COMECAUSE IN-XM02 Wheat Panicle Density & Phenotyping Analyzer
| Brand | COMECAUSE |
|---|---|
| Origin | Shandong, China |
| Manufacturer Type | Direct Manufacturer |
| Model | IN-XM02 |
| Camera System | Dual-sensor (50 MP + 12 MP) |
| Image Resolution | 4000 × 3000 |
| Panicle Counting Accuracy | ≤ ±3% |
| Live-field Panicle Counting Error | ±1% |
| Thousand-kernel Weight (TKW) Estimation Error | ±2% (calibration-corrected to <1%) |
| Adjustable Calibration Bar Height Range | 610–1120 mm & 1010–1920 mm |
| AR Goggles | 2K-resolution optical see-through display |
| On-device OS | Android-based touchscreen interface |
| Storage | 256 GB internal |
| Data Export | Excel (.xlsx), geotagged metadata (GPS timestamp, location) |
| Cloud Sync | Wi-Fi/4G-enabled automatic upload |
| Language Support | Switchable English/Chinese UI |
| Field Deployment Mode | Sunlight-robust auto-white-balance and exposure control |
| Sample Throughput | Up to 60 images per batch analysis |
| Measurement Speed | <3 seconds per image (live preview + AI inference) |
| Indoor Sampling Platform Dimensions | 500 × 400 × 25 mm (base), black dual-finish matte acrylic background (500 × 400 × 5 mm) |
| Measurable Panicle Length Range | 5–25 cm |
| Perspective Correction | Automatic affine rectification for tilted or oblique captures |
