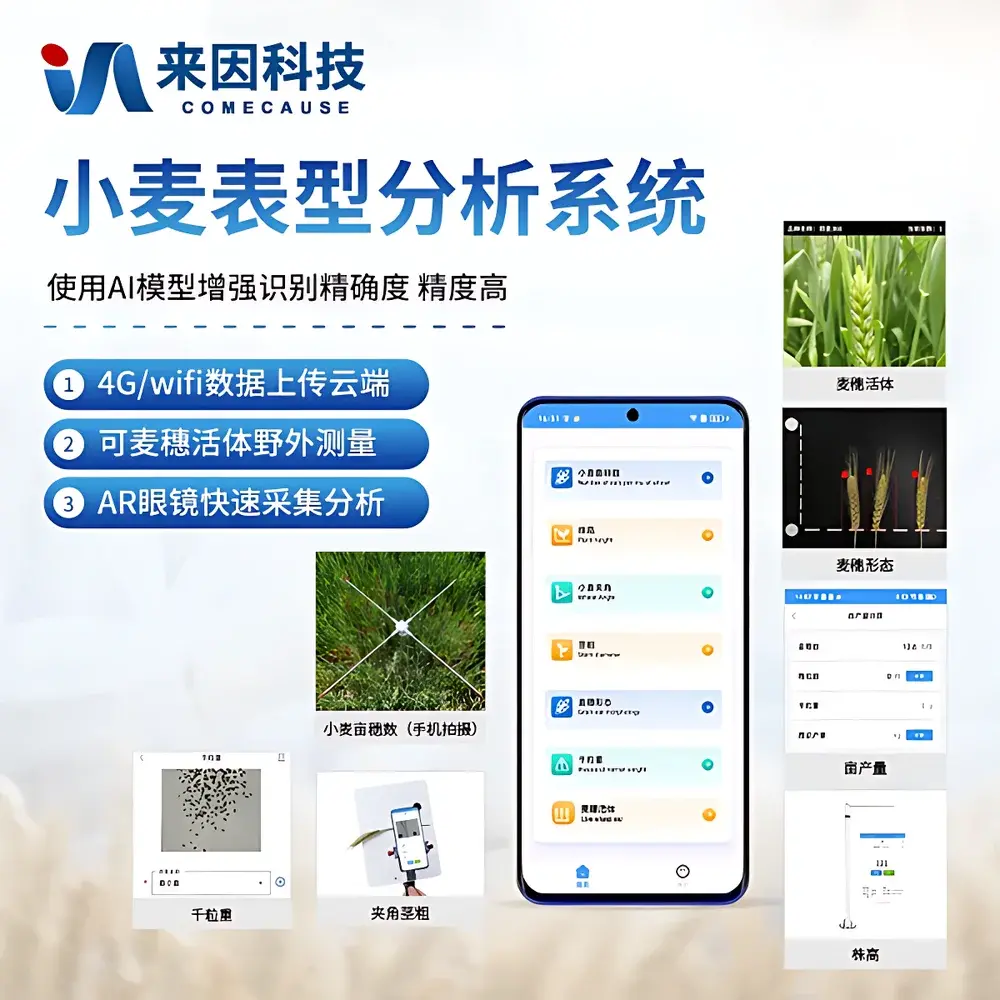
COMECAUSE IN-XM04 Wheat Phenotyping System
| Brand | COMECAUSE |
|---|---|
| Origin | Shandong, China |
| Manufacturer Type | Direct Manufacturer |
| Country of Origin | China |
| Model | IN-XM04 |
| Price | USD 4,200 (FOB) |
Overview
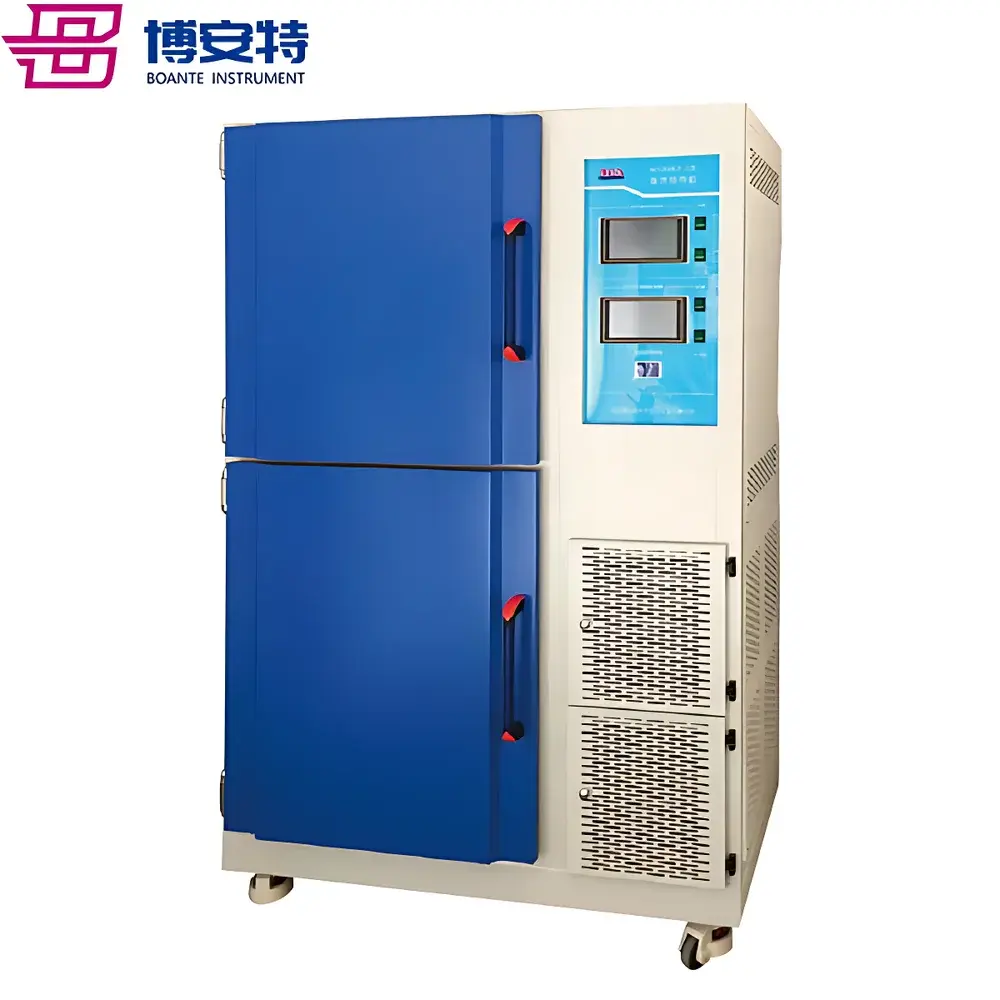
The COMECAUSE IN-XM04 Wheat Phenotyping System is a field-deployable, modular imaging-based phenotyping platform engineered for high-throughput, non-destructive quantification of key wheat morphological traits. It operates on the principle of computer vision–guided geometric and photometric analysis, integrating dual high-resolution cameras (50 MP + 12 MP), adaptive AI inference engines, and calibrated optical geometry to extract biologically meaningful metrics from raw imagery. Unlike conventional manual or destructive sampling methods, the IN-XM04 enables in situ, real-time acquisition of structural phenotypes—including tiller density, spike architecture, leaf–stem angle, stem diameter, plant height, and thousand-kernel weight—across multiple developmental stages (from seedling to physiological maturity). Its design adheres to FAO-recommended wheat growth staging protocols (Zadoks scale), ensuring temporal alignment with agronomic decision windows. The system supports both live-field measurement and post-harvest lab-based analysis, delivering traceable, audit-ready data suitable for QTL mapping, genomic selection pipelines, and regulatory-compliant variety trials.
