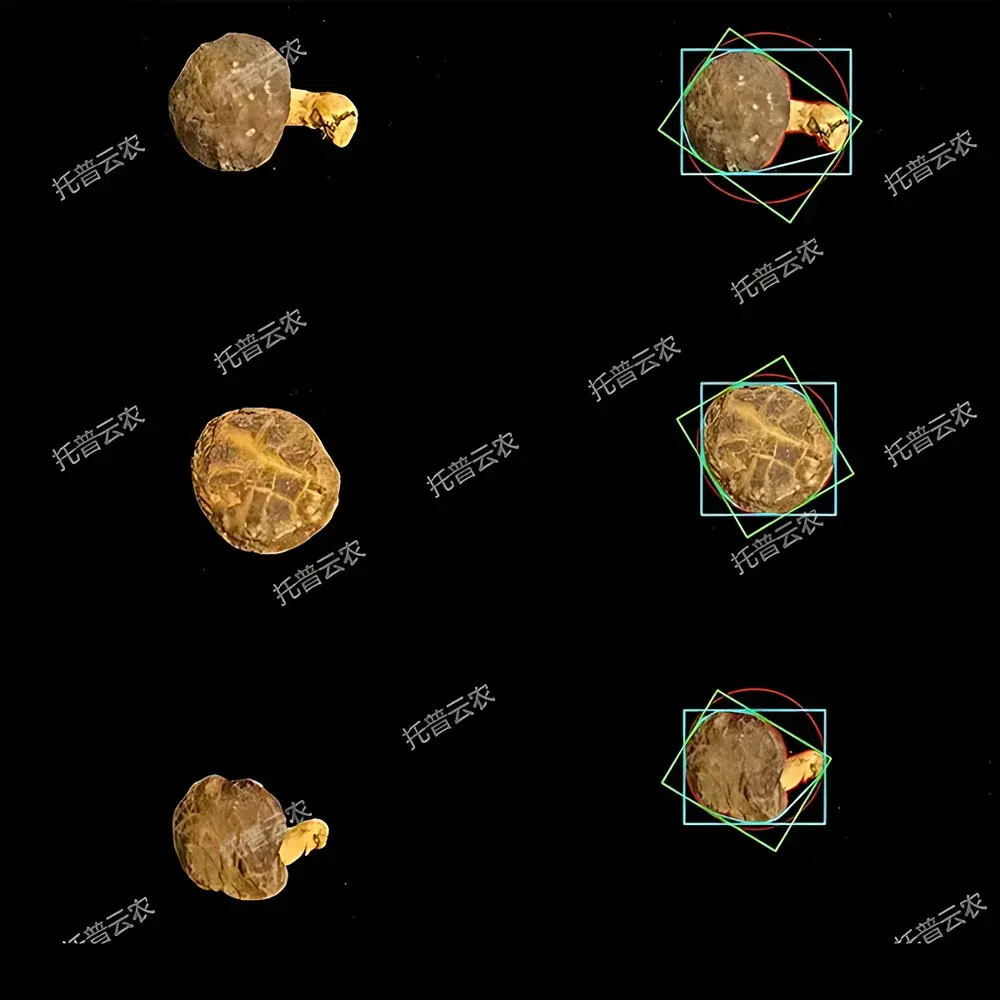
Top Cloud-agri TP-XT3D-J1 Edible Mushroom Phenotyping System
| Brand | Top Cloud-agri |
|---|---|
| Origin | Zhejiang, China |
| Manufacturer Type | Manufacturer |
| Product Origin | Domestic |
| Model | TP-XT3D-J1 |
| Pricing | Upon Request |
Overview
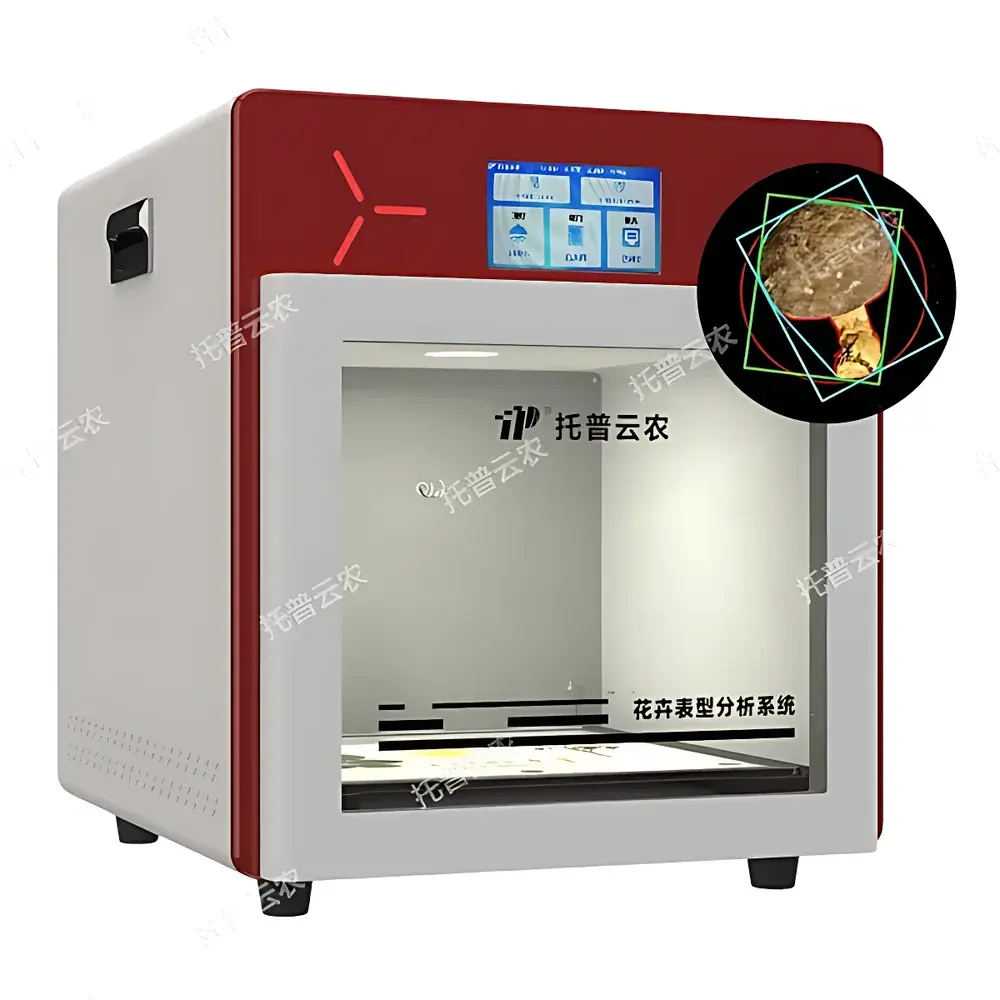
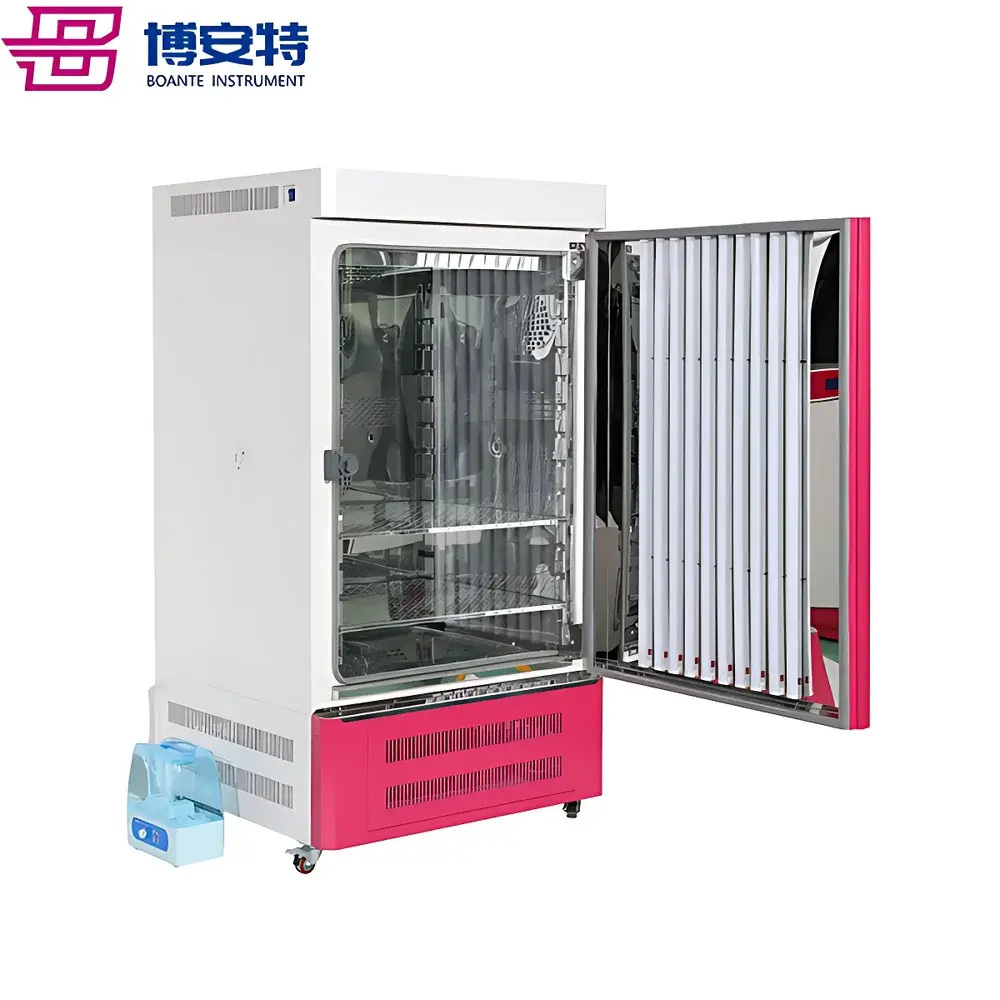
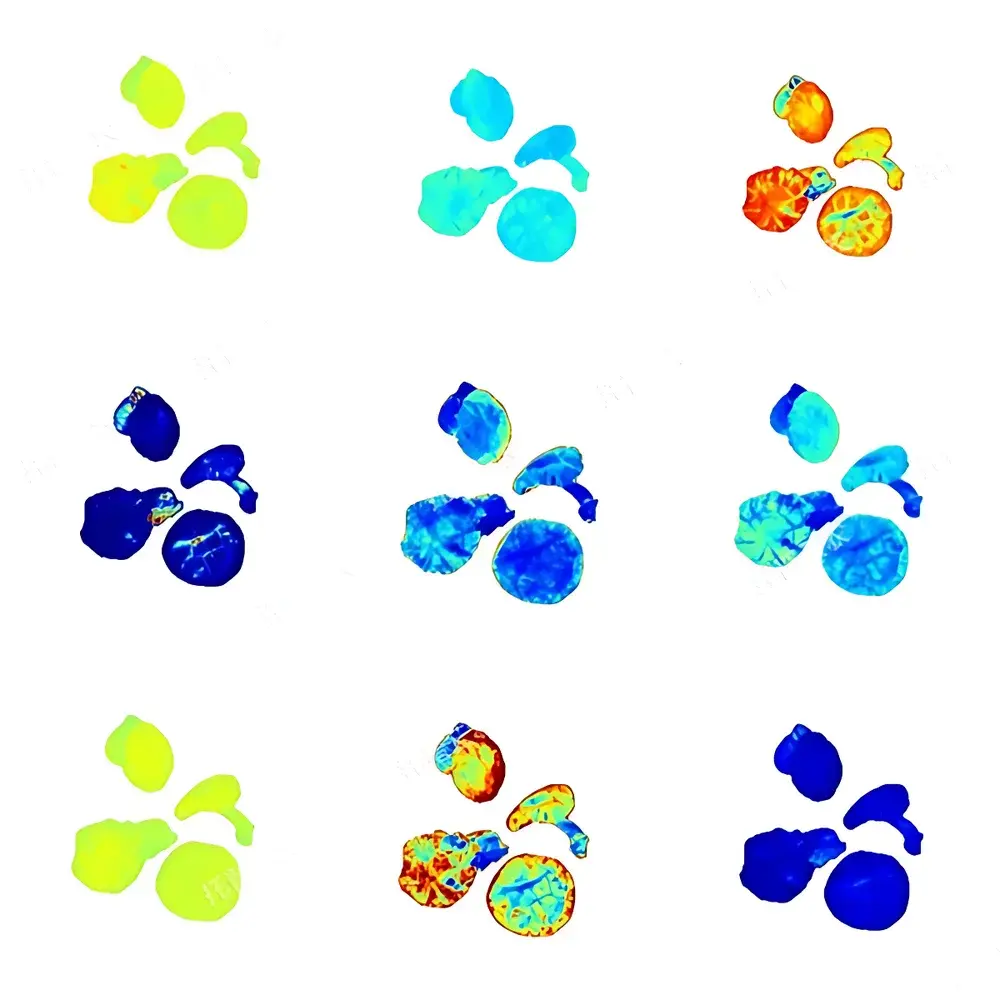
The Top Cloud-agri TP-XT3D-J1 Edible Mushroom Phenotyping System is a purpose-built, integrated imaging and metrology platform engineered for high-throughput, non-destructive phenotypic characterization of macrofungi—including Agaricus bisporus, Lentinula edodes, Pleurotus ostreatus, and other commercially relevant basidiomycetes. Operating on the principle of controlled-environment visible-light digital morphometrics, the system combines calibrated RGB imaging, precision mass measurement, and automated image segmentation to quantify structural and chromatic traits with traceable repeatability. Unlike manual or semi-automated approaches, this system eliminates inter-observer variability and subjective scoring by standardizing illumination geometry, background reflectance, spatial scaling, and pixel-to-metric calibration—enabling statistically robust comparisons across genotypes, growth stages, or environmental treatments. Designed for deployment in breeding laboratories, germplasm banks, QC facilities, and academic mycology research units, it supports GLP-aligned data acquisition workflows without requiring user expertise in computer vision or image processing.