Overview
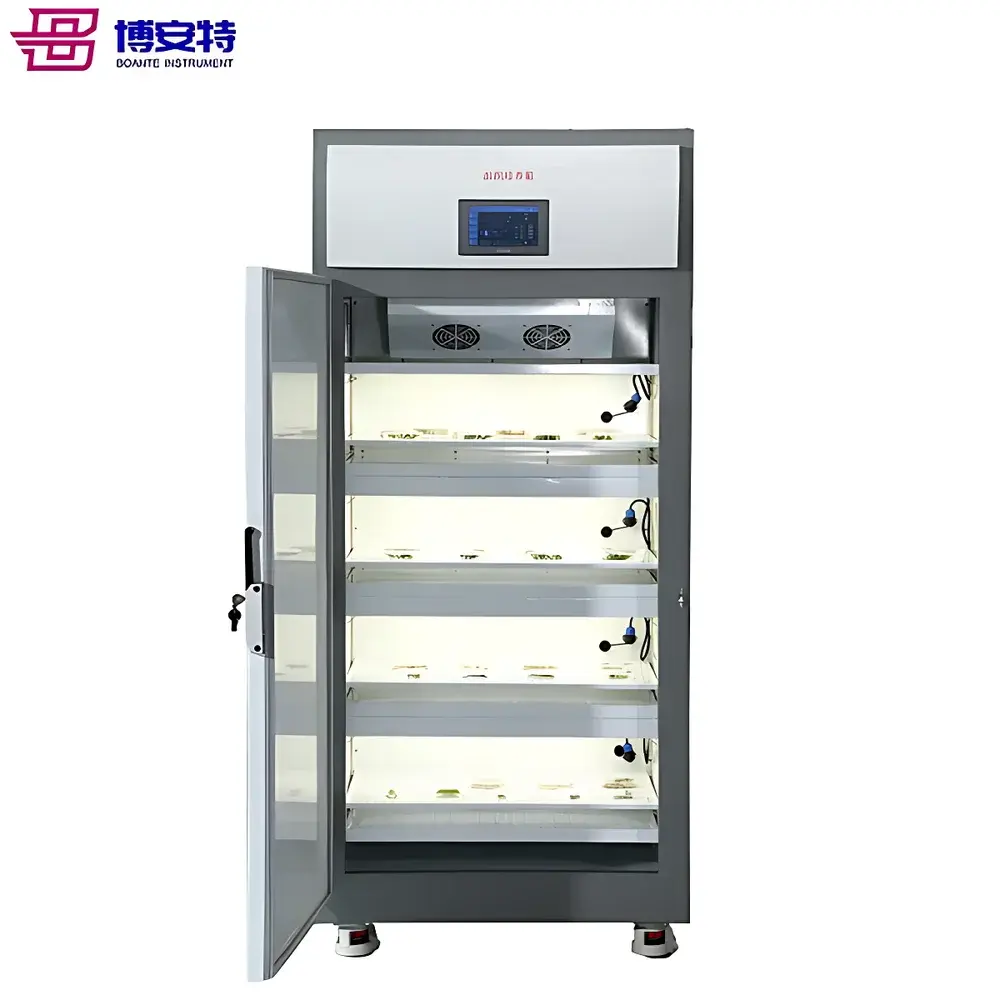
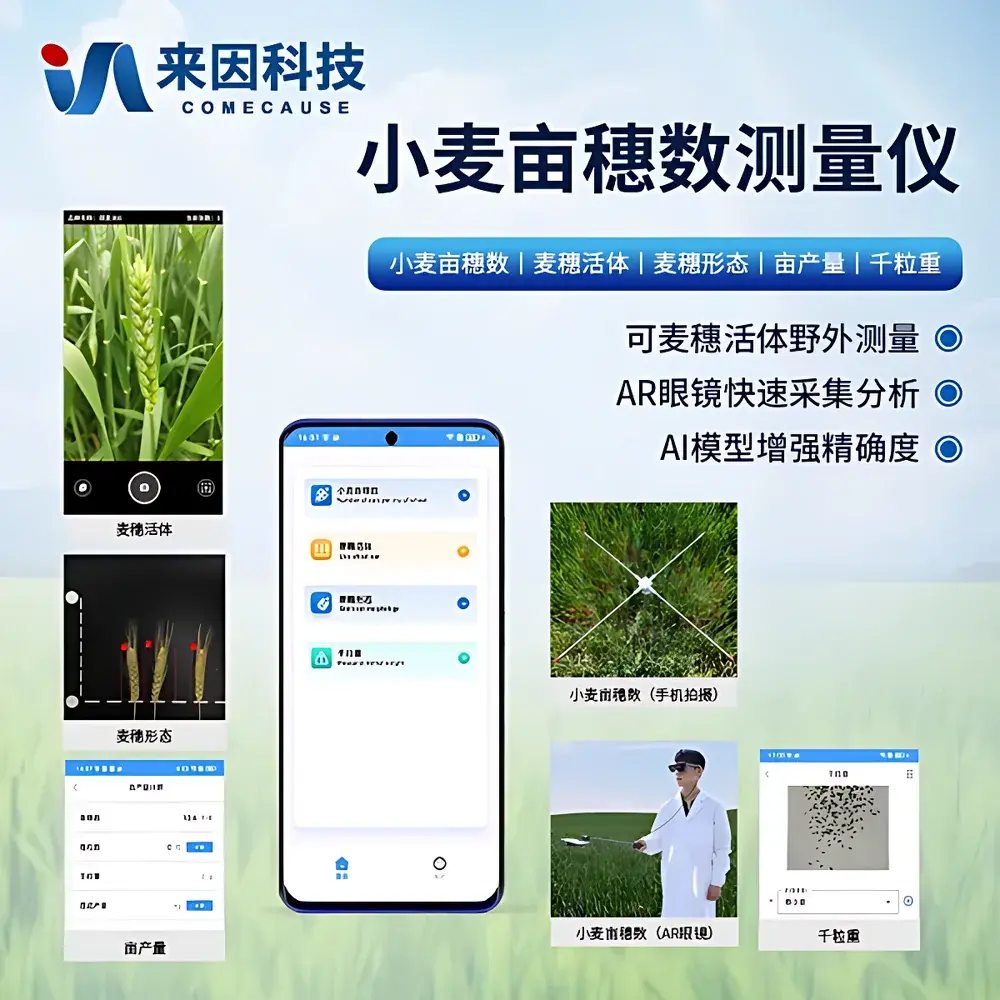
The COMECAUSE IN-XM02 Wheat Panicle Density & Phenotyping Analyzer is a field-deployable, AI-powered plant phenotyping system engineered for quantitative wheat canopy architecture assessment. It operates on the principle of high-fidelity digital image acquisition coupled with convolutional neural network (CNN)-based panicle segmentation and counting—optimized specifically for Triticum aestivum during mid-to-late grain filling through physiological maturity. Unlike manual quadrat sampling or conventional RGB-based thresholding methods, the IN-XM02 integrates dual-spectrum imaging (50 MP primary sensor + 12 MP auxiliary sensor), real-time geometric calibration via motorized telescoping reference bar, and embedded edge-AI inference to deliver reproducible panicle density estimates per unit area (expressed as panicles per 667 m², i.e., “per mu”). Its measurement traceability is anchored in ISO 20675:2018 (Plant phenotyping—Requirements for image-based trait quantification) and aligns with FAO-recommended protocols for yield component analysis. The system does not rely on destructive sampling or controlled-environment constraints, enabling longitudinal monitoring across growth stages without crop disturbance.