Key Features
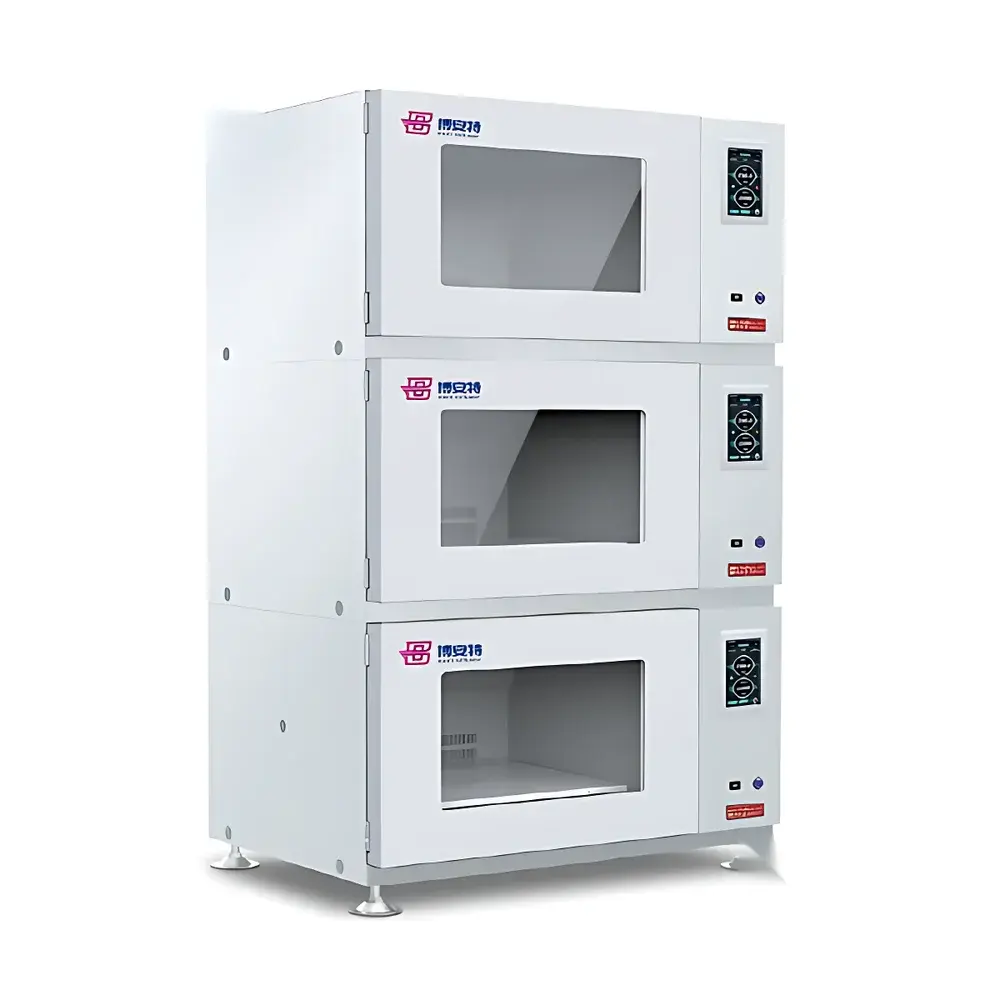
- Modular four-in-one architecture: independent yet interoperable modules for panicle count per unit area, spike morphology, leaf–stem angle & stem diameter, and plant height—each optimized for stage-specific deployment.
- Dual-sensor imaging platform: 50 MP main camera for macro-scale field sampling (1 m² frame), 12 MP auxiliary camera for close-range spike and stem detail capture; both equipped with auto white balance, distortion correction, and tilt-compensated perspective rectification algorithms.
