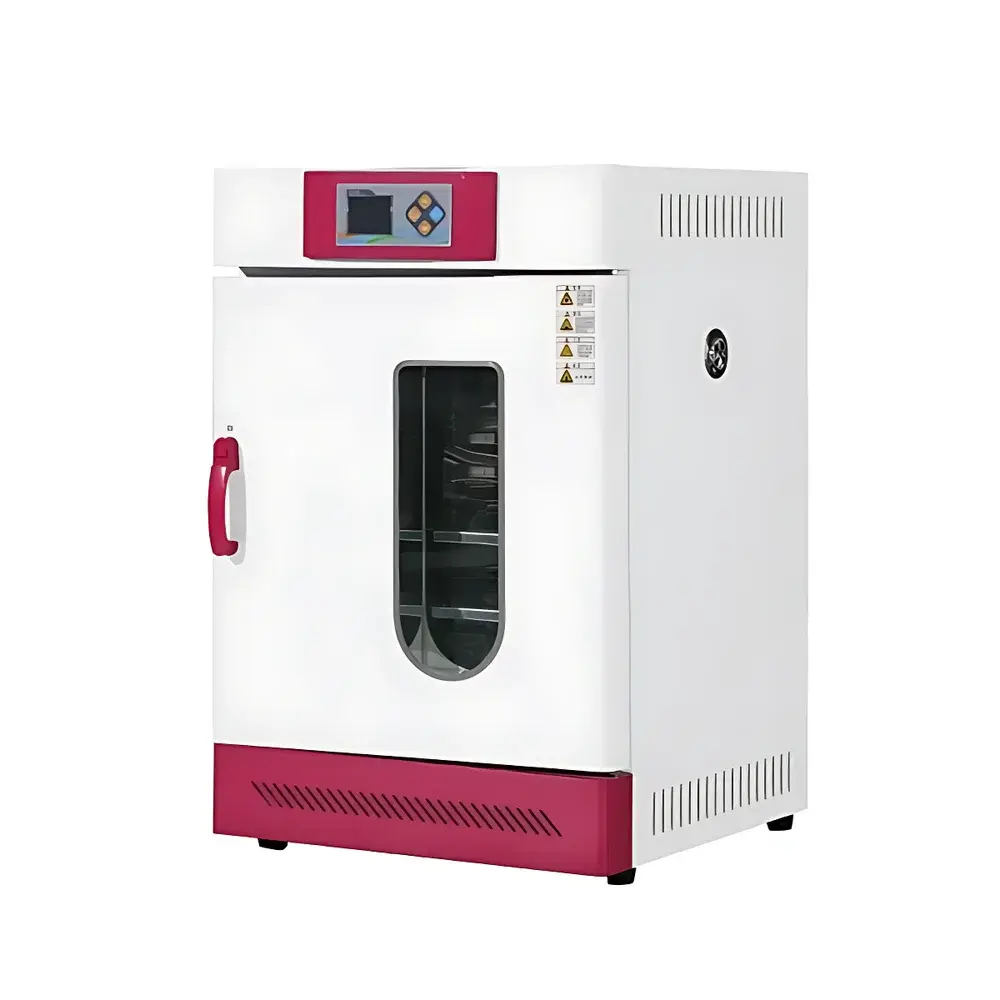
COMECAUSE IN~Pheno50 Plant Phenotyping Imaging System
| Brand | COMECAUSE |
|---|---|
| Origin | Shandong, China |
| Manufacturer Type | Direct Manufacturer |
| Country of Origin | China |
| Model | IN~Pheno50 |
| Price | USD 76,500 (FOB Qingdao, ex-works) |
| Max Plant Height | 300–1000 mm |
| Max Plant Width | 100–500 mm |
| Max Fresh Weight | 100–75,000 g |
| Enclosure Dimensions | 1400 × 830 × 2140 mm (L×W×H) |
| Camera Resolution | Up to 12 MP |