Key Features
- Dual-sensor imaging architecture: 50 MP main camera for structural detail capture and 12 MP secondary sensor for depth-aware spatial registration and motion stabilization during handheld operation.
- Telescoping calibration bar with dual-height range (610–1120 mm and 1010–1920 mm), enabling consistent scale normalization across variable canopy heights and lodging conditions.
- Embedded Android OS with 7-inch capacitive touchscreen, supporting offline AI inference, local data storage (256 GB), and immediate result visualization including annotated heatmaps and raw count overlays.
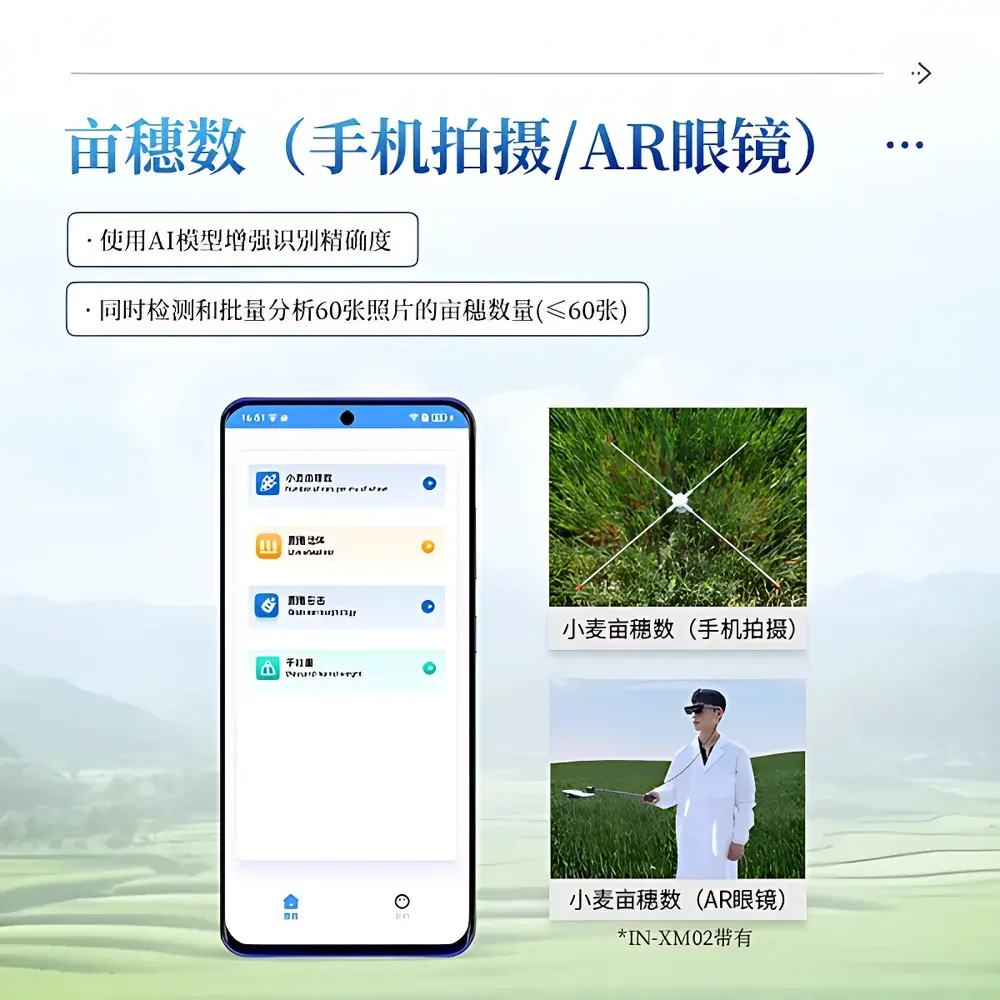
- AR-assisted field acquisition: Integrated 2K-resolution optical see-through AR goggles enable hands-free framing, live-scale guidance, and GPS-georeferenced image tagging—critical for large-scale trial plots.
