COMECAUSE IN~Pheno200 High-Spectral Plant Phenotyping Imaging System
| Brand | COMECAUSE |
|---|---|
| Origin | Shandong, China |
| Manufacturer Type | Original Equipment Manufacturer (OEM) |
| Country of Origin | China |
| Model | IN~Pheno200 |
| Price | USD 136,000 (FOB Qingdao) |
| Plant Height Range | 300–1000 mm |
| Plant Width Range | 100–500 mm |
| Fresh Weight Capacity | 100–75,000 g |
| Hyperspectral Range | 400–1000 nm |
| Spectral Resolution | <2.5 nm |
| Spectral Bands | 1200 |
| Light Source | Low-flicker, high-CRI halogen illumination |
| Enclosure Dimensions (L×W×H) | 1500 × 830 × 2140 mm |
| Imaging Sensor Resolution | Up to 12 MP (visible), hyperspectral line-scan camera |
| Acquisition Time per Plant | 40 s |
| 3D Reconstruction Time per Plant | ≤30 s |
| Software Platform | Windows 10/11, NVIDIA GPU ≥8 GB VRAM, Intel Core i5 or higher, RAM ≥8 GB |
Overview
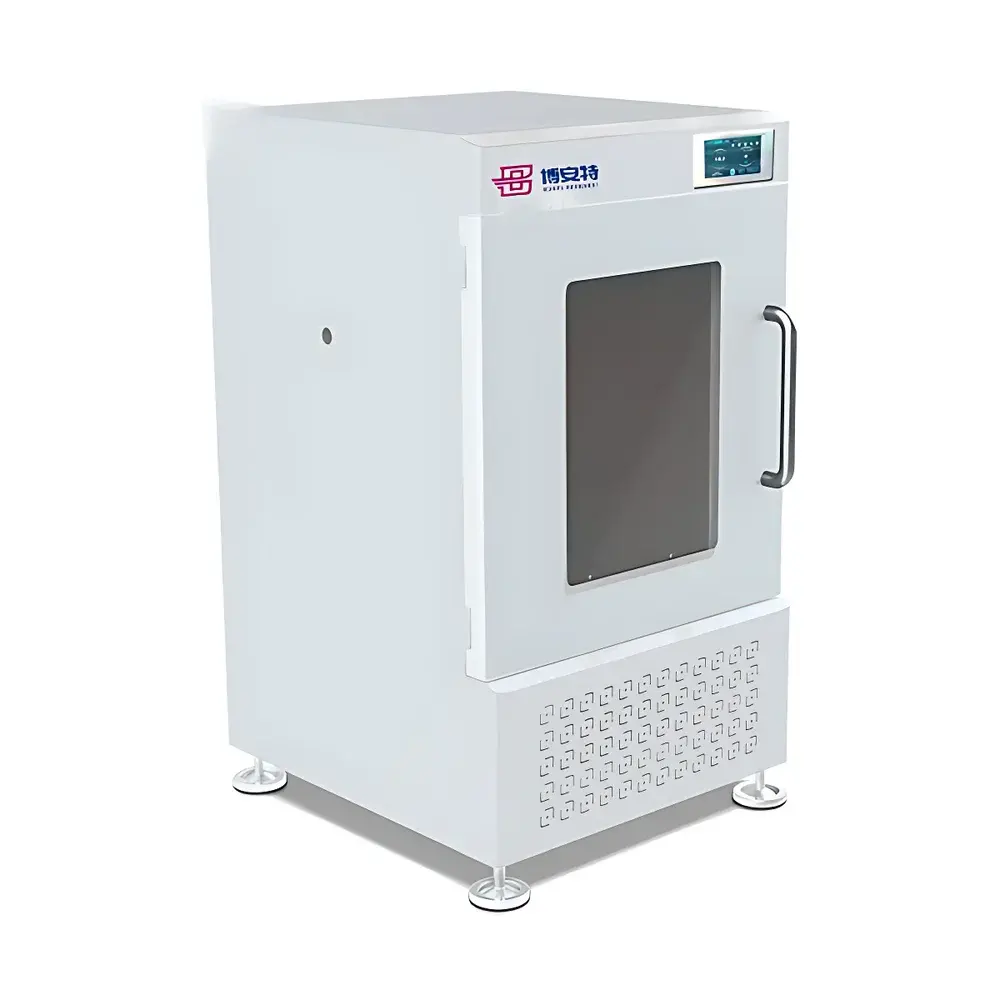
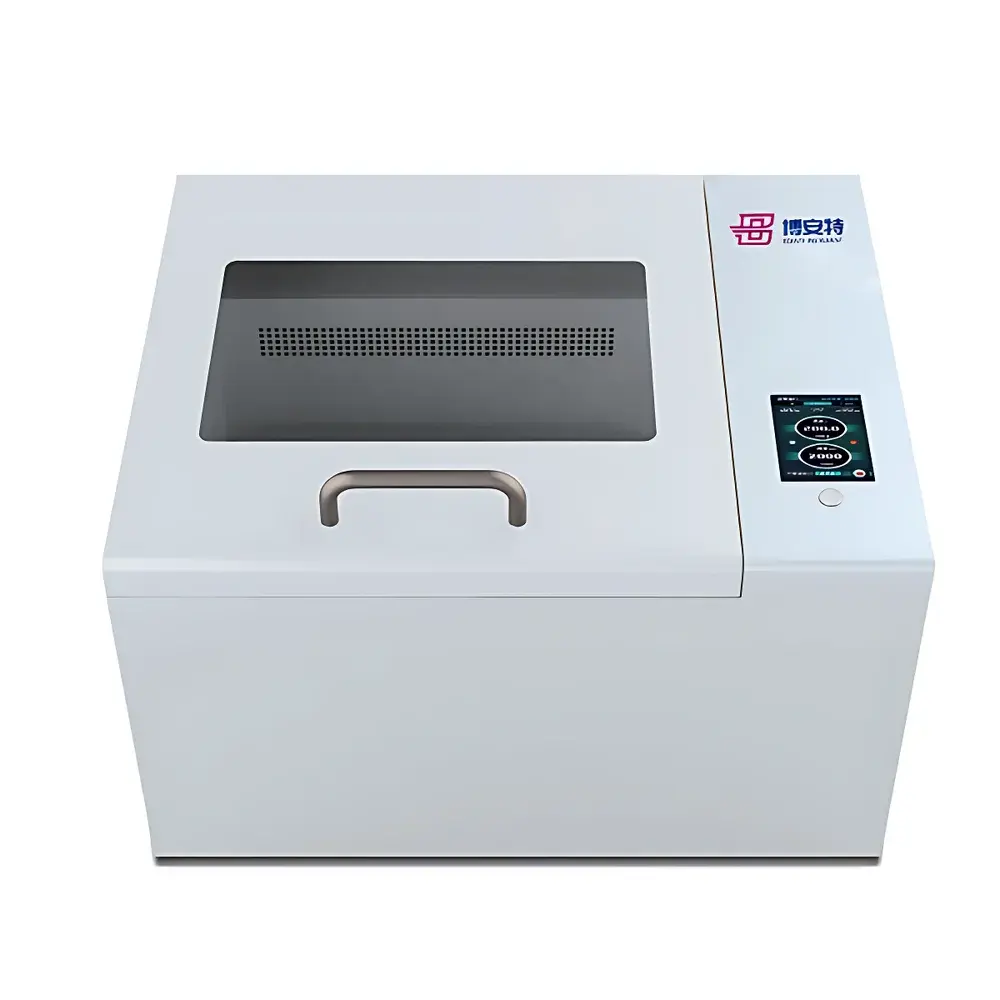
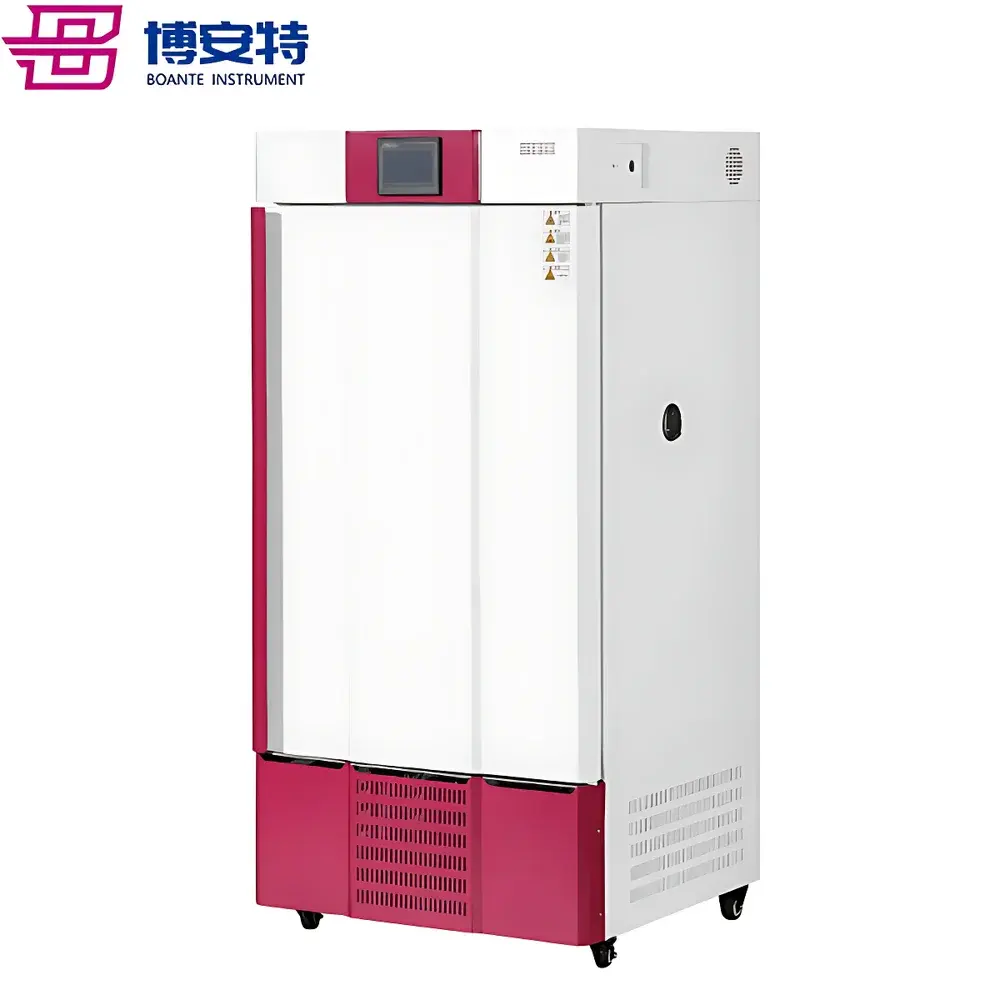
The COMECAUSE IN~Pheno200 High-Spectral Plant Phenotyping Imaging System is a fully integrated, multi-modal phenotyping platform engineered for non-destructive, quantitative assessment of plant structural, optical, and biochemical traits under controlled environmental conditions. It combines synchronized visible-light 2D/3D imaging with push-broom hyperspectral acquisition (400–1000 nm, <2.5 nm resolution, 1200 bands) to capture spatially registered morphological, chromatic, textural, and spectral signatures across the entire plant canopy. The system employs structured multi-angle imaging—top-view, side-top, and dual side-bottom configurations—coupled with a motorized rotating turntable and vertically adjustable optics to acquire full 360° volumetric data. Its AI-accelerated 3D reconstruction pipeline uses photogrammetric and stereo-vision algorithms to generate geometrically accurate, texture-mapped point clouds and mesh models, enabling true 3D parameterization (e.g., canopy volume, branch angle distribution, surface curvature) without reliance on depth sensors or laser scanning. Designed for reproducible trait extraction in breeding, stress physiology, and functional genomics studies, the IN~Pheno200 operates as a GLP-aligned instrument—supporting audit trails, user access control, and metadata-embedded data export compliant with FAIR principles.
