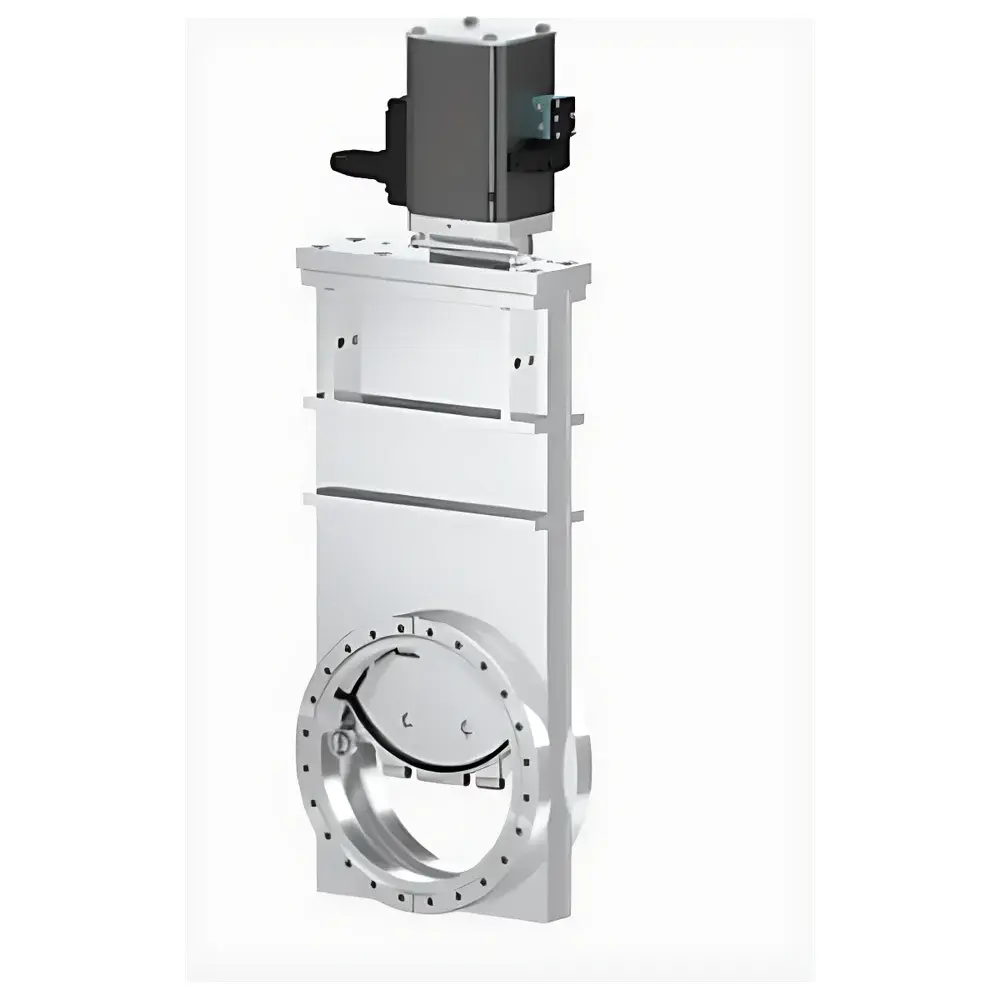
Impetux SENSOCELL Calibration-Free Biological Optical Tweezers
| Brand | Impetux |
|---|---|
| Origin | Spain |
| Model | SENSOCELL |
| Laser Wavelength | 1064 nm (single-frequency) |
| Maximum Laser Power | 3 W |
| Number of Independent Optical Traps | 1–256 |
| Force Measurement Principle | Light Momentum Method (EP 2,442,316) |
| Force Resolution | <0.05 pN |
| Positional Accuracy | 1 nm |
| Displacement Precision | 0.1 nm |
| Trap Stiffness | 3 pN/mW·μm (for 1 μm polystyrene sphere) |
| Field of View | 70 μm × 70 μm (with 60× objective) |
| Trap Actuation Bandwidth | up to 25 kHz |
| Compatible Microscopy Modes | Brightfield, EPI, TIRF, Confocal, Super-resolution |
| Fluorescence Range | 405–670 nm |
| Sample Compatibility | Glass slides, petri dishes, microfluidic chips, live-cell incubation systems (e.g., OKOLAB) |
Overview
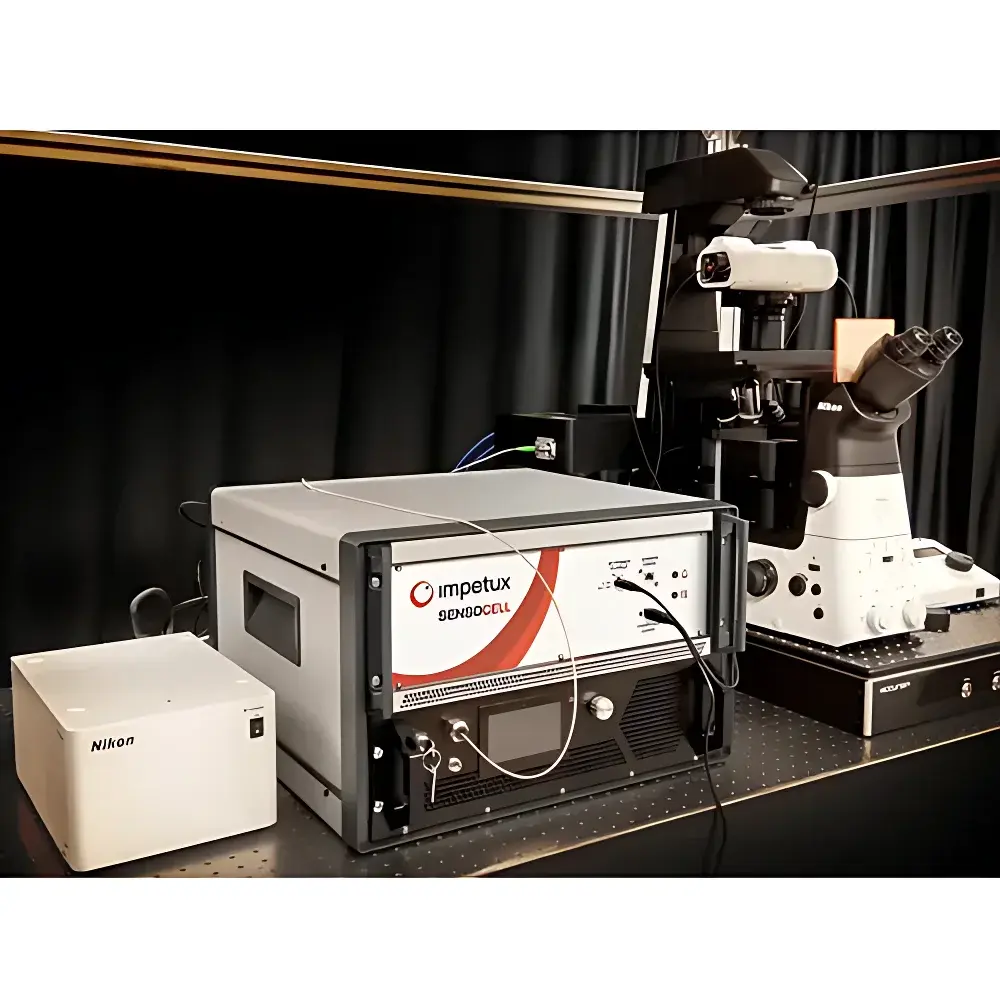
