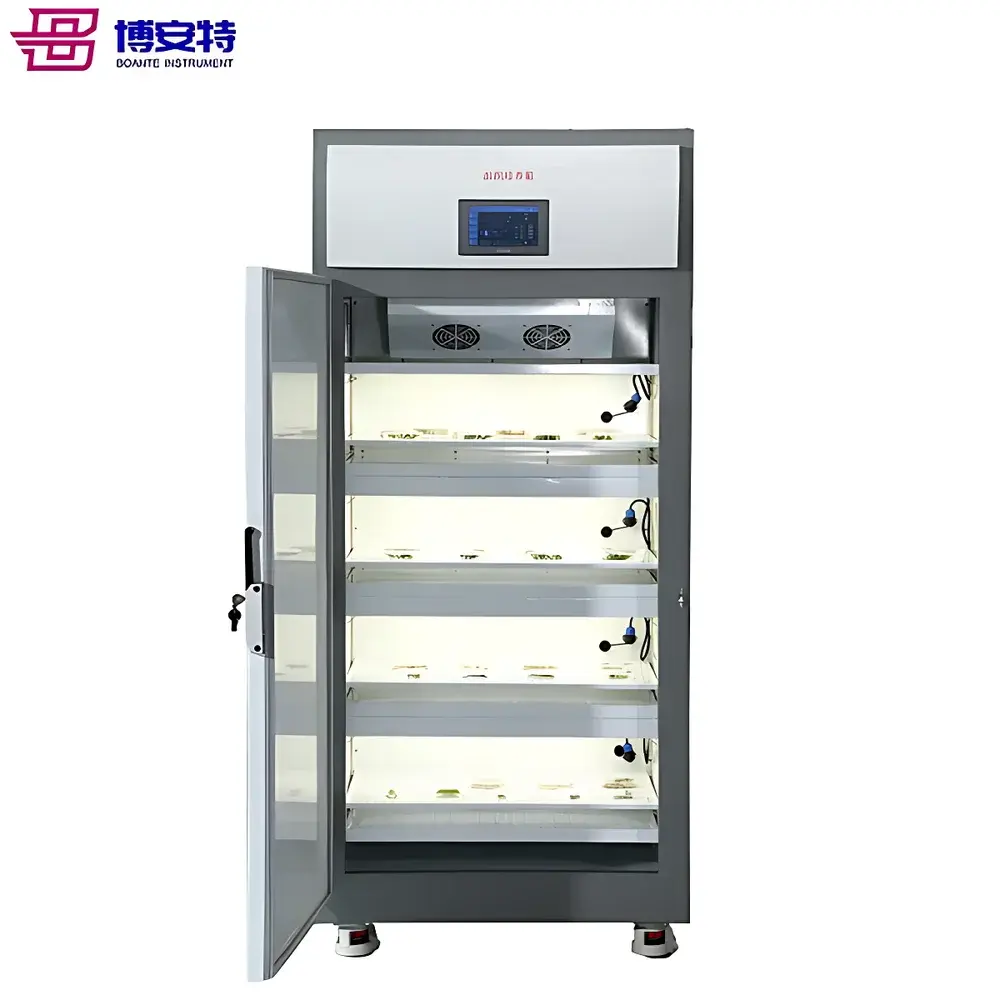
Cytiva Ettan 2D DIGE Fluorescent Differential In-Gel Electrophoresis System
| Brand | Cytiva |
|---|---|
| Origin | Sweden |
| Model | Ettan 2D DIGE |
| Instrument Type | Two-Dimensional Fluorescent Differential Gel Electrophoresis System |
| Core Components | Ettan IPGphor 3 Isoelectric Focusing Unit, Amersham DIGE Unit LF24 Vertical SDS-PAGE Electrophoresis System, Typhoon Laser Scanner with Multi-Channel Fluorescence Detection, CyDye DIGE Fluorophores (Cy2/Cy3/Cy5), Melanie 9 Image Analysis Software |
| Sample Capacity | Up to 3 samples co-separated per 2D gel |
| Sensitivity | Quantitative detection of ≤10% protein abundance change with ≥95% statistical confidence |
| Minimum Sample Requirement | As low as 5 µg total protein per sample |
| Compliance | Designed for GLP-compliant proteomics workflows |
Overview
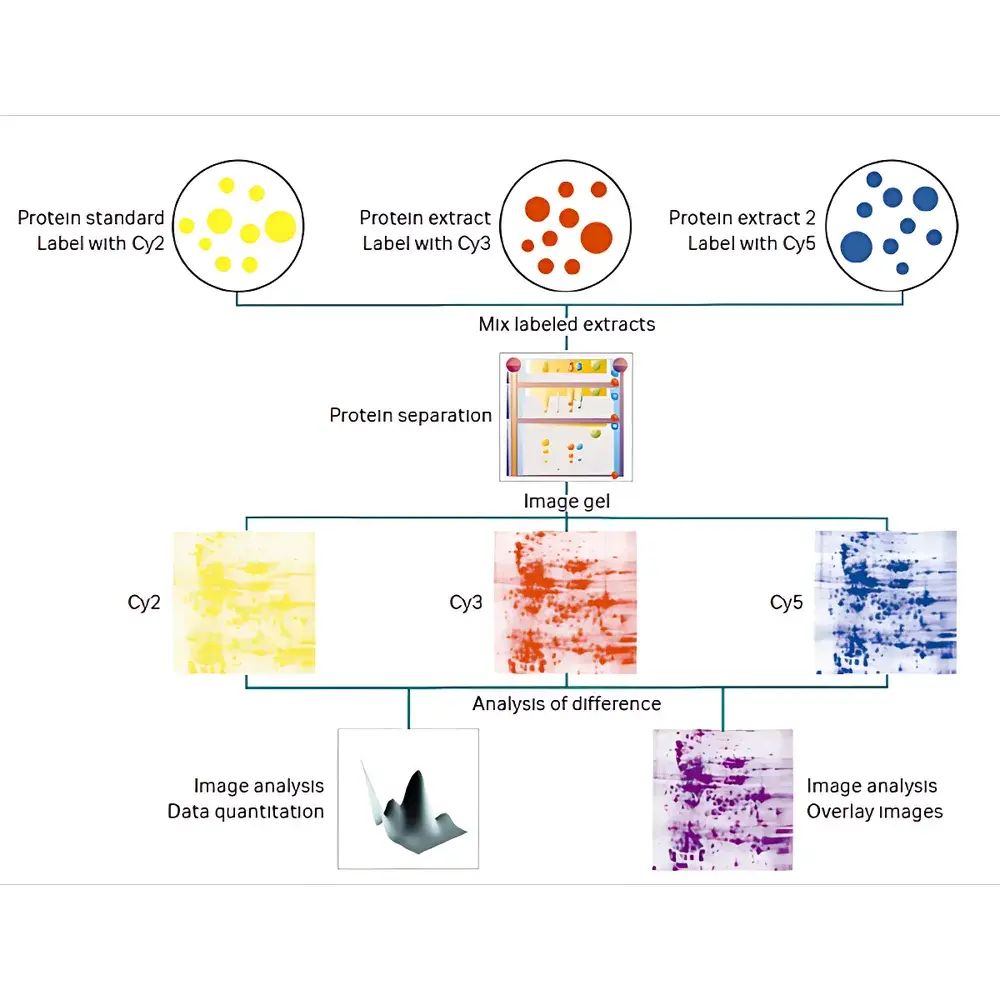
The Cytiva Ettan 2D DIGE Fluorescent Differential In-Gel Electrophoresis System is an integrated platform engineered for high-precision, quantitative comparative proteomics. It implements the validated DIGE (Differential In-Gel Electrophoresis) methodology—a rigorously standardized adaptation of two-dimensional electrophoresis (2-DE) that eliminates inter-gel variability through co-electrophoresis of multiple samples on a single gel. At its core, the system relies on charge- and mass-matched cyanine-based fluorophores (Cy2, Cy3, Cy5) to differentially label protein extracts prior to mixing and simultaneous electrophoretic separation. This ensures identical first-dimension isoelectric focusing (IEF) and second-dimension SDS-PAGE conditions across all compared samples—thereby removing technical variance inherent in traditional sequential 2-DE. The Ettan IPGphor 3 delivers precise, programmable IEF with active temperature control and real-time voltage/current monitoring, while the Amersham DIGE Unit LF24 provides reproducible vertical SDS-PAGE under optimized buffer and cooling conditions. The Typhoon laser scanner captures fluorescence signals with spectral separation fidelity and calibrated linearity across channels, enabling direct quantitation without normalization artifacts.
