MCL Think Nano Nano-Cyte® Single-Molecule Imaging System
| Brand | MCL Think Nano |
|---|---|
| Origin | USA |
| Manufacturer Type | Authorized Distributor |
| Origin Category | Imported |
| Model | Nano-Cyte® |
| Pricing | Upon Request |
Overview
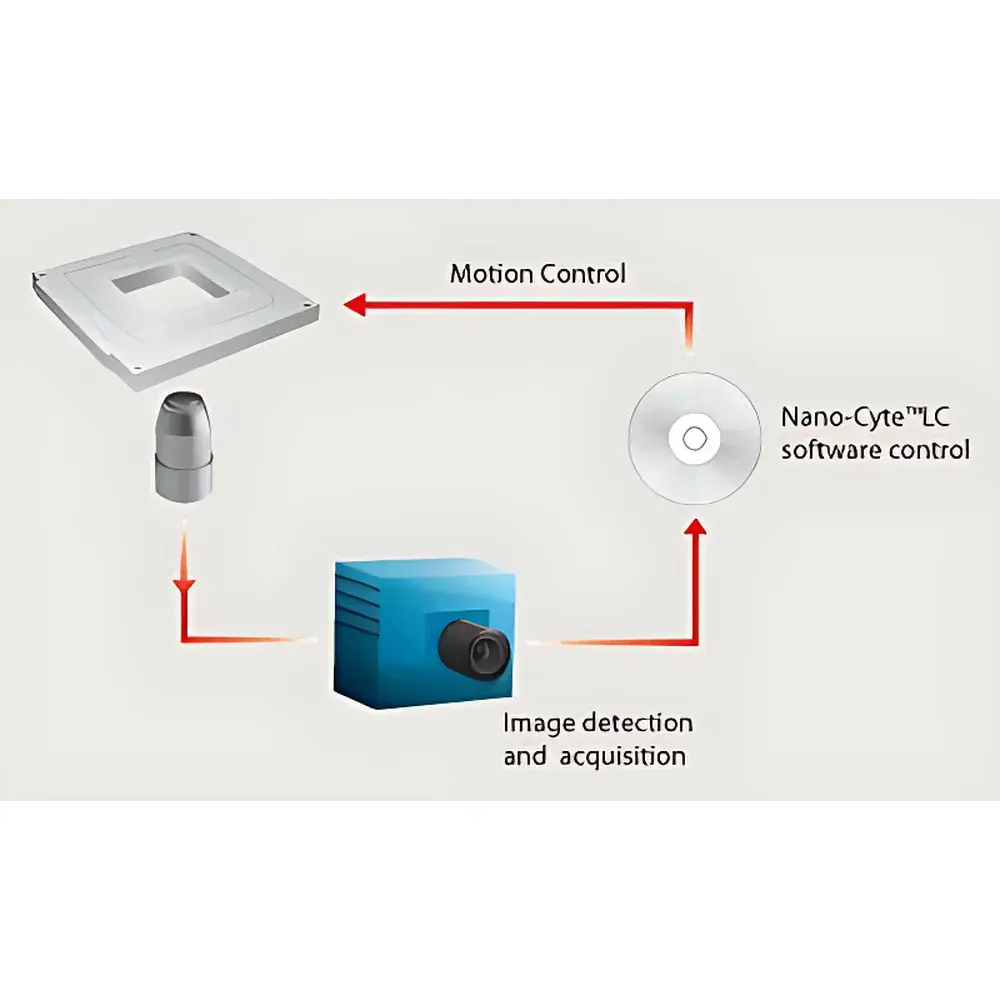
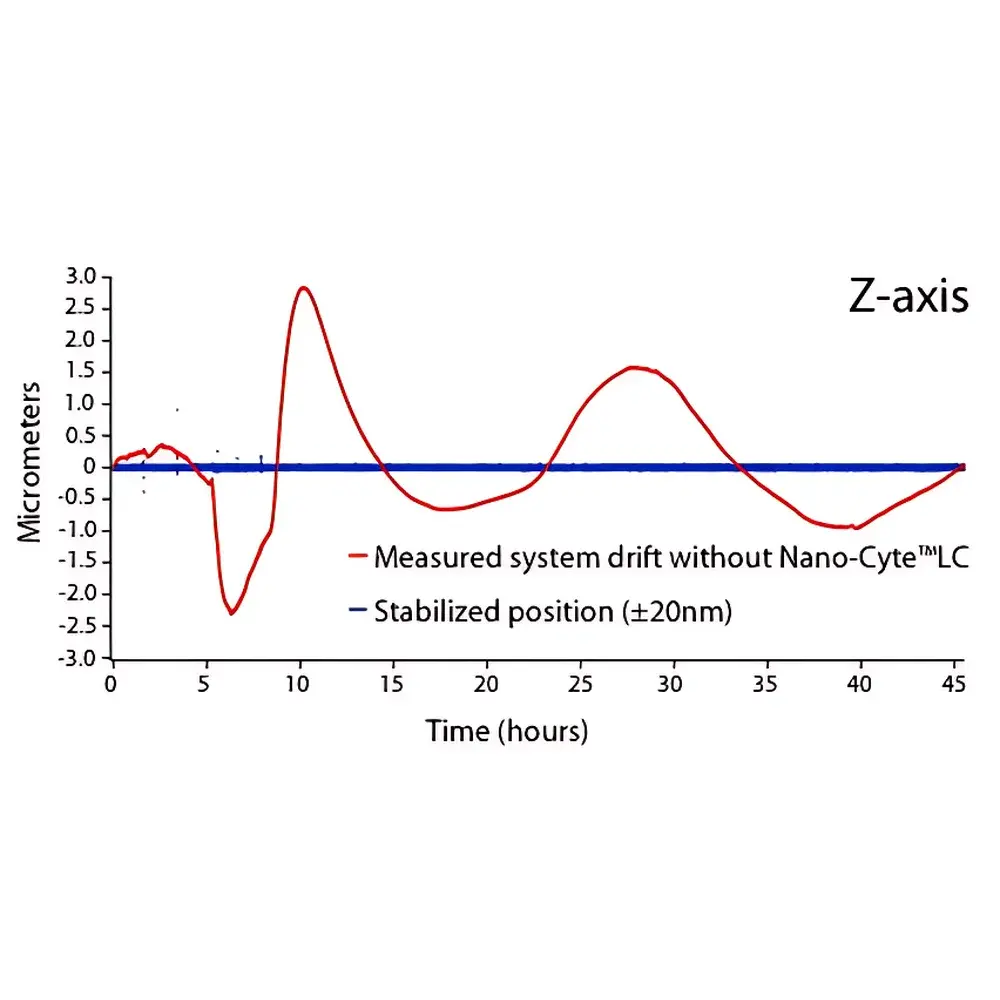
The MCL Think Nano Nano-Cyte® Single-Molecule Imaging System is a precision-engineered, closed-loop 3D active stabilization platform designed specifically for quantitative single-molecule fluorescence microscopy. It operates on the principle of real-time fiducial-based feedback control: sparse fluorescent reference emitters—embedded within or near the sample—are continuously imaged and localized in all three spatial dimensions (X, Y, Z) using sub-pixel centroid fitting algorithms. These 3D position measurements serve as error signals to drive a high-bandwidth nanopositioning system, which dynamically corrects for mechanical drift, thermal expansion, and stage instability at the nanometer level. Unlike passive drift compensation methods, Nano-Cyte® delivers true active positional control with metrological traceability, enabling extended-duration super-resolution imaging (e.g., PALM, STORM, DNA-PAINT), long-term tracking of molecular dynamics, and multi-color co-localization experiments where spatial fidelity over hours is non-negotiable.
Key Features
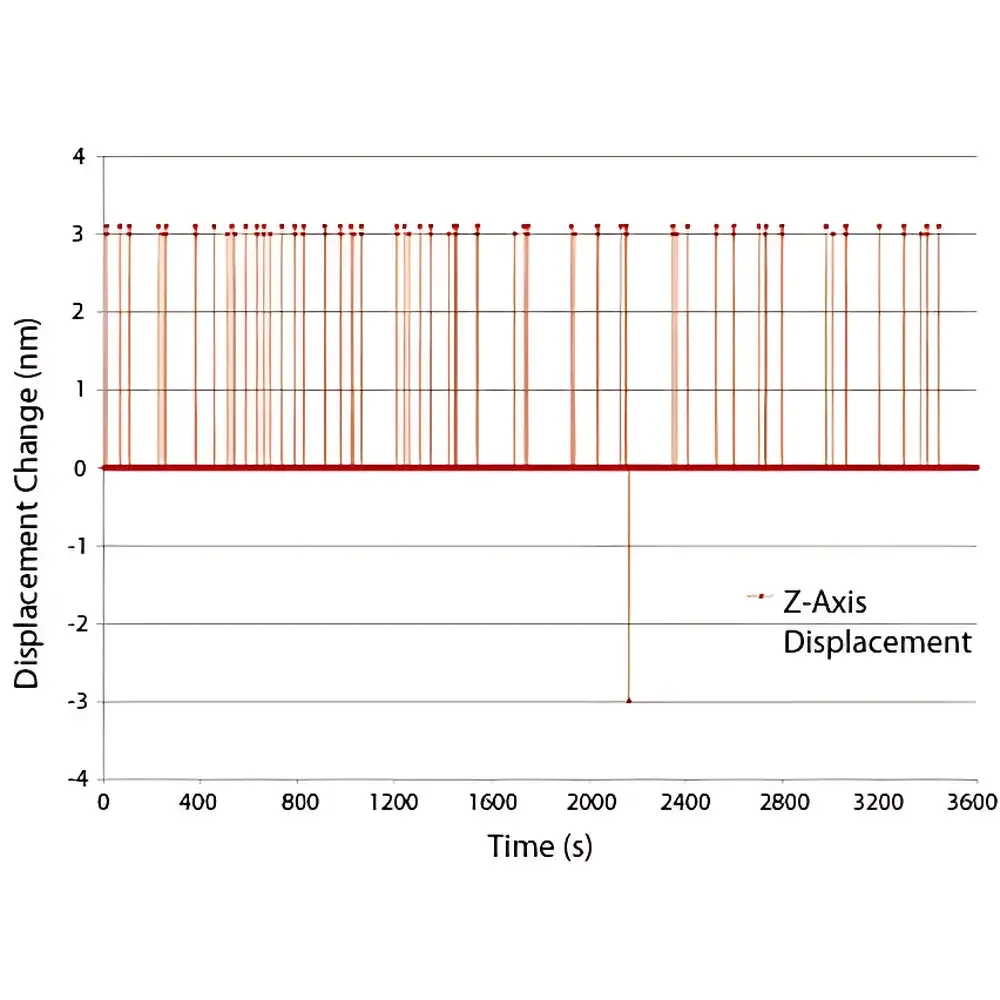
- Sub-3 nm RMS stability across X, Y, and Z axes over multi-hour acquisitions (US Patent 9,019,363)
