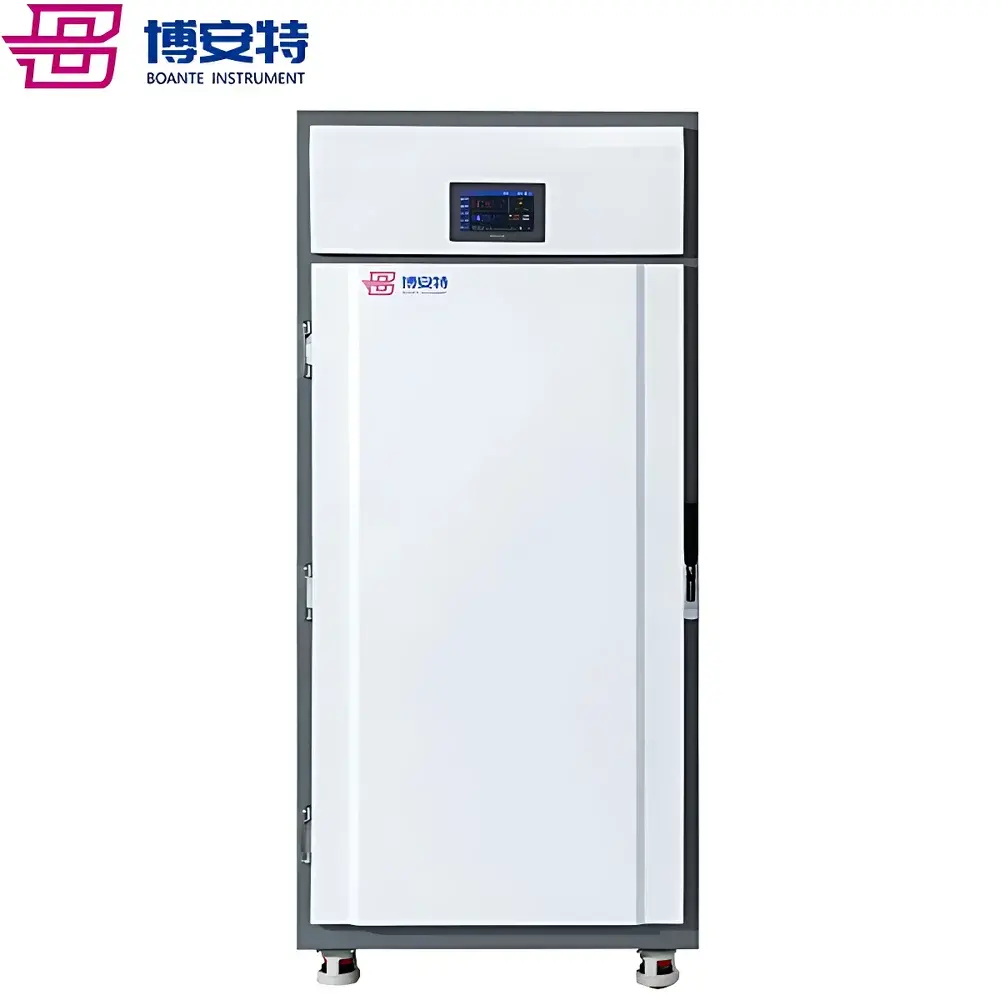
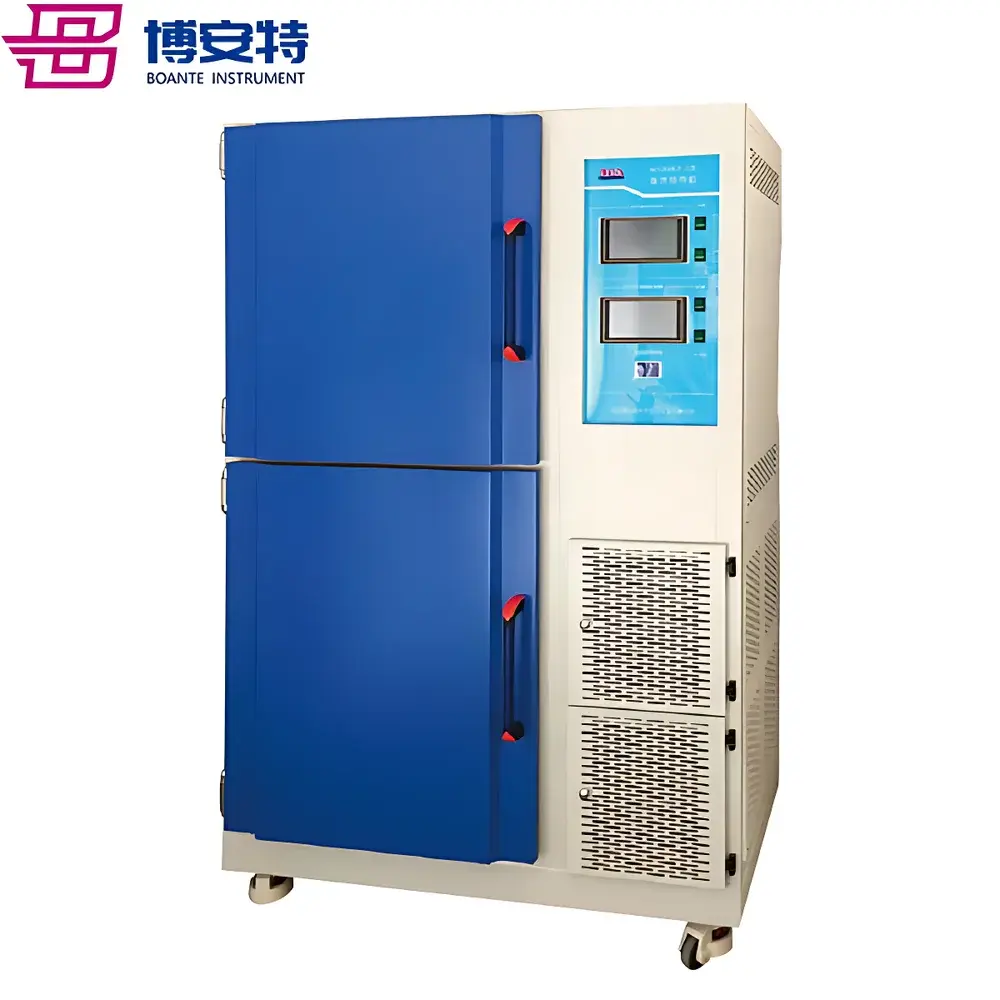
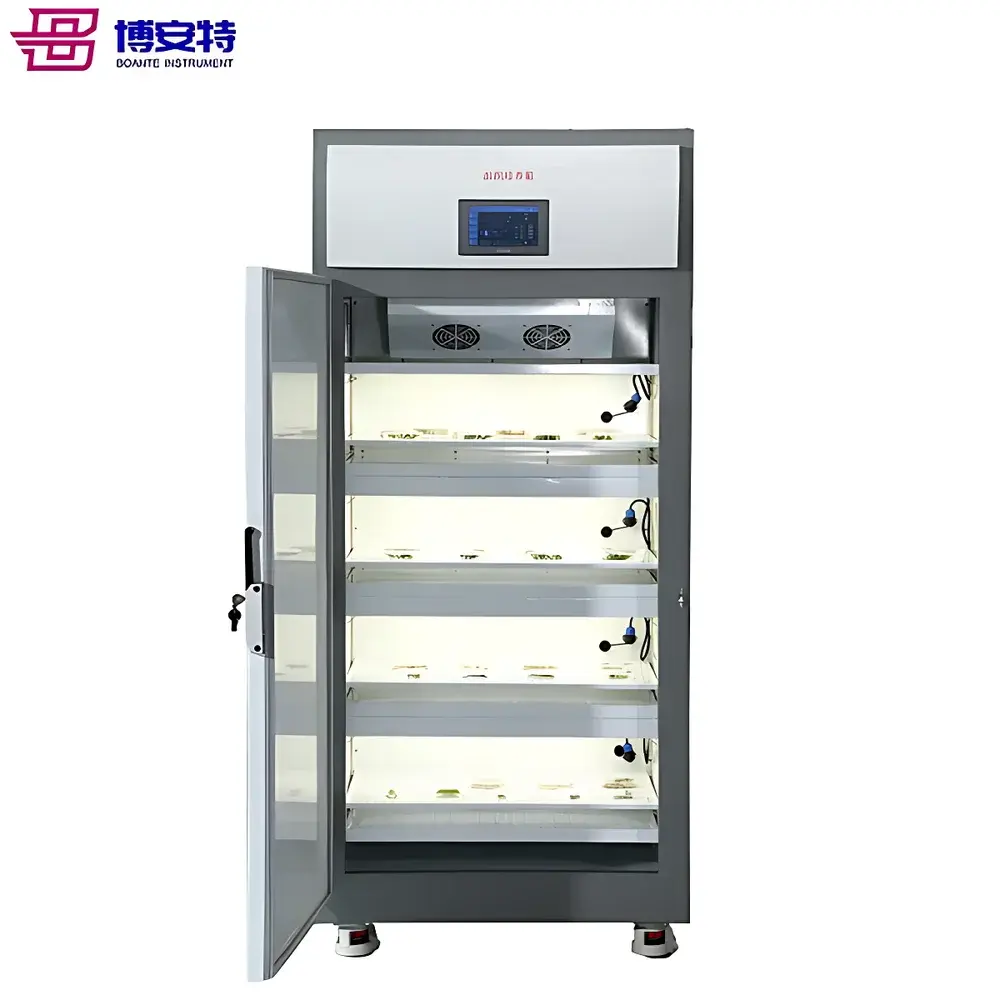
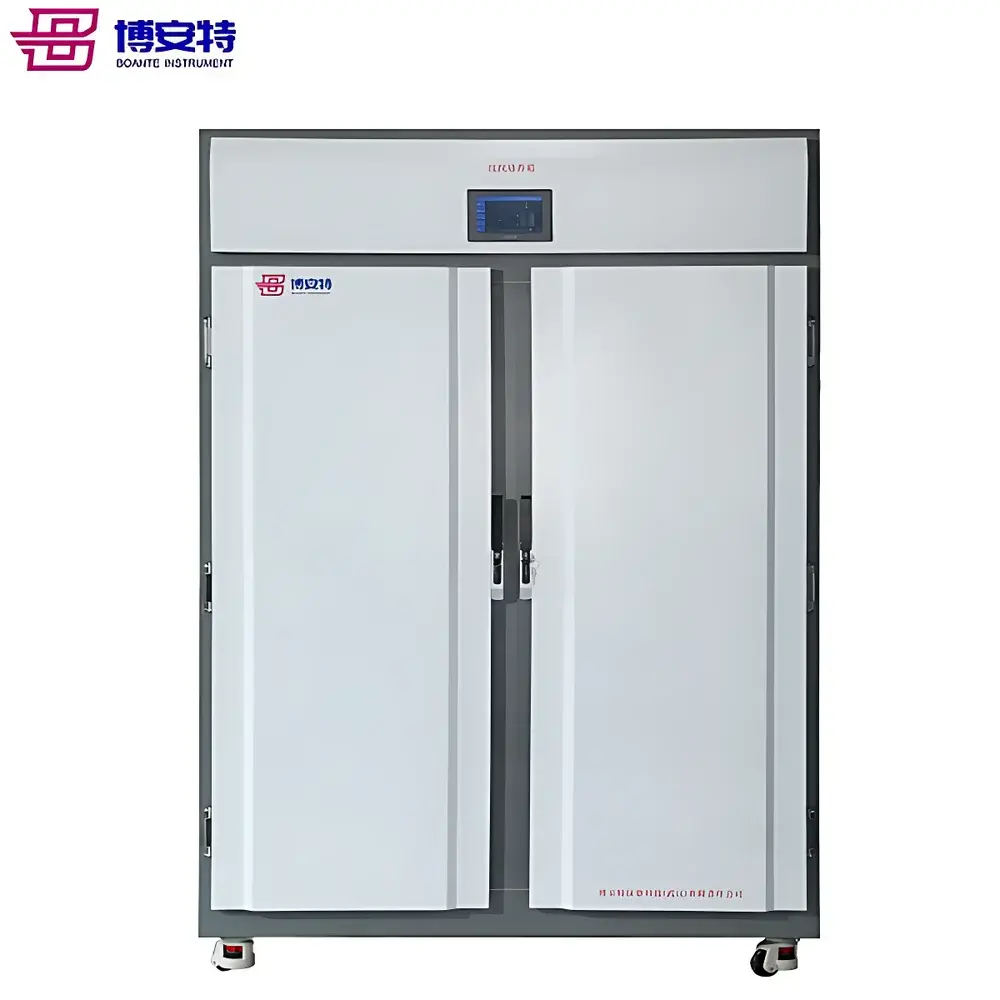
ICOMES Pipetty Semi-Automatic Electric Pipette-Based Nucleic Acid Extractor
| Brand | ICOMES |
|---|---|
| Origin | Japan |
| Manufacturer Type | Authorized Distributor |
| Origin Category | Imported |