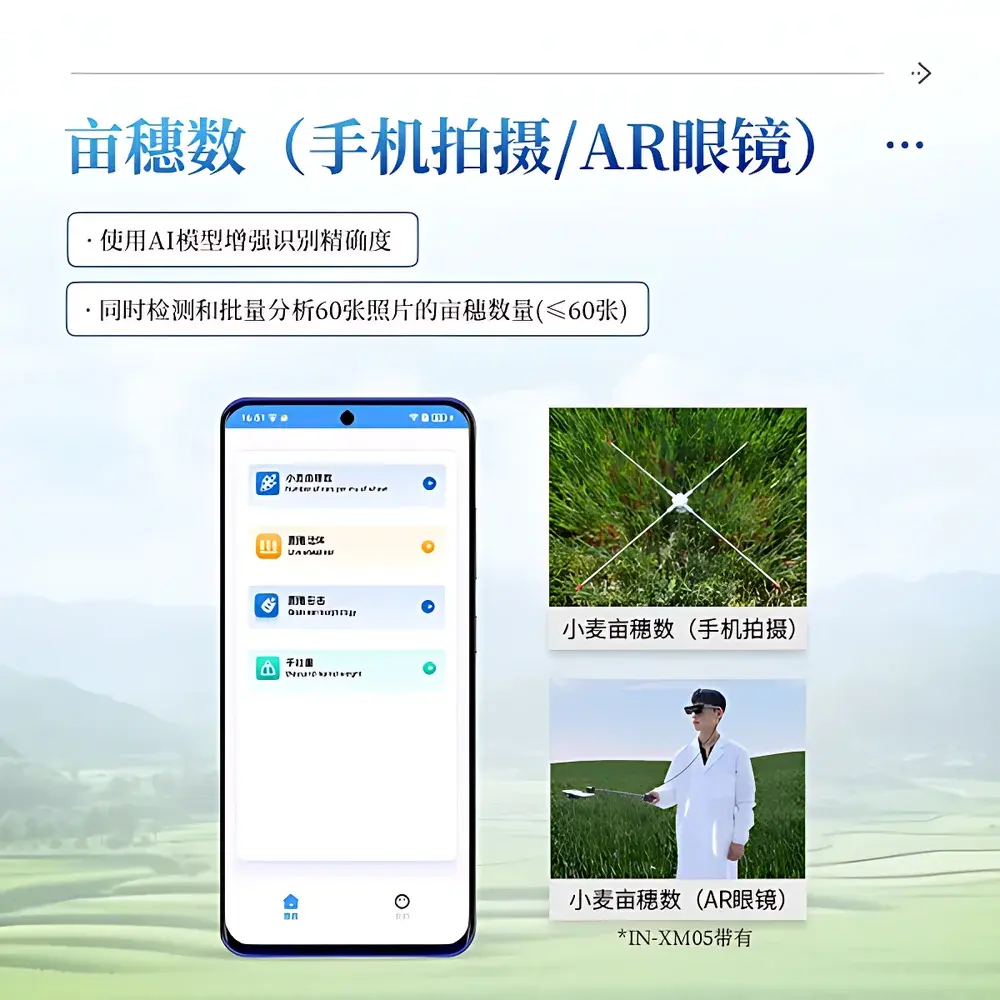
- AI-accelerated trait extraction: proprietary deep learning models trained on >200,000 annotated wheat images across 17 cultivars and 5 growing regions; models support batch processing of up to 60 images per session with ≤3% relative error in panicle counting.
- Field-rugged hardware: aluminum alloy chassis, stainless-steel support structures, matte-black acrylic reference backboards, and sealed optical path design for consistent performance under variable solar irradiance, wind, and ambient humidity.
- Android-based embedded OS: 10.1″ capacitive touchscreen interface with offline operation capability; local storage (256 GB SSD for imaging modules, 128 GB for height module); GPS geotagging and timestamping for every measurement.