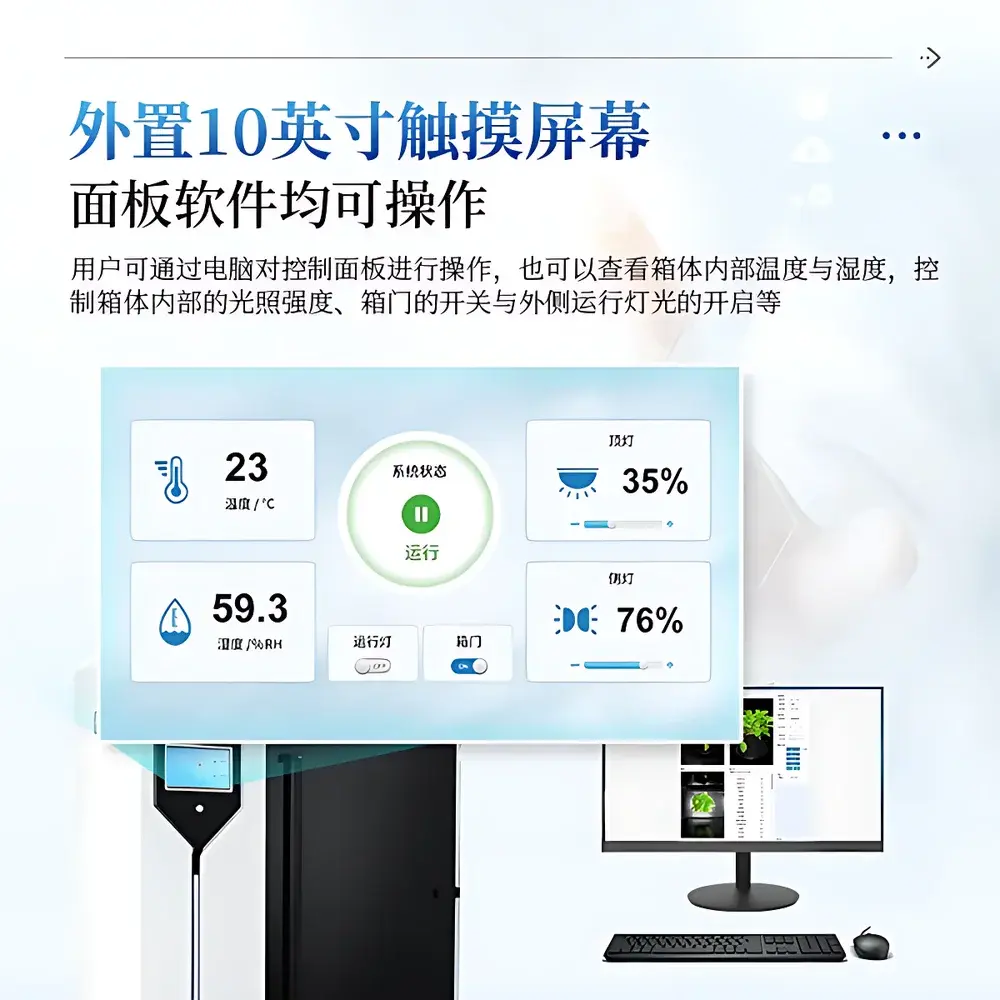
| Display | 10-inch external touchscreen interface |
| OS Requirement | Windows 10/11, Intel Core i5 or higher, ≥8 GB RAM, NVIDIA GPU with ≥8 GB VRAM |
| Environmental Sensors | Integrated temperature & humidity monitoring |
| Compliance | Designed for GLP-aligned workflows |
Overview
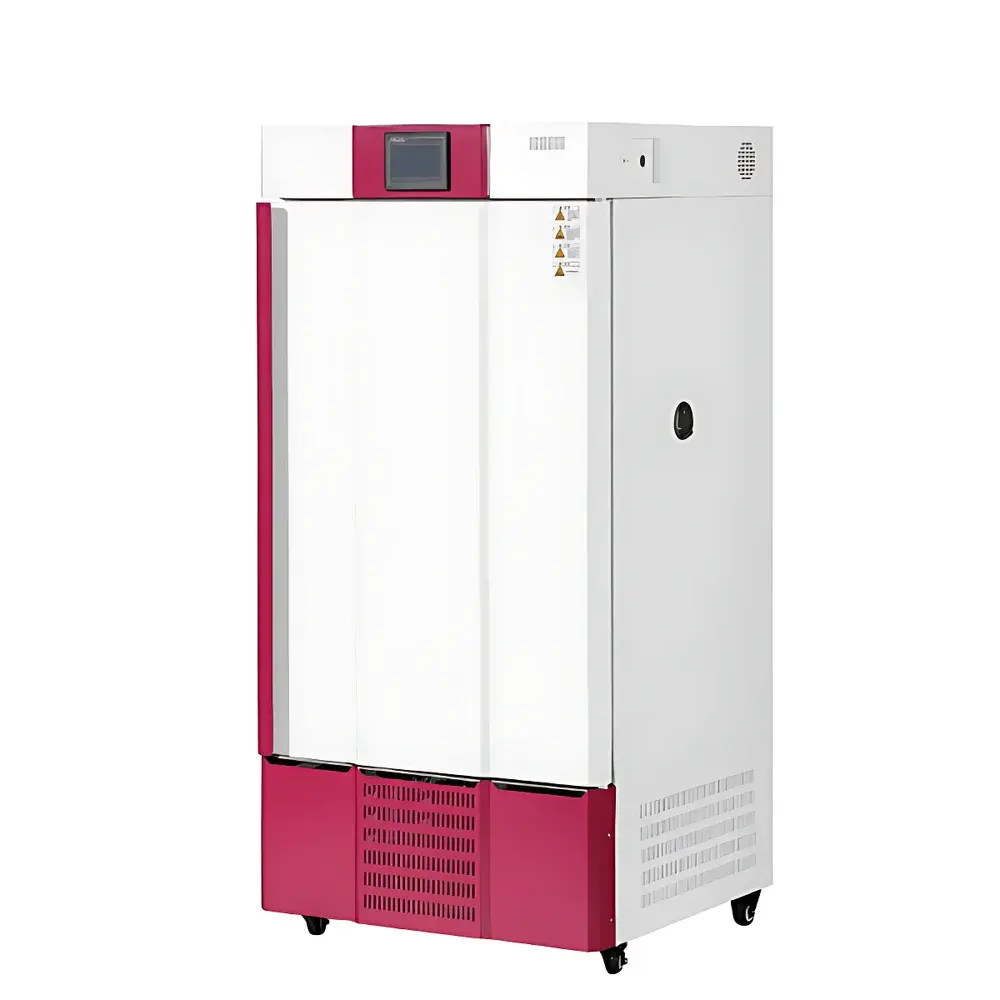
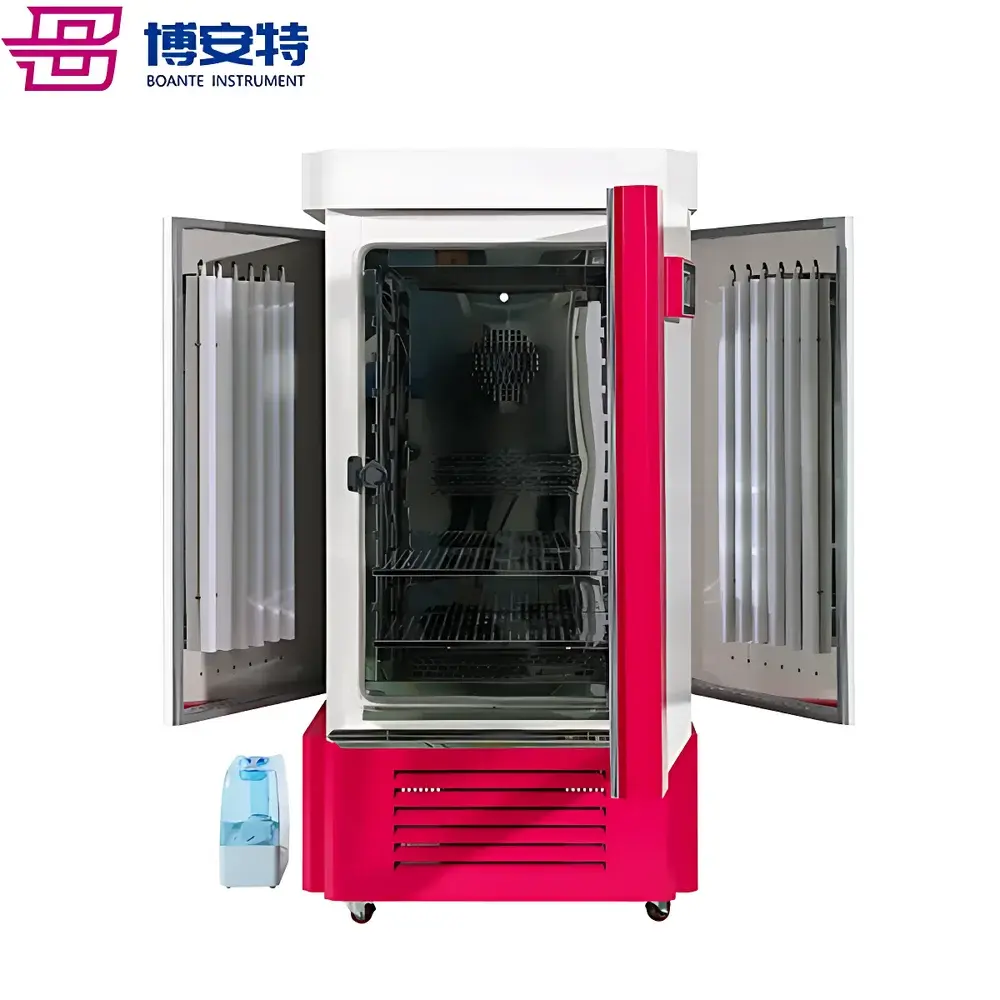
The COMECAUSE IN~Pheno50 Plant Phenotyping Imaging System is a modular, enclosed high-throughput phenotyping platform engineered for non-destructive, quantitative assessment of plant morphological, chromatic, and textural traits across developmental stages. It operates on a multi-view structured-light imaging principle combined with AI-driven computer vision algorithms, enabling dynamic reconstruction of plant geometry from synchronized top, upper-side, and lower-side visible-light image sequences. Unlike conventional single-angle scanners, the IN~Pheno50 integrates a motorized rotating turntable with programmable vertical lens positioning, allowing full 360° circumferential acquisition and adaptive focal plane alignment. The system delivers both rapid 2D trait extraction and high-fidelity 3D mesh generation—each calibrated against physical reference standards traceable to NIST-compatible dimensional metrology protocols. Its architecture supports longitudinal studies under controlled environmental conditions, making it suitable for breeding programs, stress-response screening, functional genomics validation, and ecophysiological modeling.