Key Features
- Multi-modal imaging architecture: Co-registered visible RGB (12 MP), Lab-color space (D65/2°), HSV, and grayscale channels alongside calibrated hyperspectral reflectance data
- Automated adaptive imaging: Real-time plant dimension estimation triggers optimal camera positioning, exposure, and image stitching for specimens exceeding field-of-view limits
- Integrated environmental monitoring: Onboard temperature and humidity sensors log chamber conditions synchronously with image acquisition
- Motorized optical positioning: Software-controlled vertical translation of lens assemblies and gantry to maintain consistent working distance across variable plant sizes
- High-throughput 2D analysis module: Processes single-plant images in ≤60 seconds, generating standardized reports for 42 morphological, 12 colorimetric, and 14 textural parameters
- Parallel 2D/3D workflow: 3D modeling runs asynchronously in background queue—enabling continuous 2D screening while high-fidelity 3D reconstructions are generated offline
- Cloud-enabled data management: Encrypted upload of analysis results (JSON, CSV, SPE, OBJ/GLB) via device-unique ID; role-based access and versioned storage
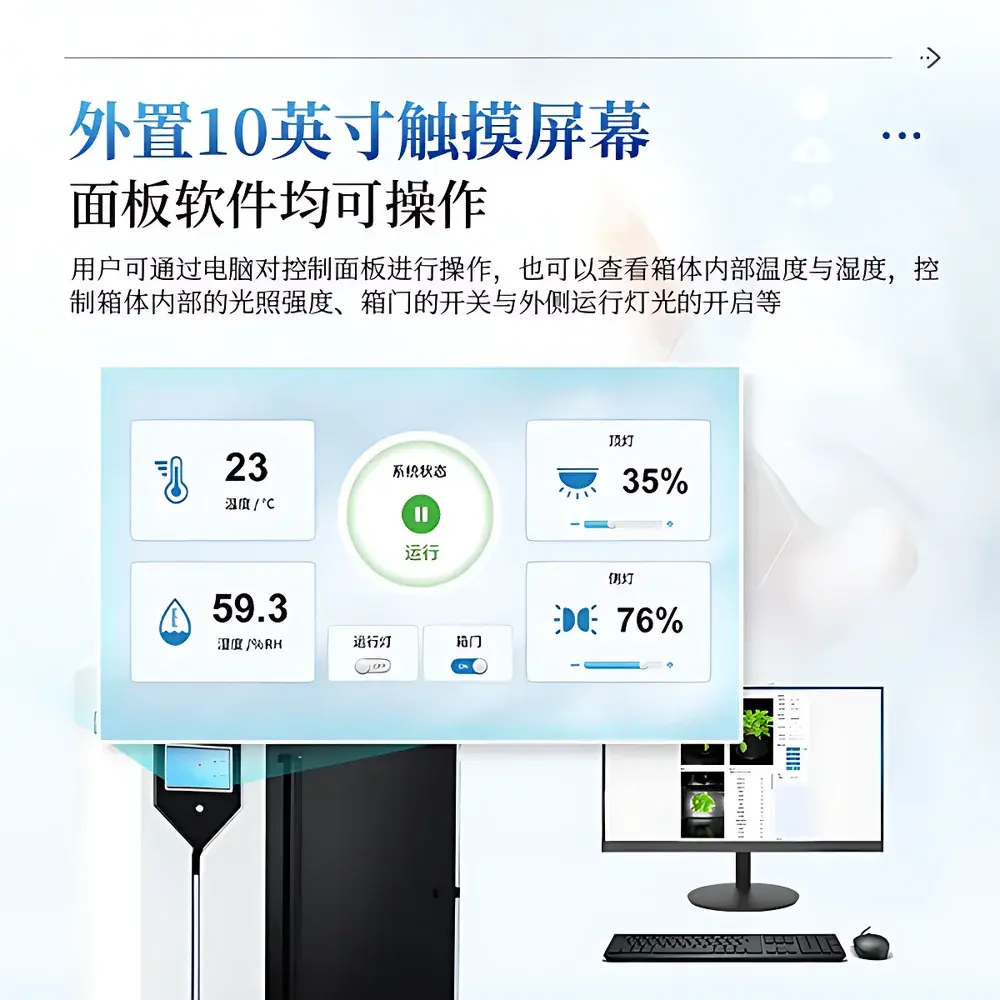
- Full hardware-software integration: 10-inch industrial touchscreen for local control (light intensity, door actuation, ambient lighting); remote operation via dedicated Windows client
- Comprehensive calibration traceability: Factory-calibrated hyperspectral sensor with NIST-traceable reference panels; geometric distortion correction applied in real time
- Mechanical safety architecture: Limit switches, IR proximity sensors, and emergency stop circuitry conforming to IEC 61000-6-2/6-4 electromagnetic compatibility standards
Sample Compatibility & Compliance
The IN~Pheno200 accommodates intact, living plants within height (300–1000 mm) and width (100–500 mm) envelopes typical of model species including Oryza sativa, Solanum lycopersicum, Arabidopsis thaliana, and Glycine max. Its low-heat, high-CRI halogen illumination ensures minimal photobiological perturbation during repeated measurements. All optical components meet ISO 9022-3:2015 (environmental testing for optical instruments) and IEC 61326-1:2013 (EMC for laboratory equipment). Data acquisition workflows support 21 CFR Part 11-compliant electronic records when deployed with validated Windows domain authentication and audit-log configuration. The system’s metadata schema aligns with MIAPPE v1.1 (Minimum Information About a Plant Phenotyping Experiment) and supports direct ingestion into BreedBase and Gramene databases.
Software & Data Management