Key Features
- Integrated darkroom enclosure: Light-tight chamber with uniform 2600K–6500K tunable LED diffuse surface illumination ensures consistent spectral output and eliminates ambient interference.
- 26 MP industrial RGB camera: High-resolution sensor with 12-bit dynamic range, mounted on fixed-focus optical path; calibrated using NIST-traceable color charts and spatial reference grids.
- Automated morphometric extraction: Proprietary segmentation algorithm identifies cap, stipe, and lamellae regions in ≤3 seconds per image; computes area, perimeter, major/minor axis length, convex hull, solidity, aspect ratio, orientation angle, and Euclidean distance metrics.
- Integrated precision weighing: Built-in load cell (0.1 g resolution, 3 kg capacity) with auto-tare and zero-tracking functionality; weight values synchronized and timestamped with each image capture.
- Modular sample handling: Sliding tray mechanism with standardized 450 mm × 450 mm matte-white background plate; optimized for rapid placement, repositioning, and cross-sample consistency.
- Batch processing engine: Supports drag-and-drop import of multi-image datasets; applies identical calibration and segmentation parameters across all frames for longitudinal or comparative studies.
Sample Compatibility & Compliance
The TP-XT3D-J1 accommodates fresh, dried, or preserved fruiting bodies up to 200 mm in height and 150 mm in cap diameter. Its standardized imaging geometry supports repeatable measurement of morphological traits defined in ISO 21501-4 (particle size analysis via image analysis), ASTM E2928-22 (digital imaging for biological specimen documentation), and FAO/IBPGR descriptors for edible fungi. All image metadata—including exposure time, white balance, lens distortion coefficients, and scale bar calibration—is embedded in EXIF headers. Data export formats (CSV, TIFF, PNG, DOCX) comply with FAIR principles (Findable, Accessible, Interoperable, Reusable). While not FDA 21 CFR Part 11-certified out-of-box, audit trails, user access logs, and version-controlled analysis protocols can be configured to meet GxP-aligned validation requirements upon installation.
Software & Data Management
The proprietary desktop application runs on Windows 10/11 (64-bit) and provides full local control over acquisition, annotation, and reporting. Each acquired image is automatically assigned a unique alphanumeric ID, stored with associated weight, timestamp, operator ID, and environmental metadata (if connected to optional external sensors). Annotation tools include polygonal ROI drawing, label tagging, and color histogram overlay. Comparative analysis modules enable side-by-side trait visualization across samples or groups, with statistical summaries (mean, SD, CV%, ANOVA-ready export). Cloud synchronization (via HTTPS-encrypted REST API) enables centralized repository management; exported reports include raw images, annotated overlays, tabulated metrics, and summary graphs—all embeddable in institutional LIMS or ELN platforms.
Applications
- Strain selection and hybrid performance evaluation in commercial mushroom breeding programs
- Post-harvest quality grading based on cap symmetry, stipe straightness, and chromatic uniformity (L*a*b* space)
- Germplasm characterization for national fungal biodiversity inventories
- Phenotypic response profiling under controlled abiotic stress (e.g., CO₂, temperature, substrate composition)
- Validation of gene-editing or mutagenesis outcomes via quantitative morphological deviation scoring
- Regulatory documentation for variety registration under UPOV guidelines
FAQ
Does the system require external calibration standards for each session?
No—factory-applied geometric and photometric calibration is retained across sessions; optional daily verification using included reference targets is recommended for ISO-compliant audits.
Can the software distinguish between overlapping or partially occluded specimens?
Yes—the algorithm employs watershed-based separation and contour hierarchy analysis to resolve touching or stacked structures, though optimal results require single-layer placement as specified in the operating manual.
Is raw image data export supported in scientific formats?
Yes—TIFF (16-bit, uncompressed) and PNG (8/16-bit) are natively supported; EXIF metadata includes focal length, sensor gain, and embedded scale information.
What level of technical support is provided post-installation?
Top Cloud-agri offers remote diagnostics, firmware updates, and protocol customization services; on-site training and IQ/OQ documentation packages are available upon request.
Are third-party integrations possible (e.g., with LabArchives or Benchling)?
Yes—CSV and JSON exports are structured for direct ingestion; API documentation and Python SDK are provided under NDA for enterprise integration projects.