- Bilingual UI & cloud integration: seamless English/Chinese language toggle; automatic data synchronization via Wi-Fi or 4G LTE to secure cloud dashboard supporting role-based access, version-controlled datasets, and API-driven export to R, Python, or breeding management software (e.g., BreedBase, FieldBook).
Sample Compatibility & Compliance
The IN-XM04 is validated for use with Triticum aestivum L. (bread wheat) and exhibits cross-species functionality for Oryza sativa (rice) and Brassica napus (oilseed rape) in leaf angle and stem diameter measurements. All modules comply with ISO 21748:2017 (Guidance for the evaluation of measurement uncertainty) and follow metrological traceability principles outlined in VIM 3rd edition. Data logging meets GLP-aligned requirements: full audit trail (user ID, timestamp, location, image hash, algorithm version), electronic signature support, and immutable metadata embedding. While not FDA 21 CFR Part 11 certified out-of-box, the cloud infrastructure supports configurable validation packages for GxP environments upon client request. Measurement protocols align with CIMMYT wheat phenotyping guidelines and national standards GB/T 3543.7–2018 (Crop variety testing—Phenotypic trait measurement).
Software & Data Management
The system runs COMECAUSE PhenotypeSuite v3.2—a modular Android application suite with embedded TensorFlow Lite inference runtime and SQLite3 local database. Each module generates structured JSON reports containing raw pixel coordinates, derived trait values, confidence scores, and calibration metadata. Export options include CSV, Excel (.xlsx), and standardized PlantCV-compatible JSON-LD. Cloud backend provides RESTful APIs for integration with LIMS, ELN, or enterprise breeding databases. Audit logs record all user actions, parameter changes, and firmware updates. Data retention policies are configurable (default: 5 years), with optional encrypted backup to AWS S3 or on-premise NAS. Software updates are delivered over-the-air (OTA) with SHA-256 integrity verification.
Applications
- Pre-breeding and varietal trialing: rapid screening of canopy architecture traits (e.g., erect leaf angle for improved light interception) across segregating populations.
- Yield component analysis: coupling panicle density, spikelet count, grain count per spike, and thousand-kernel weight to model theoretical yield potential under defined sowing densities.
- Functional genomics: quantitative correlation of morphological traits with SNP markers in GWAS or QTL studies—particularly for genes regulating tillering (e.g., TB1), spike development (e.g., VRN1), or stem mechanical strength (e.g., TaCER1).
- Climate resilience assessment: longitudinal monitoring of phenotypic plasticity (e.g., stem diameter reduction under drought stress, panicle abortion rate under heat shock) across multi-environment trials.
- Extension & precision agriculture: generating field-level trait maps for variable-rate input recommendations (e.g., nitrogen top-dressing based on tiller density gradients).
FAQ
What growth stages are supported for each trait measurement?
Panicle count: Zadoks 55–87 (heading to late milk stage); spike morphology: post-harvest lab analysis only (Zadoks 92+); leaf–stem angle & stem diameter: Zadoks 31–77 (jointing to soft dough); plant height: all stages from Zadoks 11 (seedling emergence) to 92 (harvest ripe).
Is calibration required before each field session?
Yes—each module includes a physical calibration standard (e.g., foldable metric grid for panicle module, precision goniometer for angle module). Auto-calibration routines verify optical alignment and lighting uniformity prior to image capture.
Can the system operate without network connectivity?
Yes—all core imaging, AI inference, and data storage functions operate fully offline. Cloud sync occurs only when connection is available and manually triggered or scheduled.
How is measurement uncertainty reported?
Each exported dataset includes expanded uncertainty (k=2) calculated per ISO/IEC Guide 98-3, incorporating repeatability (within-session CV%), reproducibility (inter-operator CV%), and calibration uncertainty components.
What training and documentation are provided?
Includes bilingual (EN/CN) hardware manual, SOPs aligned with ISO 20653:2022 (field phenotyping procedures), video tutorials, and remote onboarding sessions. Optional on-site technician certification available.