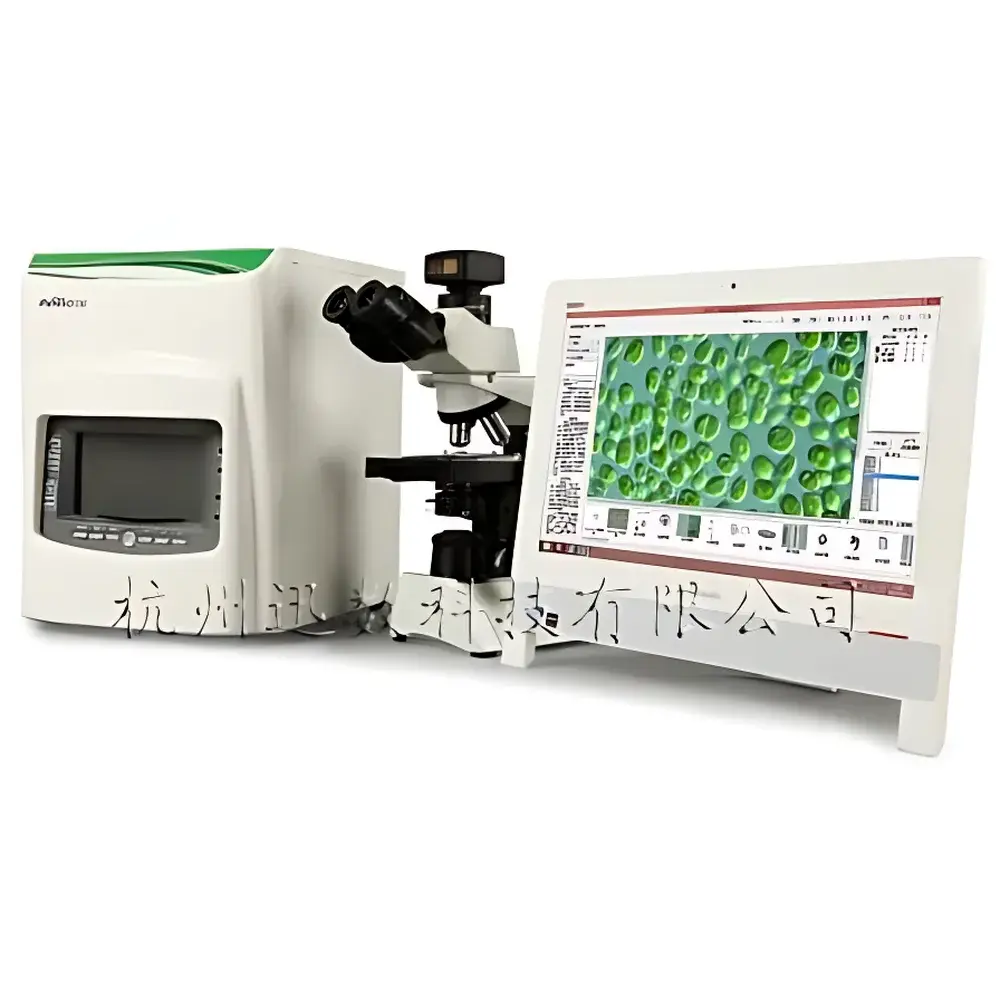
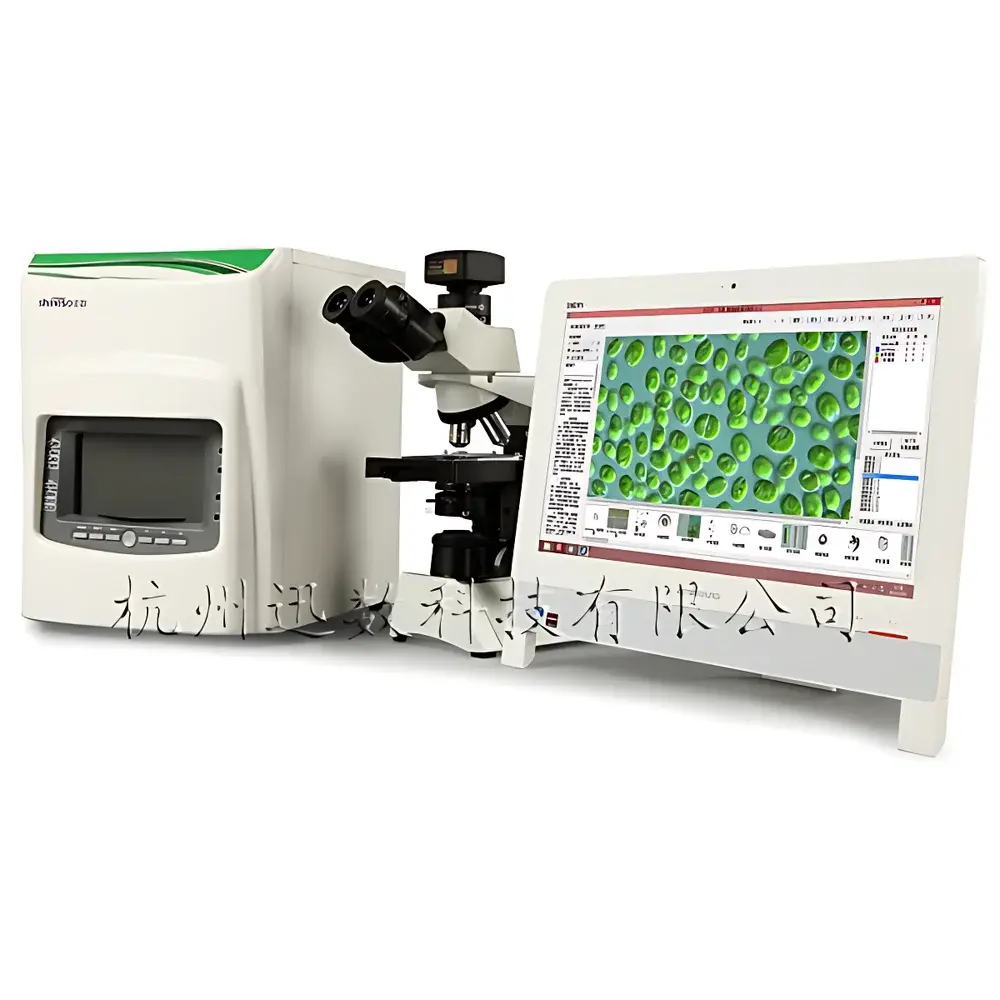
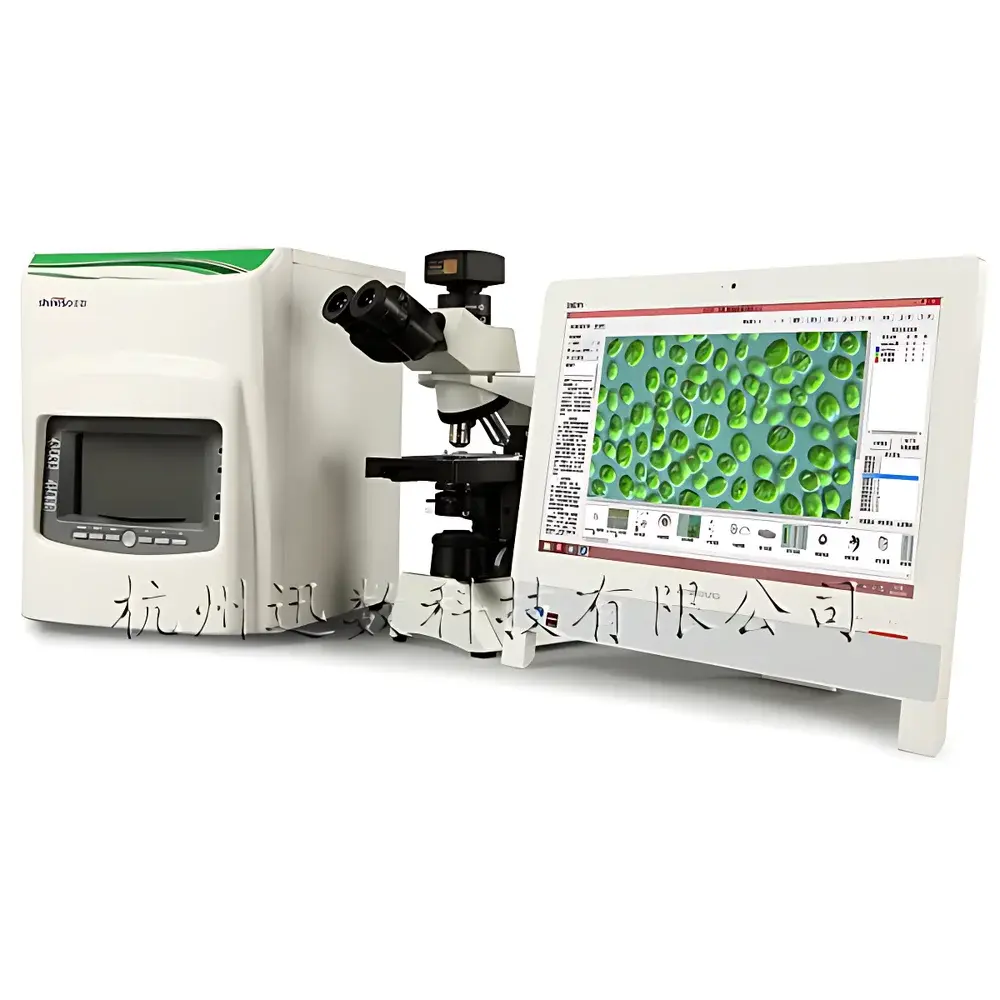
Xunshu Algacount M500 Integrated Microbial Colony Counter and Plankton Analysis System
| Brand | Xunshu |
|---|---|
| Origin | Zhejiang, China |
| Manufacturer Type | OEM Manufacturer |
| Product Category | Domestic |
| Model | Algacount M500 |
| Instrument Type | Algal Counter and Plankton Analyzer |
| Price Range | USD 150–15,000 (FOB) |
Overview
The Xunshu Algacount M500 is an integrated digital imaging platform engineered for high-precision, standardized biological monitoring in aquatic environments. It combines automated microbial colony enumeration with advanced image-based plankton analysis—spanning phytoplankton (algae) and zooplankton—within a single instrument architecture. The