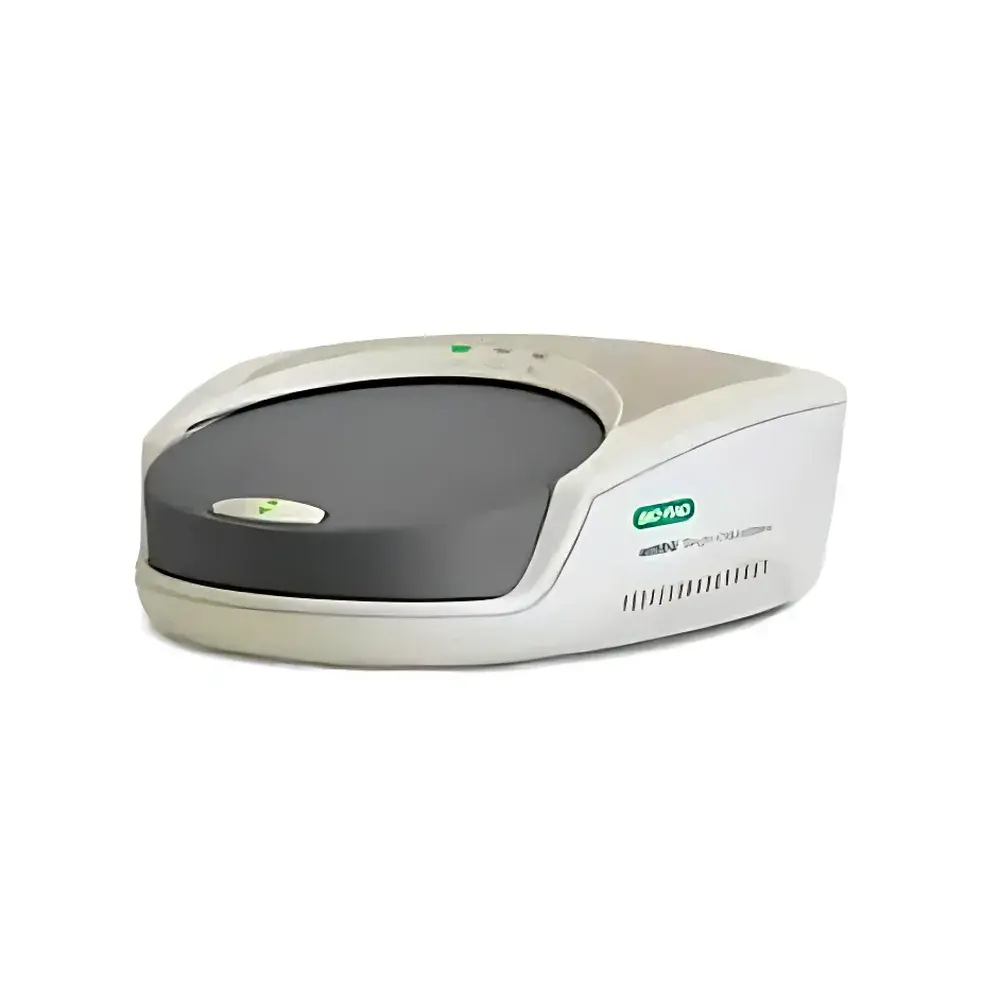
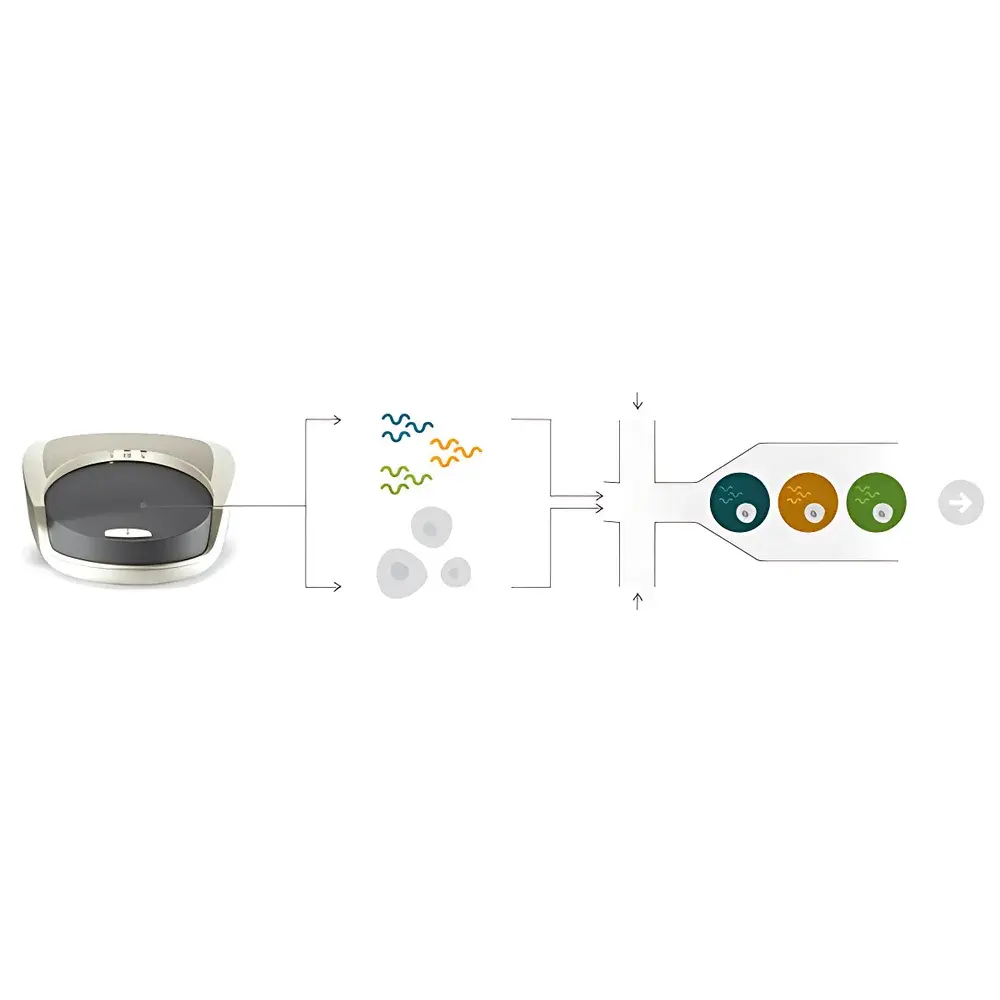
ddSEQ Single-Cell Droplet Generation System
| Brand | Bio-Rad |
|---|---|
| Origin | USA |
| Model | ddSEQ |
| Regulatory Classification | Import (CE-IVD compliant for research use only) |
| Intended Use | Automated microfluidic droplet generation for single-cell RNA sequencing library preparation |
Overview
The ddSEQ Single-Cell Droplet Generation System is a benchtop microfluidic platform engineered for reproducible, high-throughput encapsulation of individual cells into monodisperse water-in-oil droplets—enabling scalable 3′-end single-cell RNA sequencing (scRNA-seq). Leveraging deterministic droplet microfluidics—not stochastic partitioning—the system precisely co-encapsulates single cells with barcoded gel beads and lysis reagents in nanoliter-scale droplets. This architecture ensures uniform droplet size (CV < 5%), consistent cell lysis kinetics, and highly efficient reverse transcription within each compartment. Designed in collaboration with Illumina, the ddSEQ integrates seamlessly into the SureCell WTA 3′ workflow, delivering robust transcript capture across diverse mammalian cell types without bias toward cell size or viability. The system operates under ambient laboratory conditions and requires no specialized infrastructure, making it suitable for core facilities and translational research labs pursuing GLP-aligned scRNA-seq studies.
Key Features
- Automated droplet generation: Processes up to four samples simultaneously in ≤5 minutes per run, with minimal hands-on time and operator-independent reproducibility.