The proprietary IN~PhenoSuite software (v4.2+) provides a unified interface for instrument control, acquisition scheduling, and multi-layered analysis. It implements ISO/IEC 17025-aligned uncertainty propagation for derived indices (e.g., NDVI, PRI, REIP), with all calculations executed using open-source spectral libraries (PySpectral, scikit-image) and documented mathematical definitions per ASTM E2595-20 and ISO 19036:2020. Raw hyperspectral cubes (.spe) and processed results are stored with embedded EXIF-like metadata—including timestamp, illumination spectrum, sensor gain, and calibration coefficients. Export options include FAIR-compliant NetCDF4 files, tabular CSV with SI units, and interactive 3D web viewers (Three.js-based). Audit logs record user actions, parameter modifications, and software updates with SHA-256 hash verification. Language localization supports EN/zh-CN switching without restart.
Applications
The IN~Pheno200 serves as a core phenotyping tool in academic and industrial plant science laboratories conducting: drought and salinity stress response profiling via temporal NDVI/REIP trajectories; nitrogen-use efficiency quantification through DATT and chlorophyll-sensitive indices (TCARI, MTCI); senescence dynamics mapping using PSRI and PRI570; disease progression monitoring (e.g., RSRI for rice blast); fruit ripening staging via Maturity Index1/2 composites; and QTL mapping studies requiring high-reproducibility 3D architectural descriptors (branch angle variance, fractal dimension, skeletonized biomass distribution). Its capacity for longitudinal, non-invasive measurement makes it suitable for GLP-compliant regulatory submissions under OECD TG 111 and EFSA guidance on pesticide residue metabolism studies.
FAQ
What spectral calibration standards are used for hyperspectral data validation?
NIST-traceable white reference panels (Spectralon® 99% reflectance) and dark-current references are acquired before each session; spectral response curves are corrected using vendor-provided radiometric calibration coefficients.
Does the system support integration with external climate chambers or growth rooms?
Yes—the IN~Pheno200 enclosure features RS-485 and Ethernet ports for bidirectional communication with third-party environmental controllers (e.g., Conviron, Snijders), enabling synchronized light/temperature/humidity protocols.
How is 3D model accuracy verified?
Geometric fidelity is validated using certified gauge blocks and anthropomorphic phantoms; root-mean-square deviation (RMSD) from ground-truth dimensions remains ≤0.35 mm across the operational volume.
Can users implement custom vegetation indices or machine learning models?
The software SDK provides Python API access to raw spectral cubes and 3D meshes; users may deploy trained scikit-learn or PyTorch models via containerized inference modules with deterministic execution environments.
Is remote maintenance and software update supported?
Yes—secure TLS 1.3 remote desktop access (with prior authorization) enables diagnostics and patch deployment; all firmware updates undergo cryptographic signature verification prior to installation.