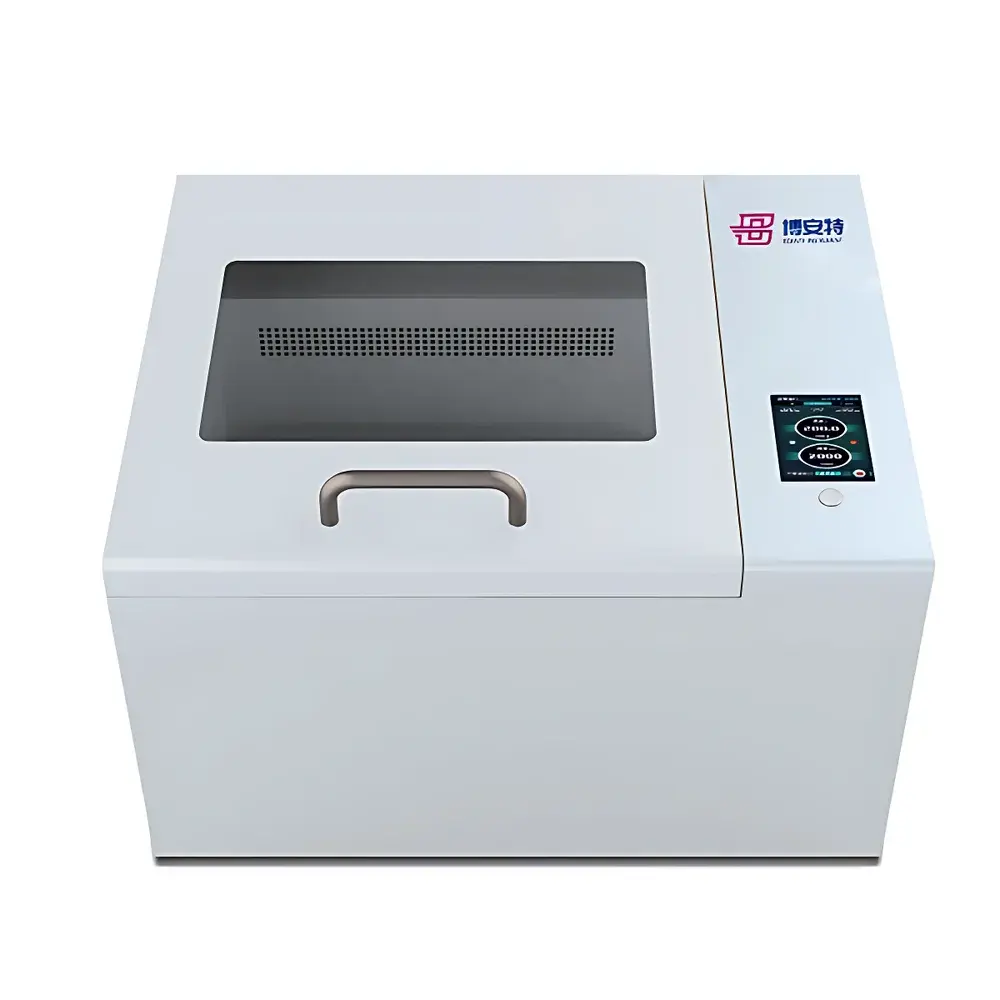
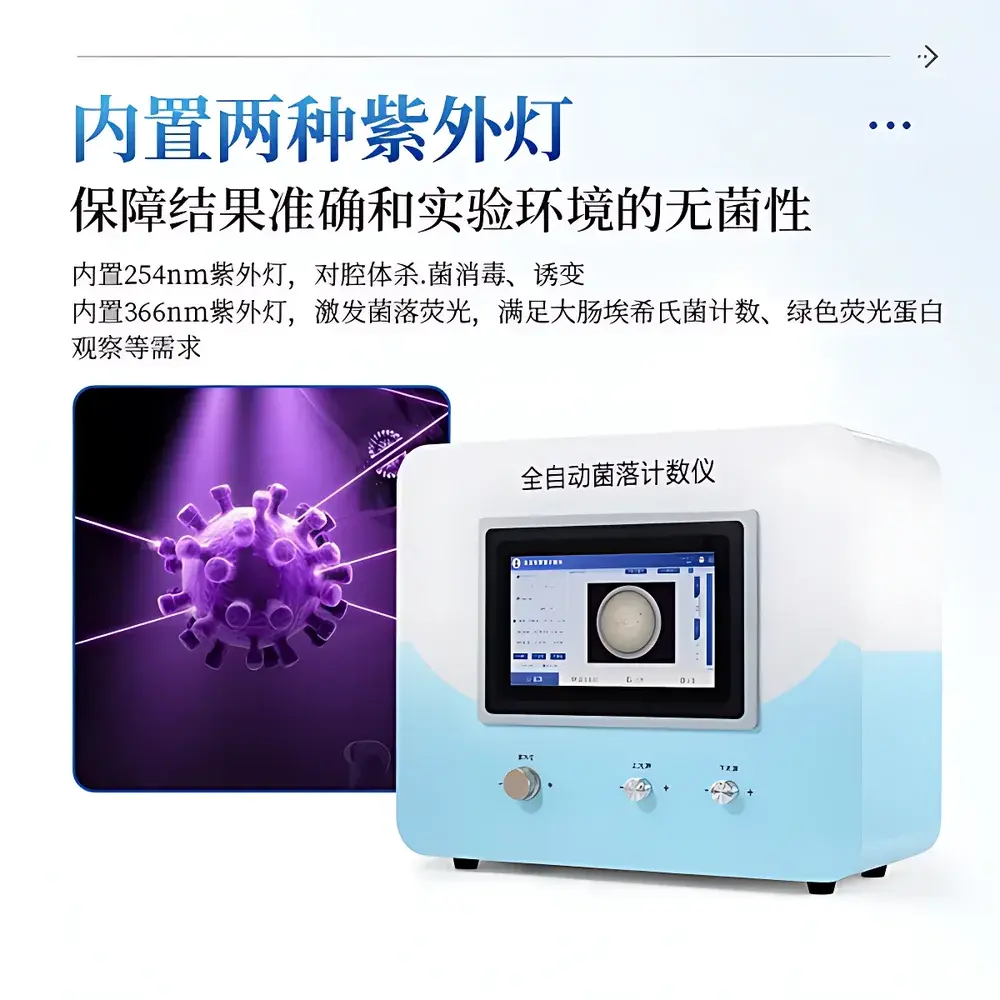
COMECAUSE IN-J100 Automated Colony Counter
| Brand | COMECAUSE |
|---|---|
| Origin | Shandong, China |
| Manufacturer Type | Direct Manufacturer |
| Instrument Type | Fully Automated Colony Counter |
| Sample Format | Petri Dish (90–110 mm diameter) |
| Counting Accuracy | ≤ ±10% |
| Imaging Resolution | 1920 × 1080 pixels |
| Key Software Functions | Threshold Adjustment, ROI Selection (Circle/Rectangle/Freehand), Automatic Calibration, Dual Illumination Control, PDF Report Export, Multi-User GLP-Compliant Data Management |
Overview
The COMECAUSE IN-J100 Automated Colony Counter is an image-based microbiological enumeration system engineered for precision, reproducibility, and regulatory compliance in routine and research-grade microbial analysis. It operates on the principle of digital machine vision combined with adaptive image segmentation algorithms—specifically optimized for detecting, distinguishing, and quantifying microbial colonies grown on standard agar-based Petri dishes (90–110 mm). Unlike manual counting or semi-automated systems, the IN-J100 integrates hardware-level optical control (dual independently adjustable top/bottom LED illumination) with real-time software processing to mitigate common artifacts—including uneven lighting, surface glare, colony overlap, background heterogeneity, and low-contrast morphology. Its core imaging engine employs a factory-calibrated 8-megapixel color CMOS sensor housed within a light-tight enclosure, enabling consistent capture under controlled conditions. The system automatically performs spatial calibration using embedded optical parameters (lens focal length, working distance, sensor pixel pitch), allowing accurate size measurement (down to 0.5 mm colony diameter) without user intervention—critical for ISO 4833-1:2013, FDA BAM Chapter 3, and USP compliant workflows.
