COMECAUSE IN+Pheno200 High-Throughput Hyperspectral Plant Phenotyping Imaging System
| Brand | COMECAUSE |
|---|---|
| Origin | Shandong, China |
| Manufacturer Type | OEM Manufacturer |
| Country of Origin | China |
| Model | IN+Pheno200 |
| Price | USD 77,000 (FOB) |
| Max Plant Height | 300–1000 mm |
| Max Plant Width | 100–500 mm |
| Max Fresh Weight | 100–75,000 g |
| Enclosure Dimensions | 1400 × 830 × 2140 mm (L×W×H) |
| Camera Resolution | Up to 12 MP |
| Acquisition Time per Plant | ≤40 s |
| Analysis Time per Plant | ≤30 s |
| Spectral Range | 400–1000 nm (VIS–NIR) |
| Number of Spectral Bands | 58 calibrated indices |
| 3D Reconstruction | Multi-view stereo photogrammetry with AI-assisted mesh generation |
| Environmental Sensors | Integrated temperature & humidity monitoring |
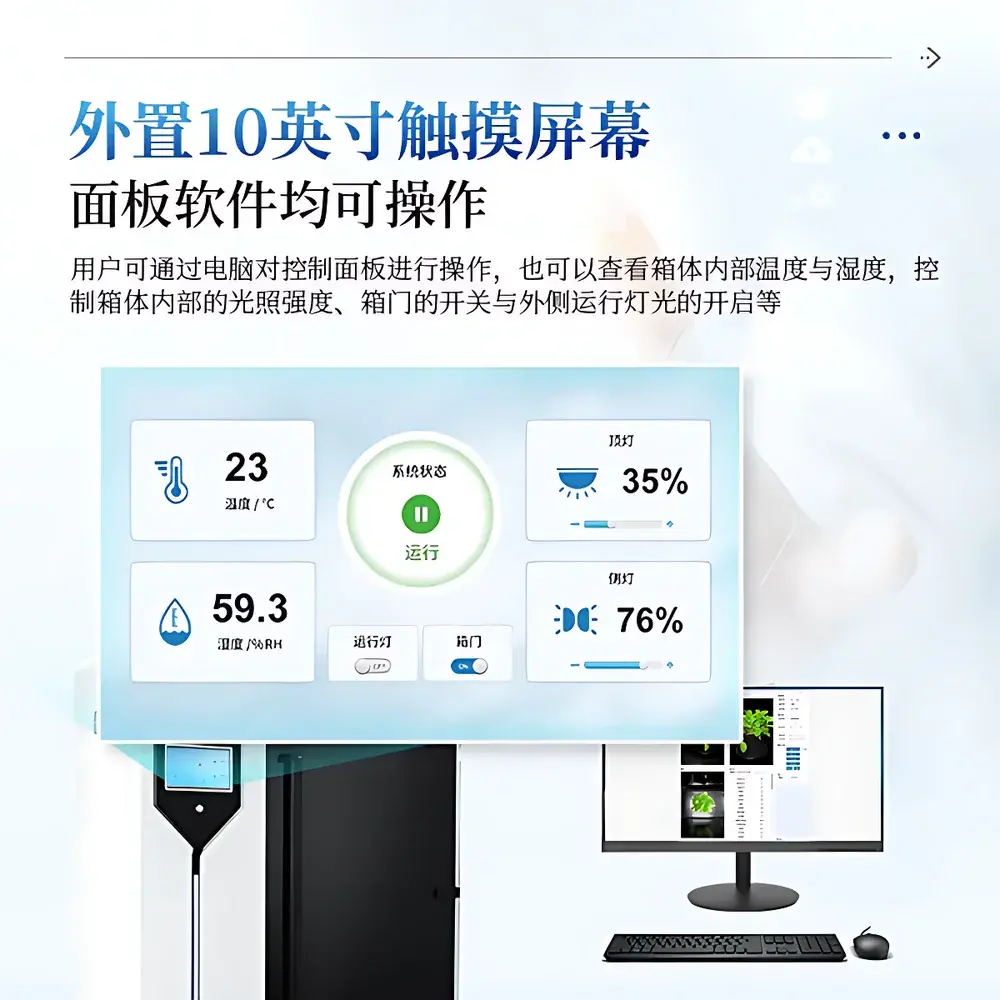
| Control Interface | 10-inch industrial touchscreen + Windows-based desktop software (Intel i5+, 8 GB RAM, NVIDIA GPU ≥8 GB VRAM) |
| Data Export Formats | JSON (analysis_result.json), SPE (hyperspectral raw), CSV, MP4 (3D model video), PNG/TIFF (2D/3D visualizations) |
| Compliance | Designed for GLP-aligned workflows |
Overview
The COMECAUSE IN+Pheno200 is a fully integrated, high-throughput hyperspectral plant phenotyping imaging system engineered for non-invasive, quantitative, and temporally resolved characterization of morphological, physiological, and stress-responsive traits across the entire plant growth cycle. Operating on the principles of multi-spectral reflectance spectroscopy combined with structured-light-free 3D photogrammetric reconstruction, the system captures synchronized visible (RGB), near-infrared (NIR), and hyperspectral (400–1000 nm) data from top, side-top, and side-bottom perspectives—enabled by a motorized rotating turntable and programmable optical positioning. Unlike conventional single-modality scanners, the IN+Pheno200 functions as a unified measurement platform where structural geometry, pigment composition, water status, and photosynthetic efficiency are co-registered in spatial and temporal domains. Its architecture adheres to metrological traceability standards common in academic phenomics facilities and commercial breeding programs, supporting repeatable acquisition under controlled-environment conditions (e.g., growth chambers or climate rooms). The system does not require sample labeling, physical contact, or destructive sampling—ensuring longitudinal tracking of individual genotypes without biological perturbation.