system operates on the principle of high-resolution digital microscopy coupled with supervised machine learning algorithms for morphological classification and quantitative image segmentation. Its dual-domain capability supports regulatory-compliant water quality assessment under ISO 19458 (water microbiology), ASTM D5763 (algal enumeration), and national environmental monitoring protocols for eutrophication, harmful algal bloom (HAB) surveillance, and sanitary indicator testing. Designed for freshwater, estuarine, and marine laboratories, the M500 delivers reproducible, auditable results without requiring manual taxonomic expertise at the point of analysis.
Key Features
- Unified hardware platform supporting both colony counting (bacteria, coliforms, fungi) and plankton analysis (phytoplankton & zooplankton) via interchangeable optical modules.
- Triple-illumination system: top-mounted 360° shadowless LED illumination, bottom-mounted dark-field suspension lighting, and optional 254 nm UV for chamber sterilization and mutagenesis studies.
- High-fidelity imaging chain: 8-megapixel color CMOS sensor (1/2.5″), C-mount industrial lens (f/1.4–f/32), distortion <1%, paired with scientific-grade CCD for microscopical imaging (2580 × 1944 resolution).
- Proprietary “Biological Similarity Search” engine: integrates color, texture, and geometric descriptors into SVM-trained feature vectors for species-level algal identification across 8,093 validated taxa.
- Multi-layer depth fusion and ultra-wide field stitching for accurate analysis of filamentous, colonial, and chain-forming algae under high-magnification objectives.
