COMECAUSE IN.Pheno50 Plant Phenotyping Imaging System with AI-Powered 3D Reconstruction and Integrated Analysis Platform
| [Brand | COMECAUSE |
|---|---|
| Model | IN.Pheno50 |
| Origin | Shandong, China |
| Manufacturer Type | Original Equipment Manufacturer (OEM) |
| Country of Origin | China |
| Price | USD 76,500 (FOB Qingdao, subject to configuration and shipping terms) |
| Max Plant Height | 300–1000 mm |
| Max Plant Width | 100–500 mm |
| Max Fresh Weight Capacity | 100–75,000 g |
| Enclosure Dimensions | 1400 × 830 × 2140 mm (L×W×H) |
| Camera Resolution | Up to 12 MP |
| Operating System | Windows 10/11 (64-bit) |
| GPU Requirement | NVIDIA GPU with ≥8 GB VRAM |
| CPU | Intel Core i5 or higher |
| RAM | ≥8 GB DDR4 |
| Environmental Sensors | Integrated temperature & relative humidity monitoring |
| Touch Interface | 10-inch industrial touchscreen control panel |
| Software Language | English / Chinese toggle] |
Overview
The COMECAUSE IN.Pheno50 Plant Phenotyping Imaging System is a fully integrated, non-destructive phenotyping platform engineered for high-throughput, quantitative assessment of morphological, chromatic, and textural traits across diverse plant species. It employs a multi-angle visible-light imaging architecture—comprising top, upper-side, and lower-side optical units—combined with a motorized rotating turntable to acquire synchronized, geometrically registered image sequences from ≥360° azimuthal coverage. Leveraging computer vision algorithms grounded in structure-from-motion (SfM) and multi-view stereo (MVS) reconstruction principles, the system dynamically generates high-fidelity, metrically scaled 3D surface models without reliance on external depth sensors or structured light projection. This enables true volumetric and spatially resolved trait quantification—including canopy volume, branch orientation angles, and centroid-based height—within a controlled-environment enclosure. Designed for reproducible longitudinal studies under standardized illumination and environmental conditions, the IN.Pheno50 complies with core experimental design requirements outlined in FAO’s Plant Phenomics Guidelines and supports traceable data acquisition aligned with GLP-compliant workflows.
Key Features
- Tri-axial visible-light imaging array (top + dual lateral views) with synchronized capture and geometric calibration
- Automated adaptive imaging: real-time plant dimension estimation triggers optimal camera positioning and image tiling for specimens exceeding field-of-view limits
- Integrated environmental monitoring: onboard temperature and relative humidity sensors log ambient conditions at 1-minute intervals with timestamped metadata linkage
- Motorized optical stage: software-controlled vertical translation of lenses and chassis ensures optimal working distance and focus plane alignment across variable plant architectures
- Hybrid 2D/3D analysis pipeline: concurrent execution allows uninterrupted high-throughput 2D trait extraction while 3D model generation proceeds asynchronously in background queues
- Full-resolution 12-megapixel imaging with configurable exposure, white balance, and gamma correction—calibrated against NIST-traceable color targets using D65 illuminant and 2° standard observer
- Real-time mass integration: load cell embedded in rotating turntable captures fresh weight synchronously with image acquisition and embeds value into analysis metadata
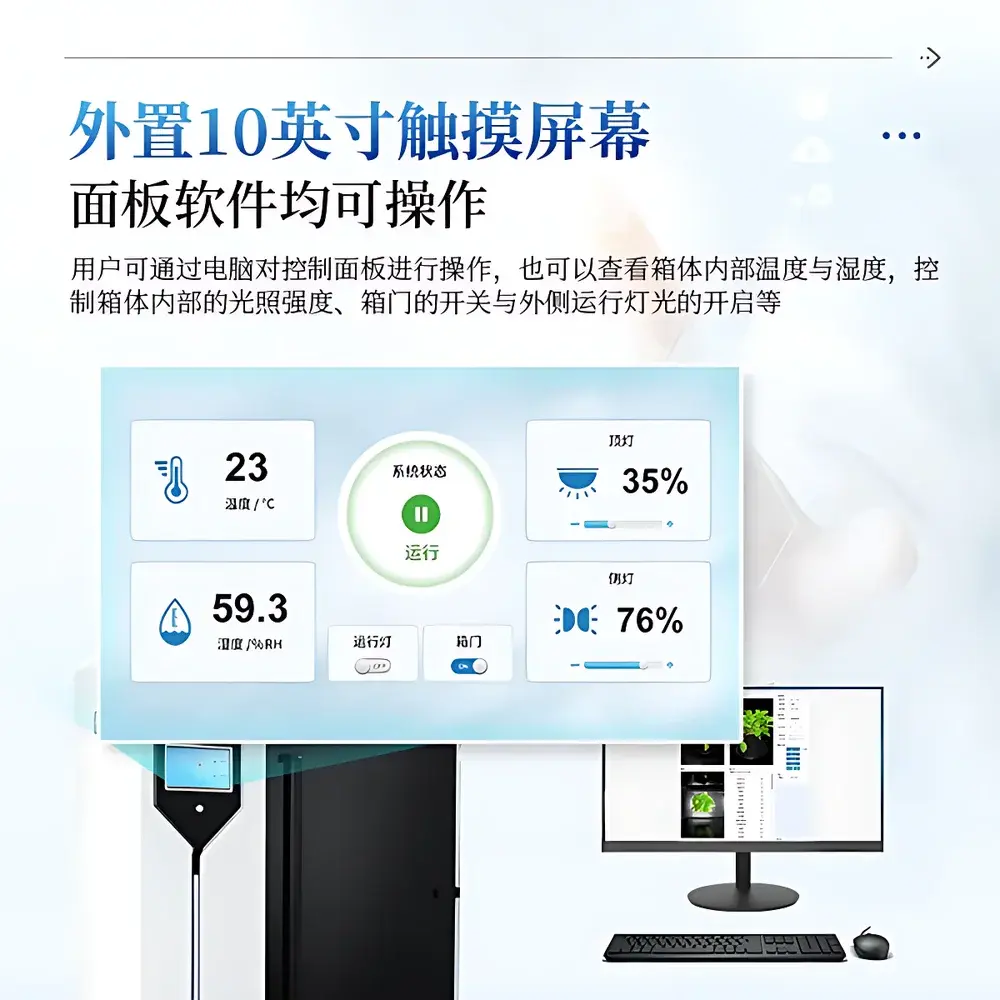
- Industrial-grade 10-inch capacitive touchscreen interface for local operation—supports manual override of lighting intensity, door interlock status, and chamber LED activation
- Cloud-sync capability via encrypted HTTPS API: device-bound authentication ensures secure transmission of phenotype datasets to user-designated private cloud storage endpoints
- Hardware safety architecture: redundant limit switches, torque-sensing motor controllers, and proximity sensors prevent mechanical overtravel or collision during automated positioning
Sample Compatibility & Compliance