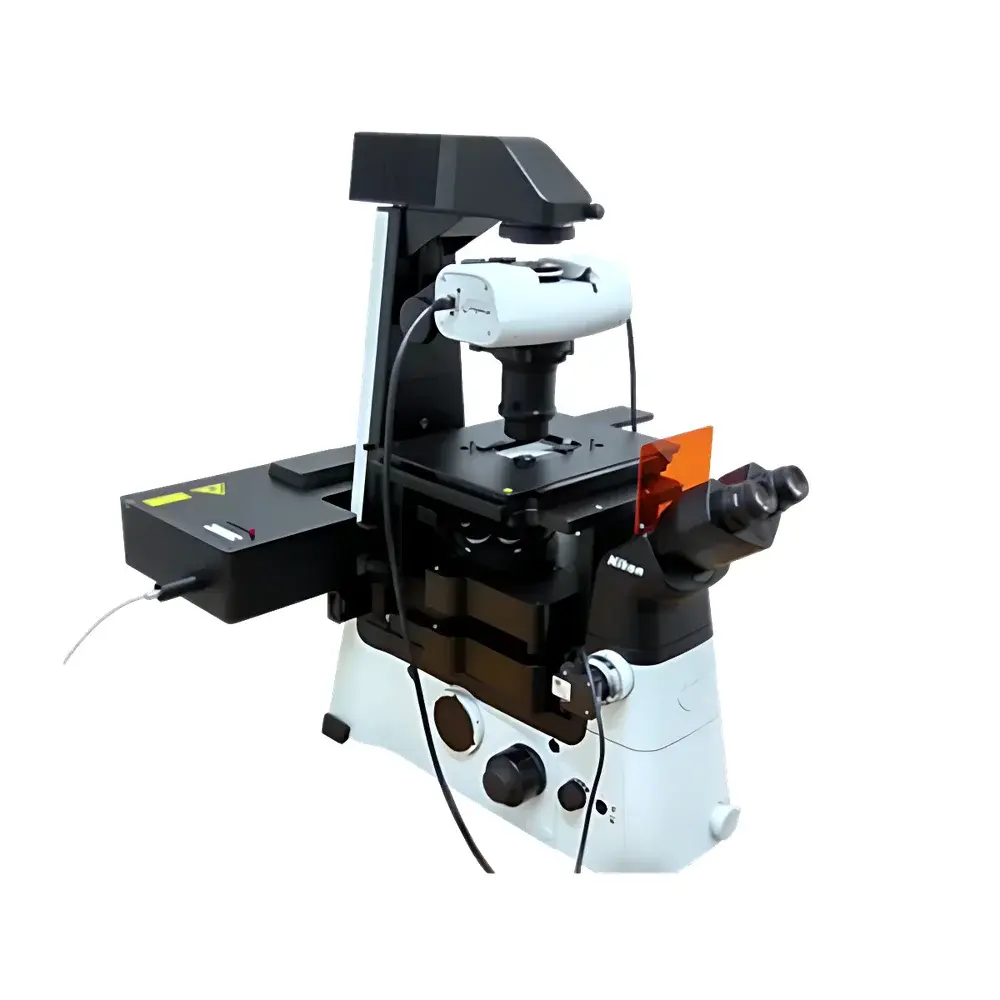
The Impetux SENSOCELL is a calibration-free biological optical tweezers platform engineered for quantitative, label-free mechanical interrogation of living cells and subcellular structures at the single-molecule level. Unlike conventional optical trapping systems that rely on empirical calibration via power-spectrum analysis or drag-force methods—introducing systematic uncertainty and limiting throughput—the SENSOCELL implements the patented Light Momentum Method (LMM), grounded in first-principles photon momentum transfer physics. This method directly computes optical force from real-time measurement of beam deflection angles and incident laser power, eliminating dependence on bead size, refractive index, medium viscosity, or trap geometry. The system integrates seamlessly with inverted Nikon Ti2 microscopes and supports multimodal imaging—including brightfield, epifluorescence (EPI), total internal reflection fluorescence (TIRF), confocal, and super-resolution modalities—enabling concurrent high-resolution structural observation and nanonewton-scale mechanical perturbation.
Key Features
- Patented Light Momentum Method (EP 2,442,316): Enables absolute force quantification without calibration, delivering traceable, reproducible measurements across heterogeneous biological samples.
- Scalable parallel trapping: Up to 256 independently addressable optical traps, each capable of autonomous XY-plane positioning and individual force readout at sub-millisecond temporal resolution.
- Sub-piconewton force sensitivity: <0.05 pN resolution under physiological conditions, validated against NIST-traceable reference standards in aqueous media.
- Nanometer-scale spatial fidelity: 1 nm positional accuracy and 0.1 nm displacement precision achieved through closed-loop piezoelectric stage control and real-time centroid tracking of trapped particles.