| Model | Pipetty |
| Automation Level | Semi-Automatic (Operator-Assisted) |
| Sample Throughput per Run | 1 |
| Sample Volume Range | 1000 µL |
| Typical Processing Time per Sample | ~15 minutes |
Overview
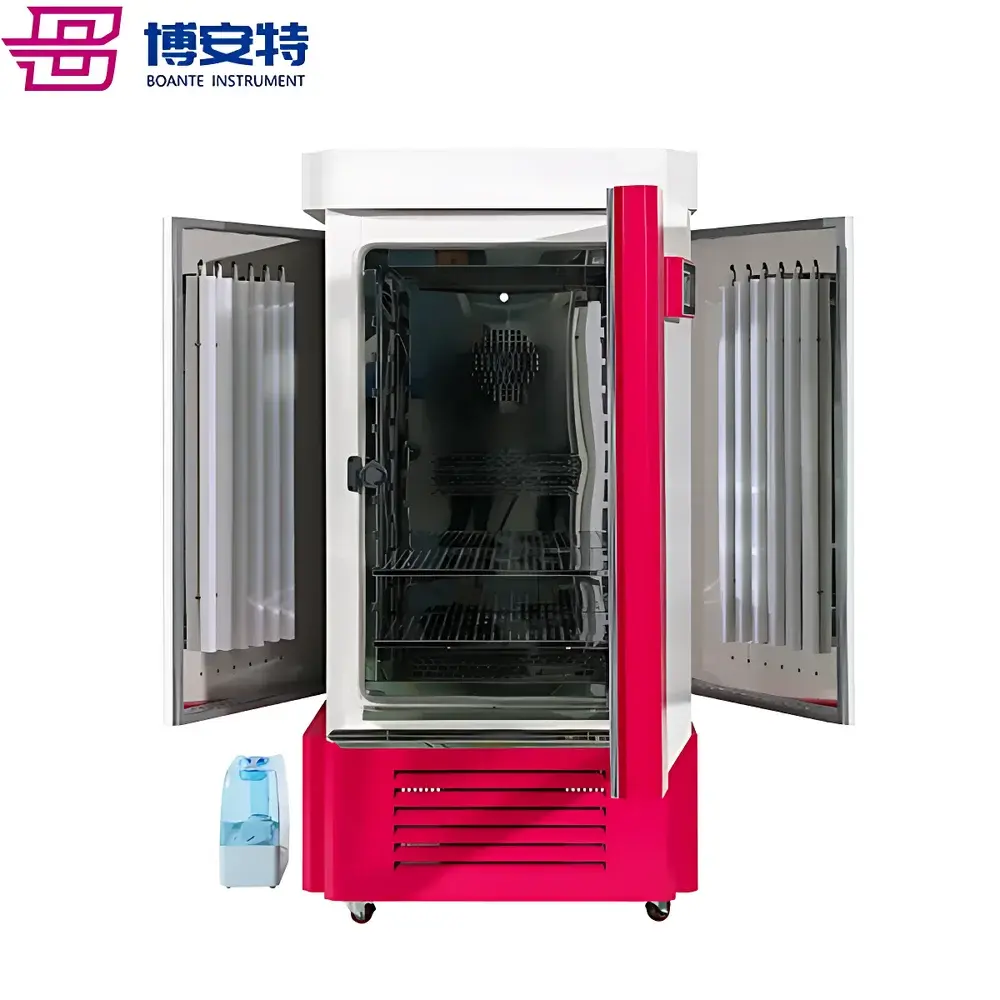
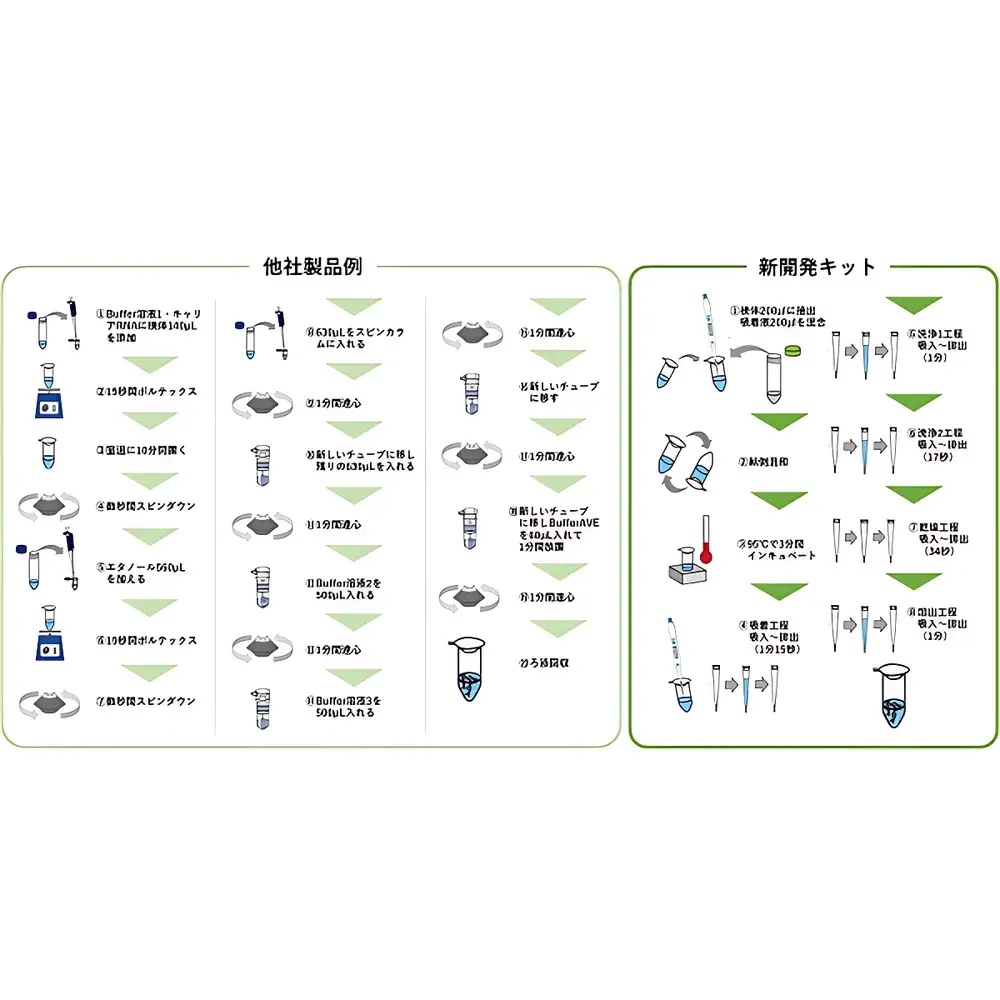
The ICOMES Pipetty Semi-Automatic Electric Pipette-Based Nucleic Acid Extractor represents a paradigm shift in benchtop nucleic acid isolation—replacing centrifugation- and vortex-dependent workflows with a precision fluidic approach grounded in controlled aspiration/dispense kinetics. Unlike conventional spin-column or magnetic bead-based systems, the Pipetty platform leverages proprietary micro-actuator-driven pipetting technology to execute integrated lysis, binding, washing, and elution steps within a single column-style pipette tip. This eliminates mechanical shear stress from high-speed centrifugation and minimizes operator-induced variability by standardizing liquid handling parameters—including flow rate, dwell time, and air-gap control—at the firmware level. Designed for laboratories requiring rapid, reproducible, and equipment-minimal nucleic acid purification—especially in resource-constrained, field-deployable, or GLP-aligned environments—the system delivers consistent DNA and RNA recovery from diverse biological matrices without reliance on external centrifuges, vortex mixers, or heating blocks.