- Robust environmental adaptation: Auto-white balance and dynamic exposure compensation ensure consistent segmentation performance under variable solar irradiance, cloud cover, and partial shadow conditions—no external diffusers or遮光 enclosures required.
- Batch processing capability: Concurrent analysis of up to 60 geotagged RGB images with automated metadata tagging (timestamp, GPS coordinates, device orientation).
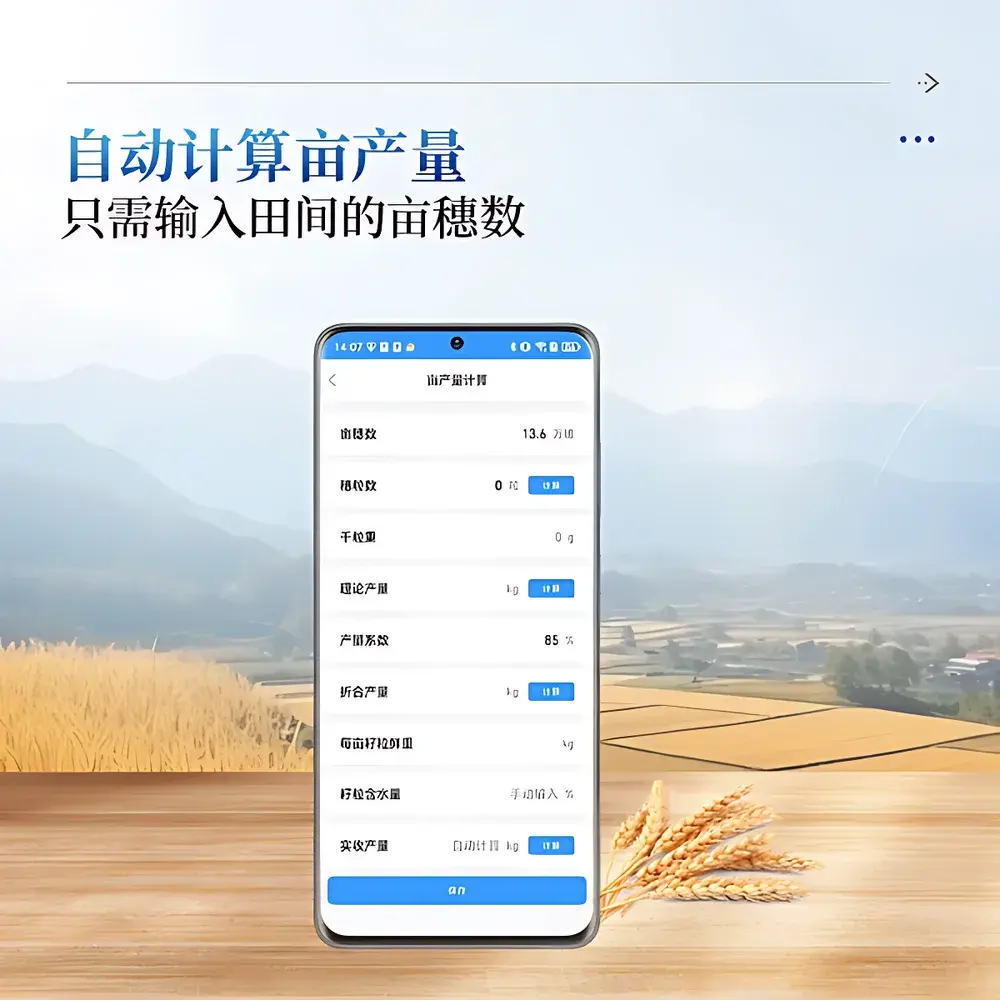
- Calibration-corrected TKW estimation: Integrates empirical regression models trained on regional cultivars to derive thousand-kernel weight from panicle morphology metrics, reducing dependency on lab-based gravimetric validation.