Key Features
- Multi-angle, multi-modal imaging: Simultaneous acquisition from three orthogonal viewpoints using calibrated RGB and hyperspectral sensors (58 spectral indices pre-validated against reference standards)
- AI-enhanced 3D reconstruction: Real-time generation of textured, georeferenced 3D point clouds and mesh models via multi-view stereo algorithms—enabling volumetric, angular, and topological trait quantification (e.g., canopy volume, branch angle distribution, fractal dimension)
- Automated adaptive imaging: Dynamic resolution scaling and image stitching for specimens exceeding field-of-view constraints; auto-focus and motorized lens elevation control ensure optimal optical geometry per sample
- Integrated environmental monitoring: Onboard temperature and relative humidity sensors log ambient conditions during each acquisition cycle—critical for disentangling genotype-by-environment (G×E) interactions
- High-throughput pipeline: 2D analysis completed in ≤60 s per plant; 3D modeling runs asynchronously in background queue—enabling continuous batch processing without workflow interruption
- Cloud-enabled data management: Secure device-bound upload of phenotype metrics (JSON), raw spectra (SPE), and 3D model videos (MP4) to encrypted cloud storage with role-based access control
- Fully localized software interface: One-click switching between English and Chinese UI; optimized for Windows OS with NVIDIA GPU acceleration (≥8 GB VRAM) for real-time rendering and batch computation
Sample Compatibility & Compliance
The IN+Pheno200 accommodates intact, living plants within defined dimensional and mass limits (height: 300–1000 mm; width: 100–500 mm; fresh weight: 100–75,000 g), including seedlings, rosettes, tillered cereals, and upright dicots from Poaceae, Solanaceae, Brassicaceae, and Fabaceae families. Its enclosed cabinet design (1400 × 830 × 2140 mm) ensures stable illumination and minimal external interference—meeting requirements for reproducible phenotyping under ISO 22716-aligned facility protocols. While not certified to FDA 21 CFR Part 11 out-of-the-box, the system’s software architecture supports ALCOA+ data integrity principles: all acquisitions are timestamped, operator-logged, and associated with environmental metadata and instrument configuration snapshots—facilitating GLP/GMP-compliant audit trails when deployed in regulated research environments. Calibration procedures follow NIST-traceable reference standards for spectral radiance and geometric scale bars.
Software & Data Management
The proprietary IN+PhenoSuite software provides modular, workflow-driven operation—from acquisition scheduling and hardware control to trait extraction and statistical export. The 2D analysis module computes 120+ morphometric, chromatic, and textural descriptors (e.g., projected area ratio, Lab color space parameters under D65/2°, ASM, SSIM, entropy, homogeneity) directly from orthorectified top/side views. The 3D analysis engine outputs spatially anchored metrics—including stem height, convex hull volume, skeletonized branch topology, and leaf inclination angles—derived from reconstructed surface meshes. All computed values are stored in structured JSON format with embedded provenance metadata. Raw hyperspectral cubes (in SPE format) and region-of-interest extractions support downstream chemometric analysis (e.g., PLSR, PCA) in third-party tools like MATLAB or R. Export options include batch CSV reports, interactive HTML trait dashboards, and annotated 3D model videos compatible with standard media players or VR-ready viewers.
Applications
The IN+Pheno200 serves as a foundational infrastructure tool across academic and industrial plant science domains. In functional genomics, it enables high-resolution QTL mapping and genome-wide association studies (GWAS) by generating dense, time-series phenotypic profiles for mutant libraries or natural populations under controlled abiotic stresses (drought, heat, nutrient deficiency). Within breeding pipelines, it supports early-generation selection for ideotype traits—such as compact architecture, delayed senescence, or stomatal conductance proxies—thereby increasing selection accuracy and shortening breeding cycles. Ecophysiologists employ its thermal-agnostic NIR reflectance indices to quantify photosynthetic performance (e.g., NDVI, PRI, MCARI) alongside structural metrics, revealing trade-offs between carbon gain and hydraulic safety. Additionally, the system supports method development for digital twin modeling—where empirical 3D morphology and spectral response inform virtual crop simulation frameworks used in climate-resilience forecasting.
FAQ
What spectral range does the hyperspectral camera cover?
The system operates across 400–1000 nm (visible to near-infrared), with 58 pre-processed spectral indices derived from calibrated reflectance measurements.
Is the 3D model exportable for use in external simulation software?
Yes—textured OBJ and PLY files, along with metadata-rich JSON summaries, can be exported for integration into Unity, Blender, or crop modeling platforms such as OpenSimRoot or APSIM.
Does the system support remote operation and monitoring?
Remote desktop access is supported via secure LAN/WAN connections; however, real-time sensor feedback and hardware actuation require local network proximity for latency-sensitive control loops.
Can the system be validated against established phenotyping benchmarks?
Yes—reference datasets from the European Plant Phenotyping Network (EPPN2020) and standardized protocols from the International Plant Phenotyping Network (IPPN) are compatible with IN+Pheno200 output formats.
What computing resources are required for full 3D reconstruction and batch analysis?
A workstation with Intel Core i7/i9 (or AMD Ryzen 7/9), ≥32 GB RAM, and NVIDIA RTX A4000/A5000 (or equivalent) is recommended for sub-minute 3D mesh generation across >100 samples per batch.