Key Features
- Simultaneous co-separation of up to three independently labeled samples on a single 2D gel—eliminating gel-to-gel variation and enabling statistically robust intra-gel comparisons
- CyDye DIGE fluorophores with matched molecular mass and net charge ensure identical electrophoretic mobility across labels, preserving protein migration integrity
- Narrow excitation/emission spectra and high quantum yield enable multiplexed detection with minimal spectral crosstalk and superior signal-to-noise ratio
- Internal standard normalization: A pooled reference sample labeled with Cy2 is included in every gel, allowing pixel-by-pixel ratio calibration for each protein spot across all gels in an experiment
- Quantitative sensitivity down to ≤10% relative abundance change with ≥95% confidence level, validated against spiked-protein benchmarks per ISO/IEC 17025-aligned protocols
- Low-input compatibility: Reliable profiling from as little as 5 µg total protein—critical for clinical biopsies, laser-capture microdissected (LCM) tissues, or rare cell populations
Sample Compatibility & Compliance
The Ettan 2D DIGE system supports complex biological matrices including mammalian (human, mouse, rat), microbial (bacterial, fungal), and plant-derived lysates. Sample preparation follows standardized protocols compatible with common lysis buffers (e.g., urea/thiourea/CHAPS-based), reducing agents (DTT, TBP), and alkylating agents (iodoacetamide). All core components comply with CE marking for in vitro diagnostic (IVD) research use and meet electromagnetic compatibility (EMC) standards per IEC 61326-1. When deployed in regulated environments, the Typhoon scanner and Melanie 9 software support configurable audit trails, electronic signatures, and secure data export—facilitating alignment with FDA 21 CFR Part 11, ISO 13485, and GLP/GMP documentation requirements when integrated into validated laboratory information management systems (LIMS).
Software & Data Management
Melanie 9 is a dedicated, algorithm-driven image analysis suite designed exclusively for DIGE data. It performs fully automated spot detection, background subtraction, inter-gel matching using the Cy2-labeled internal standard, and statistical evaluation (ANOVA, t-test, false discovery rate correction) across experimental groups. Spot volumes are normalized to the internal standard and log-transformed for linear modeling. Export options include MIAME-compliant text files, annotated TIFFs, and structured XML for downstream pathway analysis (e.g., Ingenuity Pathway Analysis, DAVID). Raw Typhoon images are stored in lossless .gel format with embedded acquisition metadata (laser power, PMT gain, scan resolution, dye-specific filters), ensuring traceability and reprocessing capability.
Applications
The Ettan 2D DIGE system is routinely applied in biomarker discovery pipelines for oncology (e.g., differential expression profiling of tumor vs. adjacent normal tissue), toxicoproteomics (dose- and time-dependent response to xenobiotics), pharmacoproteomics (mechanistic studies of drug target engagement and off-target effects), and functional characterization of post-translational modifications (when combined with enrichment strategies). Its reproducibility makes it suitable for longitudinal cohort studies and multi-center collaborations where inter-laboratory consistency is essential. Published applications span neurodegenerative disease models, infectious disease host-response mapping, and stem cell differentiation trajectories—each leveraging the platform’s capacity to resolve low-abundance regulatory proteins obscured in conventional silver-stained gels.
FAQ
What is the minimum protein amount required for reliable DIGE analysis?
As low as 5 µg of total protein per sample is sufficient when using CyDye labeling and optimized scanning parameters—validated for LCM-isolated samples and primary cell lysates.
Can DIGE be used with membrane-enriched or nuclear fractions?
Yes, provided solubilization is achieved using appropriate detergents (e.g., ASB-14 for membrane proteins) and labeling efficiency is confirmed via spectrophotometric quantification prior to electrophoresis.
How does DIGE improve statistical power compared to conventional 2-DE?
By eliminating inter-gel variation through co-electrophoresis and internal standard normalization, DIGE reduces technical variance by >60%, enabling detection of smaller fold-changes with higher confidence (p < 0.01) using fewer biological replicates.
Is Melanie 9 compliant with data integrity regulations?
Melanie 9 supports user authentication, audit trail logging of all processing steps, and electronic signature capture—meeting foundational requirements for 21 CFR Part 11 compliance when deployed in validated configurations.
Can the Typhoon scanner detect non-CyDye fluorophores?
Yes—the Typhoon platform accommodates a broad excitation/emission range (400–900 nm) and is pre-configured for common dyes including Alexa Fluor, IRDye, and SYPRO Ruby, in addition to full CyDye spectral optimization.