Key Features
- Integrated Column-Tip Architecture: Proprietary monolithic silica-matrix pipette tips enable on-tip binding and sequential buffer exchange via programmable aspiration/dispense cycles—eliminating tube transfers and column centrifugation.
- Nucleic Acid Extraction Mode Firmware: Dedicated extraction protocol embedded in the Pipetty controller ensures precise, repeatable liquid handling: adjustable aspiration speed (0.5–5 µL/s), programmable pause intervals (0.1–30 s), and calibrated elution volume delivery (10–100 µL).
- Centrifuge-Free Operation: All lysis, wash, and elution steps are completed using controlled fluid displacement—no external centrifugation, vortexing, or thermal incubation required.
- Single-Sample Flexibility: Optimized for low-throughput, high-priority applications including clinical sample triage, forensic swabs, or rare cell isolates—ideal where batch processing is impractical or contamination risk must be minimized.
- Room-Temperature Protocol Compatibility: Full workflow executes at ambient temperature (18–25°C), preserving labile RNA integrity and reducing energy consumption versus heated incubation methods.
Sample Compatibility & Compliance
The Pipetty system supports human whole blood (EDTA/K2EDTA), saliva, buccal swabs, cultured cells, and tissue homogenates (up to 25 mg). Eluted nucleic acids meet quality thresholds for downstream qPCR, RT-qPCR, Sanger sequencing, and NGS library prep (RIN ≥ 8.5 for RNA; A260/A280 = 1.8–2.0). The extraction kit reagents comply with ISO 13485:2016 (in vitro diagnostic manufacturing), and the column-tip design conforms to ISO 8573-1 Class 3 particulate specifications. For regulated environments, the Pipetty controller logs timestamped operation records—including tip ID, cycle count, and user-assigned sample ID—supporting audit readiness under FDA 21 CFR Part 11 (when paired with validated LIMS integration) and GLP/GMP documentation requirements.
Software & Data Management
The Pipetty controller features an embedded LCD interface with preloaded extraction protocols (DNA, RNA, dual-analyte), editable step parameters, and real-time progress tracking. All run data—including start/stop timestamps, total cycle duration, and tip usage counter—are stored internally (1,000-run capacity) and exportable via USB-C as CSV files. No cloud connectivity or third-party software dependencies are required. For labs implementing electronic lab notebooks (ELNs) or LIMS, the system supports manual metadata entry of extraction parameters into compliant audit trails. Firmware updates are delivered via signed binary packages to ensure traceability and version control.
Applications
- Rapid point-of-need nucleic acid prep for emergency diagnostics (e.g., respiratory virus screening in mobile clinics)
- Low-volume RNA isolation from limited clinical specimens (e.g., fine-needle aspirates, microdissected FFPE sections)
- QC-grade DNA extraction for STR profiling in forensic casework where cross-contamination avoidance is critical
- Teaching laboratories seeking transparent, hands-on nucleic acid purification principles without centrifuge infrastructure
- Field-deployable genomics workflows in biosurveillance or agricultural pathogen monitoring
FAQ
Does the Pipetty system require calibration before each use?
No—factory-calibrated piston displacement and tip recognition are verified during power-up; annual verification against gravimetric standards (ISO 8655-6) is recommended.
Can the same column-tip be reused across multiple samples?
No—each column-tip is single-use and barcode-tracked to prevent carryover; automated tip ejection prevents manual handling errors.
Is the extracted nucleic acid compatible with automated downstream platforms?
Yes—eluates are in low-EDTA TE buffer (10 mM Tris-HCl, 0.1 mM EDTA, pH 8.0) and compatible with robotic liquid handlers, thermal cyclers, and capillary electrophoresis systems.
What maintenance is required for long-term reliability?
Daily wipe-down of the pipette shaft and monthly cleaning of the tip ejector mechanism with 70% ethanol; no lubrication or internal servicing needed.
How does Pipetty compare to magnetic bead-based extractors in terms of yield consistency?
In head-to-head validation studies (n=48 replicates), Pipetty demonstrated ≤8.2% CV for DNA yield from 200 µL whole blood—comparable to high-end magnetic systems but with significantly reduced inter-operator variance (p < 0.001, ANOVA).