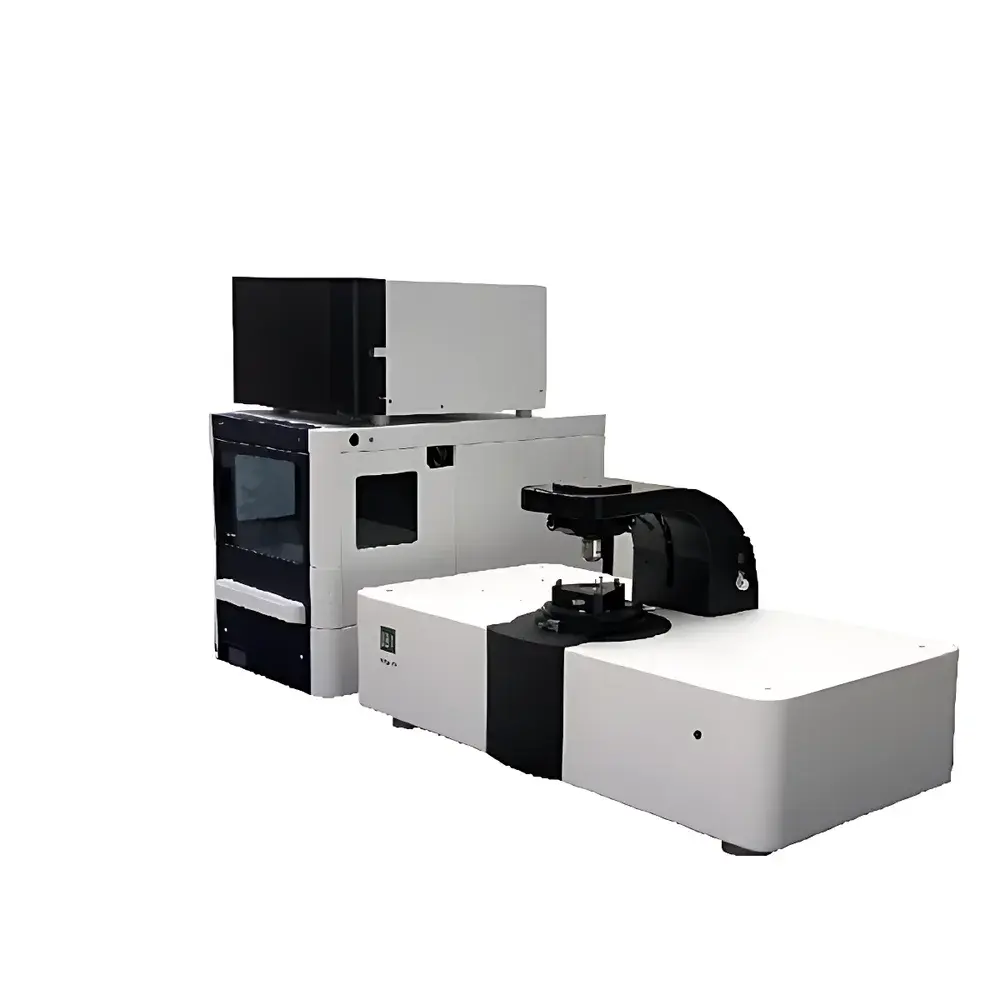
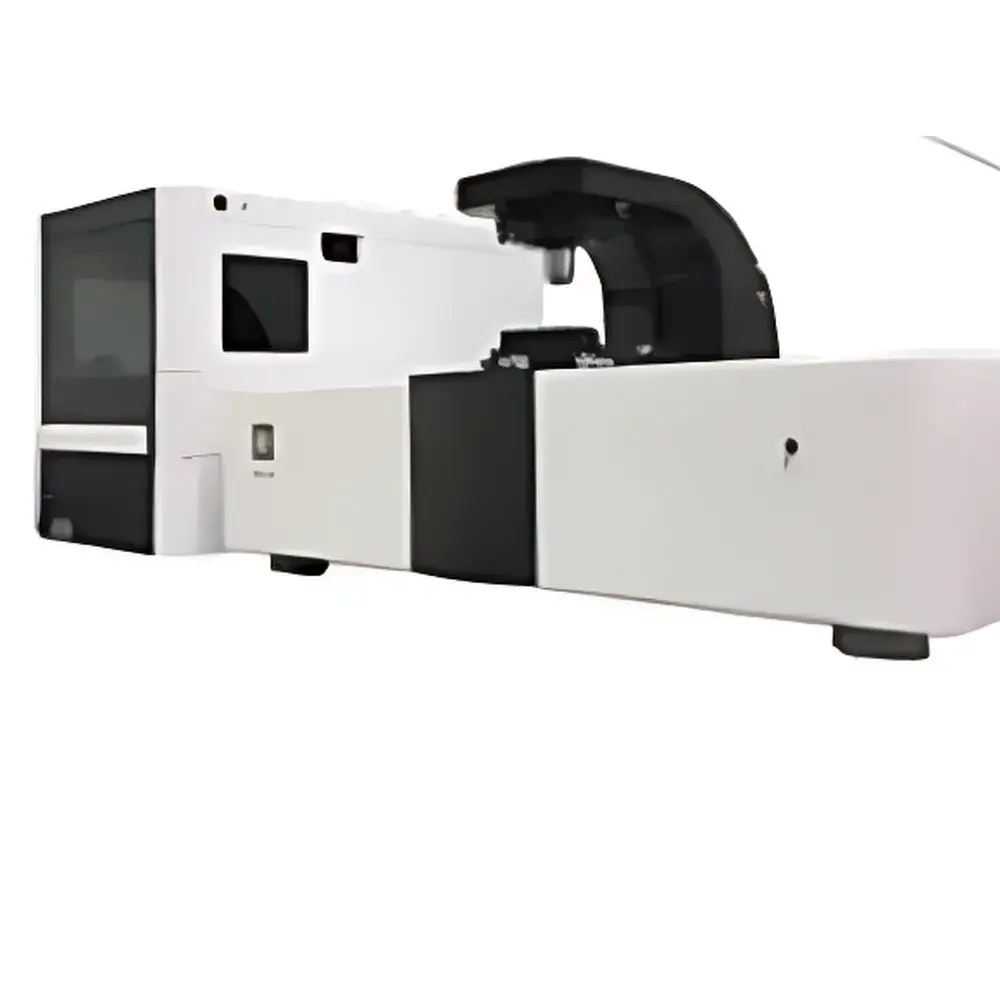
Biosensing Instrument SPRm 200 Surface Plasmon Resonance Microscope
| Brand | Biosensing Instrument |
|---|---|
| Origin | USA |
| Manufacturer Type | Authorized Distributor |
| Instrument Type | Surface Plasmon Resonance (SPR) |
| Model | SPRm 200 |
| Detection Range | 200 Da |
| Channel Configuration | 5 parallel microfluidic channels |
| Sample Throughput | 384 × 2 (dual-layer microarray format) |
| Injection Volume | 1–1500 µL (application-dependent) |
| Assay Duration | Millisecond to hour-scale (kinetically resolved) |
| Temperature Control Range | 15–40 °C |
| Detection Mode | Simultaneous SPR imaging + brightfield optical microscopy |
| Spatial Resolution | Pixel-level SPR signal mapping (< 2 µm pixel pitch) |
Overview
The Biosensing Instrument SPRm 200 Surface Plasmon Resonance Microscope is a high-resolution, label-free biosensing platform that integrates real-time surface plasmon resonance (SPR) detection with wide-field optical microscopy. Unlike conventional bulk SPR systems—which report averaged angular shifts across macroscopic sensing areas—the SPRm 200 performs quantitative, pixel-resolved SPR imaging at submicron spatial resolution. This capability enables direct visualization and kinetic quantification of molecular binding events occurring on native, unperturbed cellular membranes, intact bacteria, or immobilized nanomaterials—without fluorescent tags, enzymatic reporters, or protein purification. The instrument operates on the principle of evanescent field coupling: a p-polarized light beam incident at the Kretschmann angle excites surface plasmon polaritons (SPPs) at the gold-coated sensor chip interface. Binding-induced local refractive index changes perturb SPP propagation, generating intensity-modulated SPR contrast maps with single-pixel temporal resolution. This architecture supports true spatiotemporal mapping of heterogeneous binding landscapes across live-cell surfaces, making it uniquely suited for studying membrane receptor distribution, ligand-receptor stoichiometry, and nanoscale particle–surface interactions under physiologically relevant conditions.