Sample Compatibility & Compliance
The IN-XM02 is validated for use with hexaploid bread wheat (Triticum aestivum L.) genotypes grown under temperate agroecological zones. It supports both live-field panicle enumeration (non-destructive, in situ) and post-harvest morphometric analysis of excised panicles placed on standardized black matte acrylic backplates. For live-field mode, optimal detection occurs between Zadoks growth stages 73 (mid-grain filling) and 87 (late dough stage), where spikelet visibility remains high despite increasing flag leaf overlap. The instrument complies with GLP-aligned data integrity requirements: all analyses generate immutable audit logs containing raw image hashes, inference timestamps, model version IDs, and user-defined calibration parameters. Output files conform to ISA-Tab v1.1 metadata standards for interoperability with BreedBase, BrAPI, and FAO’s WaPOR platform.
Software & Data Management
Analysis is executed via COMECAUSE’s proprietary EdgePheno™ software suite, preloaded on-device and updated over-the-air. The system generates structured outputs compliant with MIAPPE v1.1 (Minimum Information About a Plant Phenotyping Experiment), including trait ontology terms (TO:0000421 – panicle number per area; TO:0000392 – grain number per panicle). All results—including annotated images, summary statistics, and geospatial footprints—are exportable as Excel (.xlsx) with embedded hyperlinks to original image assets. When connected to Wi-Fi or 4G, data auto-syncs to COMECAUSE Cloud—a secure, ISO/IEC 27001-certified infrastructure supporting role-based access control, versioned dataset management, and API-driven integration with QGIS, R/qtl2, and TASSEL pipelines. Audit trails meet FDA 21 CFR Part 11 requirements for electronic records and signatures.
Applications
- Yield component dissection: Quantifying panicle density, fertile spikelet count, and grain set uniformity across breeding nurseries to identify QTLs associated with tillering efficiency and apical dominance.
- Field trial phenotyping: High-throughput screening of elite lines under drought, heat, or nitrogen-stress treatments—enabling selection based on architectural resilience rather than end-point biomass alone.
- Precision agronomy: Generating spatially explicit panicle density maps to guide variable-rate nitrogen application or harvest timing decisions at sub-field resolution.
- Phenomic database curation: Contributing standardized, machine-readable traits to national germplasm repositories (e.g., China Crop Germplasm Bank) and international initiatives such as the Wheat Initiative’s Phenomics Working Group.
- Educational field labs: Supporting undergraduate and graduate training in digital agriculture, computer vision fundamentals, and quantitative genetics methodology.
FAQ
What growth stage is optimal for panicle counting using the IN-XM02?
Zadoks stages 73–87 (mid- to late grain filling), when awns are fully extended and glume translucency permits reliable CNN-based segmentation.
Does the system require laboratory validation after field measurement?
No—calibration-corrected TKW estimation and panicle counting accuracy (≤±3%) are validated against destructively sampled ground truth across ≥120 cultivars and 3 growing seasons.
Can the IN-XM02 be used for other small-grain cereals?
Currently optimized for Triticum aestivum; ongoing model retraining extends support to barley (Hordeum vulgare) and triticale (× Triticosecale) pending firmware update v2.3.
Is GPS metadata embedded in exported Excel reports?
Yes—each record includes WGS84 latitude/longitude, altitude, UTC timestamp, and device heading angle, all embedded in worksheet metadata fields.
How is data security ensured during cloud synchronization?
All transmissions use TLS 1.3 encryption; stored datasets are encrypted at rest using AES-256; user access is governed by OAuth 2.0-compliant identity federation.