- Adaptive segmentation algorithms: “Ovoid Cell Counting” and “Complex Cell Counting” modes optimized for microalgal biotechnology applications including biofuel feedstock quantification and novel food ingredient QC.
- Fully compliant with GLP data integrity requirements: role-based user permissions, audit trail logging, electronic signature support, and 21 CFR Part 11–ready export formats.
Sample Compatibility & Compliance
The Algacount M500 accepts standard microbiological media formats—including pour plates, spread plates, membrane filters (black-grid), and 3M Petrifilm™ test strips—enabling enumeration of total viable count, total coliforms, fecal coliforms, E. coli, and Staphylococcus aureus per ISO 4833-1, ISO 9308-1, and USP <61>. For plankton analysis, it processes preserved or live samples mounted on standard microscope slides or sedimentation chambers. The embedded expert database covers 11 algal phyla (Cyanophyta, Chlorophyta, Bacillariophyta, etc.), 862 genera, and 8,093 species—with separate taxonomic trees for marine (2,818 spp.) and freshwater (5,275 spp.) assemblages. Zooplankton coverage includes 23 functional/morphological classes—from protozoans to copepodids—with bilingual (Latin + Chinese) nomenclature, verified micrographs, hand-drawn schematics, and diagnostic morphometric annotations. All reference data comply with the AlgaeBase taxonomy framework and are extensible by users under version-controlled schema.
Software & Data Management
The bundled software suite comprises three core modules: MIC (Microbial Image Counting), PlanktonIQ (Plankton Identification & Quantification), and BioStat (Biological Statistics & Reporting). Each module enforces traceable workflow logging, parameterized batch processing, and metadata tagging (sample ID, date, operator, magnification, staining protocol). Statistical outputs include dominance indices (Shannon-Wiener, Simpson), relative abundance heatmaps, size-class distribution histograms, and biomass estimation using geometric models (e.g., prolate spheroid for diatoms, cylinder for filaments). Raw image files are stored in lossless TIFF format; processed data exports to Excel (.xlsx) with embedded formulas for dilution factor compensation, colony-forming unit (CFU) conversion, and cell density normalization (cells/mL or cells/L). Audit trails record all user actions—including manual corrections, threshold adjustments, and database edits—with timestamps and login identifiers.
Applications
The Algacount M500 serves critical functions across environmental, public health, and industrial research domains. In municipal water utilities, it accelerates compliance reporting for coliform monitoring and algal toxin precursor screening. Ecological monitoring agencies deploy it for long-term trend analysis of phytoplankton community shifts linked to climate change or nutrient loading. Aquaculture facilities use its microalgal concentration assays to optimize feeding regimes and detect opportunistic pathogens. Academic labs apply its pigment body quantification tools (chlorophyll-a, fucoxanthin, phycocyanin localization) to study photosynthetic efficiency and stress physiology. Additionally, the system supports method validation studies for ISO/IEC 17025-accredited laboratories seeking accreditation for phytoplankton enumeration (ISO 14442) and microbial enumeration (ISO 7218).
FAQ
Does the Algacount M500 support FDA 21 CFR Part 11 compliance?
Yes—the software implements electronic signatures, audit trails, and role-based access controls aligned with Part 11 requirements for regulated environments.
Can users expand the built-in algal database?
Yes—users may import validated micrographs, morphological descriptions, and taxonomic metadata via structured CSV templates; all additions undergo local versioning and optional peer review flagging.
What is the minimum detectable cell size for microalgal counting?
The system reliably resolves and segments single cells ≥3 µm in diameter under 40× objective magnification, subject to contrast and staining quality.
Is calibration traceable to NIST standards?
Yes—calibration routines include pixel-to-micron mapping using certified stage micrometers; calibration logs are retained with uncertainty estimates.
How does the “Chaos-Assisted Classification” algorithm differ from conventional CNN-based identification?
It applies deterministic chaos theory to quantify morphological variability within mixed populations, enabling robust classification when training data is sparse or taxonomically ambiguous—without requiring deep neural network retraining.