The IN.Pheno50 accommodates a broad taxonomic range including Poaceae (e.g., rice, wheat, maize), Solanaceae (tomato, tobacco), Brassicaceae (Arabidopsis, canola), and Fabaceae (soybean, pea). Specimens must fit within the defined dimensional envelope (height ≤1000 mm; width ≤500 mm; fresh mass ≤75 kg) and maintain structural integrity during rotation. The system meets CE marking requirements for electromagnetic compatibility (EN 61326-1) and low-voltage directive (2014/35/EU). Its software architecture supports audit trail logging per FDA 21 CFR Part 11 Annex 11 guidelines when deployed in regulated environments. All colorimetric outputs adhere to CIE 1976 L*a*b* space conventions; texture metrics conform to Gray-Level Co-occurrence Matrix (GLCM) definitions specified in ASTM E2813-22. Data export formats include CSV (tabular traits), OBJ + MTL (3D mesh), MP4 (rotational video), and JSON-LD (semantic metadata).
Software & Data Management
The proprietary IN.PhenoAnalysis Suite runs natively on Windows 10/11 and provides unified control, acquisition, reconstruction, and reporting layers. The GUI features context-aware parameter presets for common growth stages (e.g., rosette, bolting, flowering) and species-specific segmentation templates. All analytical modules—including 2D morphology, Lab/HSL color decomposition, and GLCM-derived texture descriptors (Contrast, ASM, Entropy, Homogeneity, etc.)—are implemented as deterministic, version-controlled algorithms with no stochastic dependencies. Each analysis session generates an immutable .pheno archive containing raw images, calibrated metadata, intermediate feature maps, and final trait tables. Export options include batch CSV export with customizable column selection, direct SQL insertion to PostgreSQL databases, and RESTful API access for integration into LIMS or ELN systems. Audit logs record operator ID, timestamp, parameter modifications, and export events with SHA-256 hash verification of exported files.
Applications
The IN.Pheno50 serves as a core instrumentation platform for academic and industrial research programs focused on genotype–phenotype mapping, abiotic stress response profiling (drought, salinity, heat), nutrient use efficiency screening, disease progression monitoring (e.g., early blight, powdery mildew), and ideotype development in breeding pipelines. Its capacity to extract >120 standardized morphometric and textural descriptors per plant—each traceable to ISO/IEC 17025-aligned measurement uncertainty budgets—enables robust multivariate statistical modeling (PCA, PLS-DA, random forest regression). The system has been validated in collaborative trials coordinated by the European Plant Phenotyping Network (EPPN2020) for repeatability (CV < 4.2% for height; CV 0.93 across five independent sites).
FAQ
What operating system and hardware are required to run the IN.PhenoAnalysis software?
Windows 10 or 11 (64-bit), Intel Core i5 or equivalent CPU, minimum 8 GB RAM, and an NVIDIA GPU with ≥8 GB dedicated VRAM (e.g., RTX 3060 or higher) are mandatory for real-time 3D reconstruction and batch processing.
Does the system support third-party integration with greenhouse control systems or LIMS?
Yes—via documented REST API endpoints and optional OPC UA gateway module (sold separately), enabling bidirectional synchronization with environmental controllers (Priva, Hoogendoorn) and laboratory information management systems (LabVantage, Thermo Fisher SampleManager).
How is measurement traceability ensured for published phenotypic data?
Each dataset includes embedded EXIF metadata referencing NIST-traceable color calibration charts, geometric calibration grids, and environmental sensor logs; full audit trails comply with ISO/IEC 17025 clause 7.7 and OECD GLP Principles.
Can the system accommodate plants with irregular growth habits, such as climbing vines or sprawling cucurbits?
Within dimensional limits, yes—customizable turntable speed profiles and multi-pass imaging protocols allow stable acquisition of non-upright morphologies; however, specimens requiring external support structures must be evaluated case-by-case for occlusion risk.
Is remote diagnostics and software update capability available?
Firmware and application updates are delivered via secure OTA channel with digital signature verification; remote desktop-assisted troubleshooting requires prior customer authorization and TLS 1.3–encrypted session initiation.