- Scalable throughput: Supports 200–10,000 single cells per sample; daily capacity exceeds 40,000 cells using standard operational protocols.
- Microfluidic precision: Integrated glass-chip cartridge ensures stable droplet formation (diameter: ~70 µm), optimized for >95% cell viability retention post-encapsulation and >90% cDNA conversion efficiency.
- No pre-amplification required: Compatible with Nextera-based library construction, eliminating bias introduced by PCR duplication and preserving native transcript abundance ratios.
- Indexing flexibility: Supports 24-sample multiplexing via dual-indexed adapters compliant with Illumina sequencing platforms (including NextSeq 550, NovaSeq 6000).
Sample Compatibility & Compliance
The ddSEQ system demonstrates validated performance across primary human and murine tissues—including PBMCs, dissociated tumor biopsies, neuronal cultures, and immortalized lines (HEK 293, NIH 3T3, A20, BJ). It exhibits no detectable size-dependent capture bias, as confirmed by orthogonal validation using TC20 Automated Cell Counter and median genes-per-cell metrics across heterogeneous populations. All reagents—including SureCell WTA 3′ Library Prep Kits—are manufactured under ISO 13485-certified quality systems. The ddSEQ platform and associated consumables are CE-marked for in vitro diagnostic (IVD) use in the European Union and cleared for research use only (RUO) in the United States under FDA guidelines. Data integrity meets ALCOA+ principles, supporting audit readiness for GxP environments when paired with BaseSpace Sequence Hub’s 21 CFR Part 11-compliant audit trail functionality.
Software & Data Management
Instrument operation is fully managed through the ddSEQ Control Software v3.x—a Windows-based application providing real-time pressure monitoring, chip priming diagnostics, and run log export (CSV/JSON). Raw droplet output is directly linked to Illumina’s BaseSpace Sequence Hub via secure HTTPS API integration. The BaseSpace Single-Cell Analysis App implements standardized preprocessing (cell barcode demultiplexing, UMI counting, gene-level quantification) using STARsolo and Seurat-compatible outputs. All intermediate and final datasets are stored encrypted-at-rest and in-transit, with role-based access control, versioned analysis history, and automated backup to AWS S3 buckets. Audit logs record user actions, parameter changes, and software updates—fully traceable for regulatory review.
Applications
- Transcriptomic profiling of rare immune subsets in tumor microenvironments
- Developmental trajectory inference during organoid differentiation
- Longitudinal monitoring of clonal evolution in hematologic malignancies
- Cross-species comparative analysis (e.g., human-mouse xenograft models)
- Quality control of iPSC-derived cell therapies via expression signature validation
FAQ
Is the ddSEQ system compatible with non-Illumina sequencing platforms?
Yes—while optimized for Illumina chemistry, FASTQ outputs are format-agnostic and can be processed by open-source pipelines such as CellRanger (10x Genomics), Alevin-fry, or kb-python for downstream analysis on any short-read sequencer.
What maintenance is required for the microfluidic cartridge?
Cartridges are single-use, sterilized, and pre-loaded with oil and surfactant. No cleaning or recalibration is needed between runs; disposal follows standard biohazard protocols.
Can frozen or fixed cells be processed?
The system is validated for fresh or cryopreserved viable cells (≥70% viability by trypan blue exclusion). Formalin-fixed samples are not supported due to compromised membrane integrity and RNA fragmentation.
How is batch effect minimized across multi-day experiments?
Each ddSEQ run includes internal technical replicates and spike-in controls (ERCC RNA standards); BaseSpace applies Combat-like normalization during cross-run integration, preserving biological variance while correcting for plate-to-plate drift.
Does the system support spatial transcriptomics integration?
While ddSEQ itself performs dissociative scRNA-seq, its gene expression matrices serve as reference anchors for deconvolution-based spatial mapping (e.g., with Visium or Xenium data) using tools like RCTD or SPOTlight.