- Real-time, simultaneous image acquisition and 3D stabilization—no trade-off between resolution and duration
- Integrated flexure-guided, piezo-driven triaxial nanopositioner with PicoQ® absolute position sensors and closed-loop ometer-level resolution
- Motorized 25 mm × 25 mm micropositioning stage for coarse sample navigation prior to stabilization engagement
- Drift compensation rate up to 10 nm/sec; typical stabilization latency ≤ 1 frame (dependent on camera frame rate)
- Native support for EMCCD cameras (Andor, Photometrics, Hamamatsu), TTL-synchronized shutters, and programmable light sources
- USB 2.0 controller interface with four configurable TTL outputs for hardware synchronization
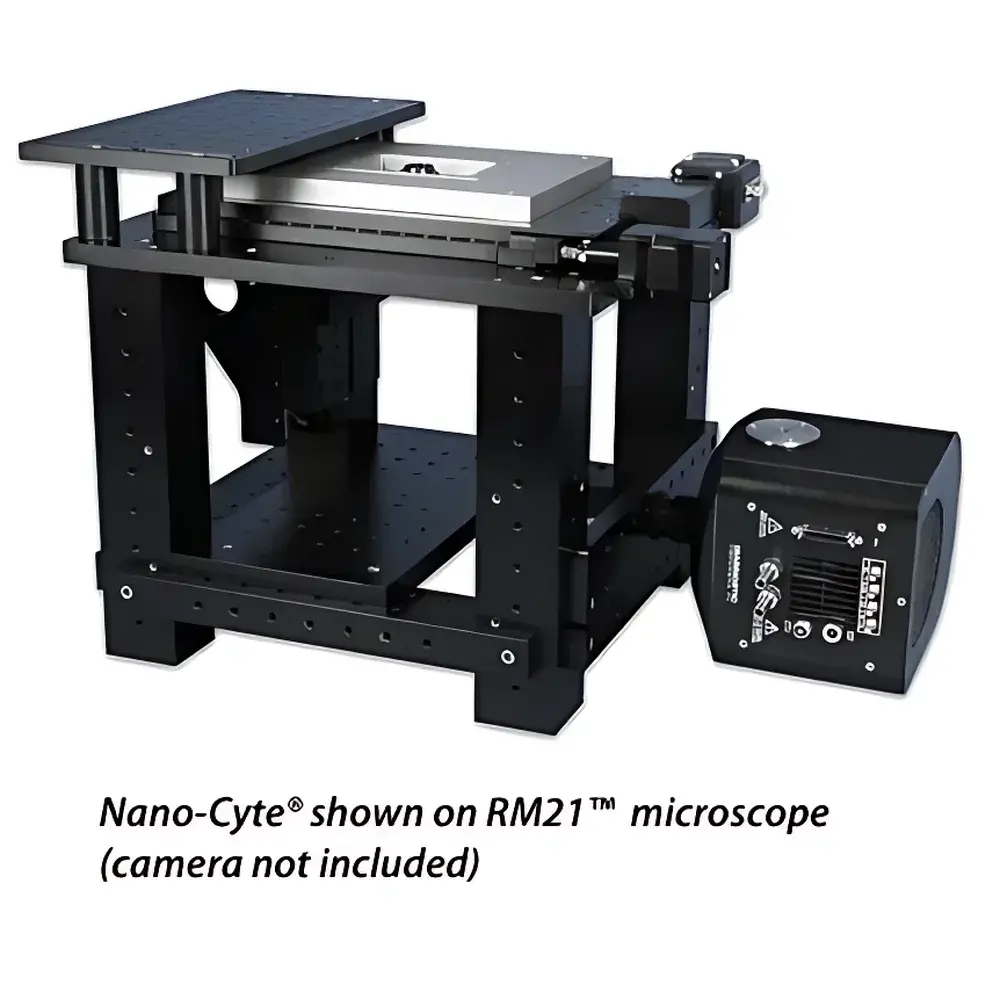
Sample Compatibility & Compliance
Nano-Cyte® is compatible with standard inverted optical microscopes including Nikon Ti/TE, Zeiss Axio Observer, Olympus IX series, and the Mad City Labs RM21™ open-platform microscope. It supports conventional glass-bottom dishes, coverslips (No. 1.5), and custom sample chambers used in TIRF, HILO, and light-sheet configurations. The system does not require modifications to existing microscope optics or illumination paths—stabilization is implemented entirely through sample-stage motion control. All firmware and software comply with GLP-aligned data integrity principles: raw TIFF images include embedded metadata (timestamp, stabilization status, reference count, RMS error), and full audit trails are maintained for position commands and calibration events. While not certified for clinical diagnostics, Nano-Cyte® meets engineering benchmarks referenced in ISO 10993-1 (biocompatibility context), ASTM E2841 (microscopy measurement uncertainty), and supports FDA 21 CFR Part 11 compliance when deployed with validated LabVIEW™ or μManager workflows.