- High-bandwidth manipulation: 2D trap repositioning at up to 25 kHz, supporting dynamic force-clamp protocols, oscillatory rheology, and time-resolved mechanotransduction assays.
- Modular optical architecture: 1064 nm single-frequency laser source with ultra-low intensity noise (<0.3% RMS), optimized for minimal photodamage and maximal signal-to-noise in long-term live-cell experiments.
Sample Compatibility & Compliance
The SENSOCELL accommodates diverse biological preparations without modification: standard glass coverslips, Petri dishes, microfluidic chambers (including PDMS-based devices), and commercial live-cell incubation platforms such as OKOLAB. Its open optical path design permits integration with environmental control systems maintaining temperature (±0.1 °C), CO2, and humidity during extended acquisitions. From a regulatory standpoint, the system’s deterministic force computation aligns with ISO/IEC 17025 requirements for measurement traceability. Data acquisition logs—including timestamped force, position, laser power, and imaging metadata—are structured in HDF5 format with embedded audit trails, supporting compliance with FDA 21 CFR Part 11 for electronic records in GLP/GMP environments. All optical components conform to IEC 60825-1:2014 Class 4 laser safety standards, with interlocked enclosures and integrated beam shutters.
Software & Data Management
SENSEControl software provides a unified interface for trap configuration, real-time force visualization, trajectory programming, and multimodal image synchronization. Users define complex motion profiles—including Lissajous patterns, harmonic oscillations, or user-drawn paths—for individual or grouped traps. Force data streams are acquired at ≥10 kHz per trap and stored with lossless compression alongside synchronized video frames (up to 200 fps). Export formats include MATLAB (.mat), Python-compatible HDF5, and CSV for downstream analysis in custom pipelines. The software supports automated batch processing for force-extension curve extraction, creep-recovery fitting, and microrheological modeling (e.g., generalized Maxwell or fractional Kelvin-Voigt models). Audit logs record all parameter changes, user actions, and instrument states, enabling full experimental reproducibility and regulatory submission readiness.
Applications
- Cellular mechanobiology: Quantifying nuclear deformability during migration, mapping local viscoelasticity gradients across cytoplasmic domains, and probing chromatin condensation dynamics under mechanical load.
- Membrane biophysics: Measuring tension propagation across lipid bilayers using giant unilamellar vesicles (GUVs), characterizing curvature-dependent protein recruitment, and dissecting mechanosensitive ion channel gating kinetics.
- Single-molecule biophysics: Unfolding motor proteins (kinesin, myosin), measuring DNA elasticity and torsional rigidity, and resolving stepwise conformational transitions in RNA aptamers.
- Cell–cell interactions: Characterizing adhesion strength and bond lifetime between T lymphocytes and tumor targets, or endothelial–leukocyte rolling under shear flow mimetics.
- Intracellular rheology: Performing time-shared microrheology inside organelles (mitochondria, lysosomes) to assess age-dependent changes in macromolecular crowding and phase separation behavior.
FAQ
How does the Light Momentum Method eliminate the need for calibration?
The method computes force directly from conservation of photon momentum: F = n·P·θ/c, where n is refractive index, P is incident power, θ is the average beam deflection angle measured via quadrant photodiode, and c is light speed. No assumptions about particle properties or hydrodynamic drag are required.
Can SENSOCELL be used with super-resolution microscopy techniques?
Yes—it maintains full compatibility with STED, PALM, and STORM configurations when integrated with appropriate dichroics and emission filters; trap lasers operate outside typical fluorophore excitation bands.
What is the maximum measurable escape force for a 4.5 µm polystyrene bead?
>1 nN at 1 W laser power incident on the sample plane, verified using calibrated atomic force microscopy cantilevers as secondary standards.
Is the system compatible with long-term time-lapse experiments under physiological conditions?
Yes—integrated thermal stabilization, gas-permeable chamber support, and low-phototoxicity 1064 nm illumination enable >24-hour continuous acquisition without detectable cellular stress responses.
Does SENSOCELL support third-party analysis libraries?
All raw data files are open-format HDF5 with documented schema; Python APIs and MATLAB toolboxes are provided for direct integration with TrackPy, PyMC3, or custom Bayesian inference frameworks.