale histogram moments), and textural (Gray-Level Co-occurrence Matrix derivatives: ASM, contrast, entropy, homogeneity, dissimilarity, and SSIM). All results are stored in structured CSV/JSON format with embedded EXIF-like metadata. Users may export individual reports, bulk datasets, or annotated 3D model videos (MP4/H.264). Path configurations for analysis output, 3D model storage, and video archives are editable per project. The interface supports seamless language switching (English/Chinese) and scales dynamically to native display resolution. Cloud synchronization uses TLS 1.2 encryption and retains versioned backups for up to 90 days.
Applications
The IN-Pheno50 serves core research and breeding applications requiring objective, longitudinal phenotypic characterization. In plant physiology, it quantifies drought-induced changes in leaf angle distribution, canopy closure dynamics, and senescence-associated color shifts (a*, b* trajectories). In environmental science, it correlates spectral texture metrics (e.g., GLCM entropy decline) with ozone exposure or nutrient deficiency severity. For quantitative genetics, high-resolution 3D branching architecture data feed QTL mapping of inflorescence complexity traits in cereals. In plant pathology, SSIM-based anomaly detection identifies early-stage biotic stress before visible symptom onset. Its portability and modularity make it suitable for deployment in field-deployable phenotyping trailers or greenhouse mobile units—enabling comparative trials across growth chambers with identical measurement geometry and calibration traceability.
FAQ
What operating system and hardware specifications are required to run the IN-Pheno Suite?
Windows 10 or 11 (64-bit), Intel Core i5 or higher, minimum 8 GB RAM, dedicated NVIDIA GPU with ≥8 GB VRAM, and 500 GB SSD storage for cache and temporary model generation.
Does the system support integration with external environmental control systems (e.g., HVAC or irrigation controllers)?
Yes—via TCP/IP or Modbus RTU protocols using optional API extension module; enables closed-loop experiments where phenotypic feedback triggers environmental setpoint adjustments.
Can the 3D models be exported in standard formats for third-party analysis (e.g., MeshLab, Blender, or R packages like rgl)?
Yes—models export as OBJ with embedded MTL textures and PLY point clouds, both compatible with open-source and commercial 3D analysis toolchains.
Is calibration verification traceable to national standards?
Geometric calibration uses NIST-traceable checkerboard targets; color calibration employs X-Rite ColorChecker Passport targets with D65 illumination validation; all calibration certificates are stored in the software audit log.
How is data security managed during cloud transmission and storage?
All data is encrypted in transit (AES-256/TLS 1.2) and at rest (AES-256); cloud accounts enforce two-factor authentication, role-based access control, and automatic session timeout after 15 minutes of inactivity.