Software & Data Management
The Nano-Cyte® native software provides six core functional modules: (1) real-time stabilization engine with adaptive reference selection logic; (2) synchronized image acquisition (TIFF output only); (3) device orchestration (nanopositioner, micropositioner, TTL peripherals); (4) particle localization analysis using Gaussian PSF fitting; (5) 3D point-cloud rendering with depth-encoded color mapping; and (6) multi-FOV trajectory stitching via fiducial-assisted field registration. Acquired datasets are stored as time-stamped, metadata-rich TIFF stacks—fully compatible with ImageJ/Fiji, rapidSTORM, and custom Python/Matlab pipelines. An exportable Windows DLL enables integration into third-party acquisition frameworks. Native interoperability with μManager and LabVIEW™ allows users to embed Nano-Cyte® control within larger experimental protocols—including automated focus maintenance (C-Focus™), multi-channel excitation sequencing, or closed-loop optogenetic stimulation paradigms.
Applications
- Single-particle tracking (SPT) of membrane proteins, cytoskeletal motors, or intracellular vesicles over >10,000 frames
- Multi-color super-resolution reconstruction requiring pixel-level registration across acquisition rounds
- Long-duration FRET or FLIM experiments where donor–acceptor distance stability must remain <5 nm
- Structural biology studies of biomolecular complexes immobilized on PEG-passivated surfaces
- Quantitative colocalization analysis across heterogeneous cell populations under physiological temperature gradients
- Calibration and validation of novel localization microscopy algorithms against ground-truth stabilized references
FAQ
What is the minimum required camera frame rate for effective stabilization?
Nano-Cyte® operates optimally at ≥10 fps for robust drift correction; however, stabilization remains functional down to 1 fps with reduced bandwidth. Frame rate dependency is governed by feedback loop latency—not camera specification alone.
Can Nano-Cyte® be used with upright microscopes?
No—it is engineered exclusively for inverted microscope configurations due to mechanical integration constraints and gravity-aligned nanopositioner orientation.
Does Nano-Cyte® require specialized fluorescent dyes or fiducials?
It uses standard photoactivatable or photoswitchable fluorophores (e.g., Alexa Fluor 647, Cy5, Janelia Fluor dyes) as fiducials; no proprietary labels are needed.
Is remote operation supported?
Yes—via USB 2.0 connection and network-shared LabVIEW™ or μManager instances; however, all real-time control loops execute locally on the host PC to ensure deterministic timing.
How is calibration performed and how often is it required?
Initial system calibration is performed during installation using a NIST-traceable step gauge. Recalibration is recommended every 6 months or after mechanical shock, and takes <15 minutes using the built-in automated routine.