Key Features
- Simultaneous dual-modality imaging: synchronized acquisition of brightfield optical images and quantitative SPR response maps at video rate (up to 30 fps)
- Pixel-level kinetic analysis: full time-resolved SPR sensorgrams extracted from user-defined regions of interest (ROIs), including single-cell subdomains or individual nanoparticles
- Integrated microfluidics: five independently addressable flow channels supporting multiplexed assays with programmable injection sequences and regeneration cycles
- High-sensitivity detection: sub-0.6 RU noise floor enabling measurement of low-molecular-weight analytes (≥200 Da) and weak-affinity interactions (KD down to 1 pM)
- Precise thermal regulation: Peltier-controlled stage maintaining temperature stability within ±0.1 °C across the 15–40 °C operating range
- Native-state compatibility: validated for adherent and suspension cells, whole bacteria (e.g., E. coli O157:H7), virus particles, liposomes, and functionalized nanoparticles (down to ~20 nm hydrodynamic diameter)
Sample Compatibility & Compliance
The SPRm 200 accommodates diverse biological samples without labeling or extraction protocols—including intact mammalian cells (e.g., SH-EP1, HEK293), primary neurons, bacterial colonies, and synthetic nanocarriers. Its open-chip design supports custom surface chemistries (e.g., SAMs, streptavidin-biotin, NHS-ester coupling) compatible with ISO 10993-5 cytotoxicity standards and ASTM F2129 electrochemical corrosion testing for implantable device interfaces. Data acquisition complies with GLP/GMP documentation requirements: all sensorgrams include embedded timestamps, environmental metadata (temperature, flow rate, buffer composition), and hardware-calibrated intensity units (RU). Audit trails, electronic signatures, and user-access controls align with FDA 21 CFR Part 11 expectations for regulated preclinical assay development.
Software & Data Management
Instrument control and data analysis are performed via the proprietary SPRm Studio software suite, which provides modular workflows for experimental design, real-time monitoring, and advanced kinetic modeling. Built-in algorithms support global fitting of heterogeneous binding models (e.g., two-state conformational change, bivalent analyte), spatial heterogeneity analysis (ROI-based kon/koff heatmaps), and nanoparticle trajectory tracking using cross-correlation SPR wavefront distortion analysis. Raw data are stored in HDF5 format with embedded MIAME-compliant metadata. Export options include CSV, PNG/TIFF (for publication-ready figures), and MATLAB-compatible structures. Software validation packages—including IQ/OQ documentation, version-controlled release notes, and traceable calibration logs—are available upon request for regulated environments.
Applications
- Label-free quantification of membrane protein–ligand kinetics on live cells (e.g., nAChR–antibody, EGFR–EGF, lectin–glycoprotein)
- Spatiotemporal mapping of receptor clustering and lateral mobility during signal transduction
- Antimicrobial susceptibility testing via real-time tracking of single-bacterial motility inhibition upon antibiotic exposure
- Characterization of nanoparticle–cell membrane adhesion dynamics and endocytic uptake pathways
- High-content screening of therapeutic antibodies against native epitopes in complex tissue-mimetic microenvironments
- Validation of CRISPR-edited cell lines by direct measurement of target receptor expression density and binding avidity
FAQ
How does SPRm 200 achieve nanoscale detection sensitivity despite optical diffraction limits?
It leverages plasmonic near-field amplification: bound nanoparticles act as localized scatterers that distort propagating surface plasmon waves, generating interference patterns (“plasmonic ripples”) magnified >100× beyond physical size—enabling detection of objects as small as 20 nm.
Can SPRm 200 distinguish between specific binding and nonspecific adsorption on heterogeneous cell surfaces?
Yes—by combining spatially resolved sensorgram deconvolution with reference-subtracted differential imaging (e.g., engineered vs. wild-type cell controls), nonspecific refractive index contributions (e.g., bulk solvent effects, cell morphology drift) are computationally isolated.
Is the system compatible with standard microfluidic consumables and surface chemistries?
Yes—it accepts commercially available SPR chips (e.g., BIACORE™-compatible gold slides) and supports amine-, thiol-, and biotin-based immobilization chemistries validated per manufacturer protocols.
What level of technical support is provided for method development in academic or pharmaceutical labs?
Biosensing Instrument offers application-specific protocol optimization, on-site training, and collaborative assay development services—including kinetic model selection, error propagation analysis, and regulatory documentation templates.
Does the instrument support long-term time-lapse experiments on sensitive primary cells?
Yes—integrated CO2-insensitive environmental chamber options (sold separately) maintain physiological pH and viability over multi-hour acquisitions while preserving SPR signal fidelity.