Key Features
- Tri-axial visible-light imaging array: Top-mounted, upper-lateral, and lower-lateral camera units synchronized with turntable rotation for complete surface coverage
- Automated adaptive imaging: Real-time size evaluation triggers optimal resolution selection, field-of-view adjustment, and image stitching for specimens exceeding standard capture volume
- Integrated environmental monitoring: Onboard temperature and relative humidity sensors log ambient conditions with timestamped metadata for environmental correlation analysis
- Dual-mode analysis pipeline: Independent 2D high-throughput module (≤60 s/sample) and background-queued 3D reconstruction engine operate in parallel without workflow interruption
- Motorized optical positioning: Software-controlled vertical translation of lenses and imaging chassis ensures optimal working distance and depth-of-field for variable plant architectures
- Full-spectrum color calibration: RGB, CIELAB (D65 illuminant, 2° observer), HSV, and grayscale channels captured with hardware-level white balance and gamma correction
- Cloud-enabled data synchronization: Secure device-bound upload of processed datasets—including 3D model files, parameter tables, and time-series reports—to encrypted cloud repositories
- Comprehensive safety architecture: Mechanical limit switches, infrared proximity sensors, and door interlocks comply with IEC 61000-6-2/6-4 electromagnetic compatibility and Class I electrical safety standards
Sample Compatibility & Compliance
The IN~Pheno50 accommodates herbaceous and semi-woody specimens from major angiosperm families including Poaceae, Solanaceae, Brassicaceae, and Fabaceae. Specimens must fit within the defined height (300–1000 mm), width (100–500 mm), and fresh mass (100–75,000 g) envelopes. The system’s non-contact methodology eliminates mechanical perturbation, preserving physiological integrity during repeated measurements. All raw image data, derived parameters, and processing logs are stored with immutable timestamps and user identifiers, supporting adherence to Good Laboratory Practice (GLP) documentation requirements. Export formats include CSV (trait tables), OBJ/GLTF (3D meshes), MP4 (rotational model videos), and JSON (structured metadata), facilitating integration into LIMS environments and regulatory submissions aligned with OECD TG 150 or ISO 20679-1 frameworks.
Software & Data Management
The proprietary IN~Pheno Suite v4.x runs natively on Windows 10/11 platforms with GPU-accelerated rendering enabled via CUDA-compatible NVIDIA drivers. It features dual-language UI (English/Chinese) with one-click switching, configurable export paths for analysis results, 3D models, and video assets, and hierarchical project management with versioned dataset archiving. Each measurement session generates a self-contained package containing original images, calibrated color profiles, segmentation masks, skeletal graphs, and statistical summaries. Audit trails record all user actions—including parameter edits, manual reprocessing, and export events—with SHA-256 hashing for integrity verification. Data exports support FDA 21 CFR Part 11-compliant electronic signatures when deployed with validated identity management infrastructure.
Applications
- Quantitative trait locus (QTL) mapping and genome-wide association studies (GWAS) requiring high-resolution structural descriptors
- Drought, salinity, and nutrient-stress phenotyping via temporal tracking of canopy expansion rate, leaf area index (LAI), and fractal dimension shifts
- Herbicide mode-of-action studies using texture-based biomarkers (e.g., ASM, SSIM, entropy) sensitive to early cellular damage
- Root-shoot allometry modeling through correlated analysis of aboveground morphology and gravimetric biomass
- Controlled-environment phenomics in growth chambers and phytotrons where reproducible lighting and microclimate control are critical
- Educational use in plant science curricula for teaching digital image analysis, geometric morphometrics, and spectral feature engineering
FAQ
What is the minimum and maximum plant size the IN~Pheno50 can accommodate?
Plants must be between 300 mm and 1000 mm in height and 100 mm to 500 mm in maximum width. Specimens outside this range require custom calibration or pre-processing steps.
Does the system support integration with third-party LIMS or breeding management software?
Yes—CSV, JSON, and XML export templates are configurable to match common schema requirements; API documentation is available under NDA for enterprise integrations.
Is the 3D reconstruction output compatible with common computational biology tools?
OBJ, PLY, and GLTF 2.0 formats are natively supported, enabling direct import into MorphoGraphX, PlantSeg, Blender, MATLAB, and Python-based morphometric pipelines.
How is measurement repeatability ensured across operators and sessions?
All optical paths are factory-aligned using laser interferometry; each session records calibration frame metadata, and software enforces standardized illumination intensity (measured in lux) and exposure duration.
Can the system operate unattended for overnight or multi-day time-series experiments?
Yes—the enclosure is thermally insulated and equipped with fail-safe power management; scheduled acquisition sequences persist through brief outages, and sensor-triggered alerts notify users of environmental excursions.