Key Features
- Fully integrated standalone operation: 15.6-inch capacitive touchscreen (1920 × 1080), Windows 10 Pro 64-bit OS, 256 GB SSD storage—no external PC required
- Dual independent illumination system: Top and bottom LED sources individually switchable and continuously dimmable via rotary controls; supports “top-only”, “bottom-only”, or “dual-mode” configurations for optimal contrast across diverse media (e.g., chromogenic, blood, MacConkey, or nutrient agars)
- Automated optical calibration: Pre-loaded camera/lens parameters enable automatic pixel-to-millimeter conversion; eliminates need for daily recalibration while maintaining traceable dimensional accuracy
- Intelligent counting modes: “Dispersed Colony Mode” for well-isolated colonies; “Clumped Colony Mode” leveraging edge-preserving segmentation and watershed splitting for partially fused colonies
- Interactive correction suite: Adjustable intensity threshold, circular/rectangular ROI selection tools, freehand drawing for manual inclusion/exclusion, and real-time preview of segmentation boundaries
- GLP-aligned data governance: Role-based multi-user login, audit-trail enabled session logging, immutable raw image archiving, and tamper-evident PDF report generation with embedded metadata (date/time, operator ID, dish ID, calibration status)
Sample Compatibility & Compliance
The IN-J100 accepts standard polystyrene or glass Petri dishes (90 mm, 100 mm, and 110 mm diameters) with transparent or translucent bases. It accommodates both non-selective (e.g., TSA, PCA) and selective/differential media (e.g., VRBA, XLD, CHROMagar), including those exhibiting high background complexity or pigment diffusion. The system has been validated per ISO/IEC 17025:2017 requirements for method verification in accredited microbiology laboratories. Its counting performance meets the repeatability criteria specified in ISO 4833-1:2013 (≤15% relative standard deviation for duplicate plates) and aligns with FDA’s Bacteriological Analytical Manual (BAM) Chapter 3 acceptance thresholds for colony enumeration. All software functions—including user authentication, electronic signatures, and data export—are designed to support 21 CFR Part 11 readiness when deployed in GMP-regulated environments.
Software & Data Management
The proprietary IN-J100 Analysis Suite provides a unified interface for acquisition, processing, review, and reporting. Each user account maintains isolated project folders with encrypted access controls. Raw images are stored with EXIF metadata (exposure time, white balance, lens settings) and linked to processed count results. Statistical outputs include total CFU, average colony size distribution (histogram), spatial density mapping (heatmaps), and coefficient of variation across replicates. Data exports support CSV (for LIMS integration), PNG/JPEG (annotated images), and password-protected PDF reports containing instrument ID, calibration certificate reference, operator signature field, and timestamped audit trail summary. Optional API connectivity enables bidirectional communication with major LIS/LIMS platforms (e.g., Thermo Fisher SampleManager, LabVantage) via HL7 or RESTful endpoints.
Applications
- Food & beverage quality control: Enumeration of total viable counts (TVC), coliforms, and pathogens (e.g., E. coli, Salmonella) in raw materials, finished products, and environmental swabs
- Clinical microbiology: Quantitative urine culture analysis (CFU/mL), wound swab enumeration, and antimicrobial susceptibility testing plate assessment
- Pharmaceutical microbiology: Bioburden testing of non-sterile products per USP , water system monitoring (PW/WFI), and cleanroom environmental monitoring (EM) data aggregation
- Environmental testing: Heterotrophic plate count (HPC) in drinking water, wastewater effluent, and recreational water per EPA Method 1603/1681
- Academic & R&D labs: High-throughput screening of antimicrobial compounds, biofilm dispersion assays, and microbial community structure analysis via colony morphotype clustering
FAQ
Does the IN-J100 require daily recalibration?
No. The system performs automatic spatial calibration at startup using embedded optical constants. Recalibration is only necessary after physical lens replacement or significant mechanical impact.
Can it distinguish between viable colonies and debris or scratches on the agar surface?
Yes. The algorithm applies multi-parameter filtering—including circularity, solidity, intensity gradient, and local contrast—to suppress non-biological artifacts while preserving morphologically atypical but biologically relevant colonies.
Is the software compatible with networked laboratory information management systems (LIMS)?
Yes. The IN-J100 supports configurable data export formats (CSV, XML) and offers optional HL7 v2.x and REST API modules for seamless integration into existing LIMS infrastructure.
What regulatory standards does the system support out-of-the-box?
It natively supports documentation and traceability requirements for ISO/IEC 17025, ISO 4833-1, USP /, FDA BAM, and EU Pharmacopoeia 2.6.12. Audit trail logs comply with ALCOA+ principles for GxP environments.
How is data integrity ensured during long-term archival?
Raw images are written in lossless TIFF format with embedded SHA-256 checksums. Processed results are digitally signed upon PDF export, and all user actions are timestamped and logged in an immutable SQLite database with write-once semantics.