ZooSCAN ZSCA04 Plankton Imaging and Analysis System
| Brand | — |
|---|---|
| Origin | France |
| Manufacturer Type | Authorized Distributor |
| Origin Category | Imported |
| Model | ZSCA04 |
| Quotation | Upon Request |
| Net Weight | 25 kg |
| Input Voltage | 110–230 VAC, 50–60 Hz |
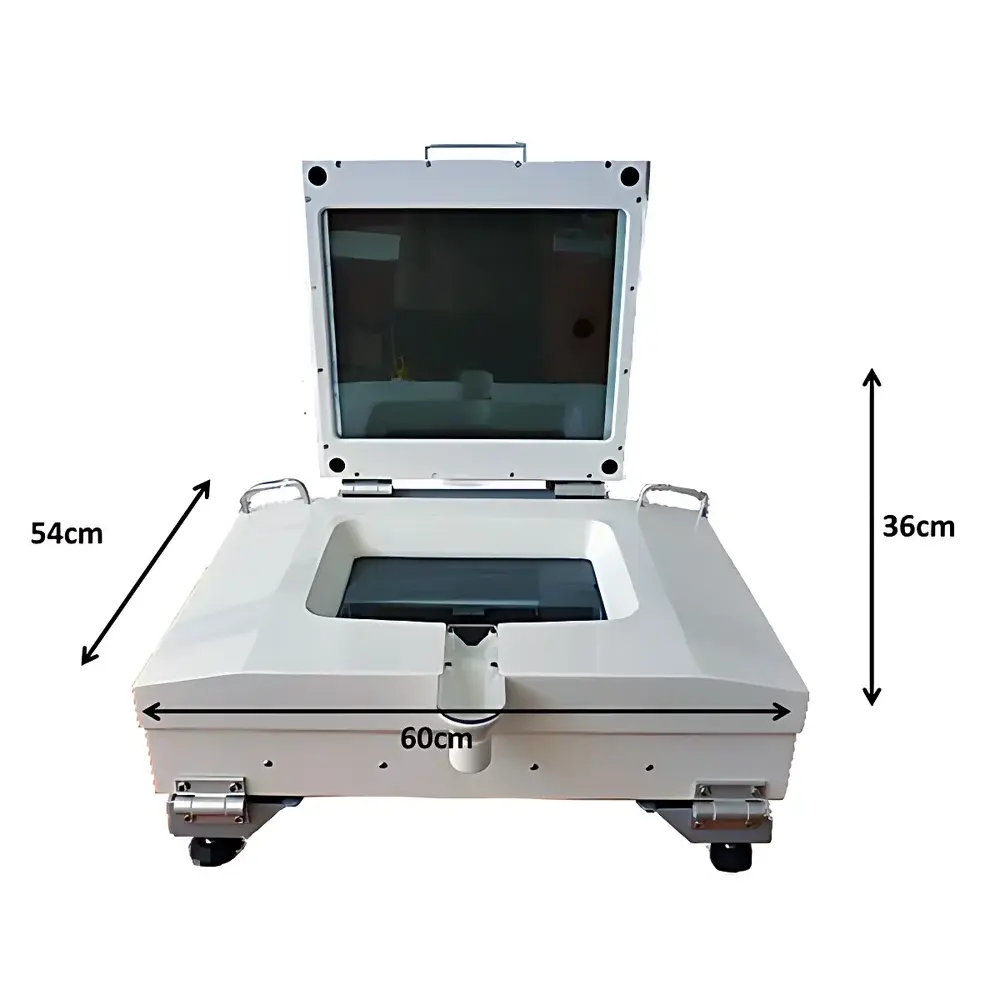
| Dimensions (L×W×H) | 60 × 54 × 36 cm (lid closed) |
| Interface | USB 2.0 |
| Optical Resolution | 4800 dpi |
| Image Dimensions | 14150 × 22640 pixels (≈320 MP, ~1 GB per scan) |
| Illumination | Uniform LED-based transmitted-light system optimized for plankton contrast and shadow-free imaging |
Overview
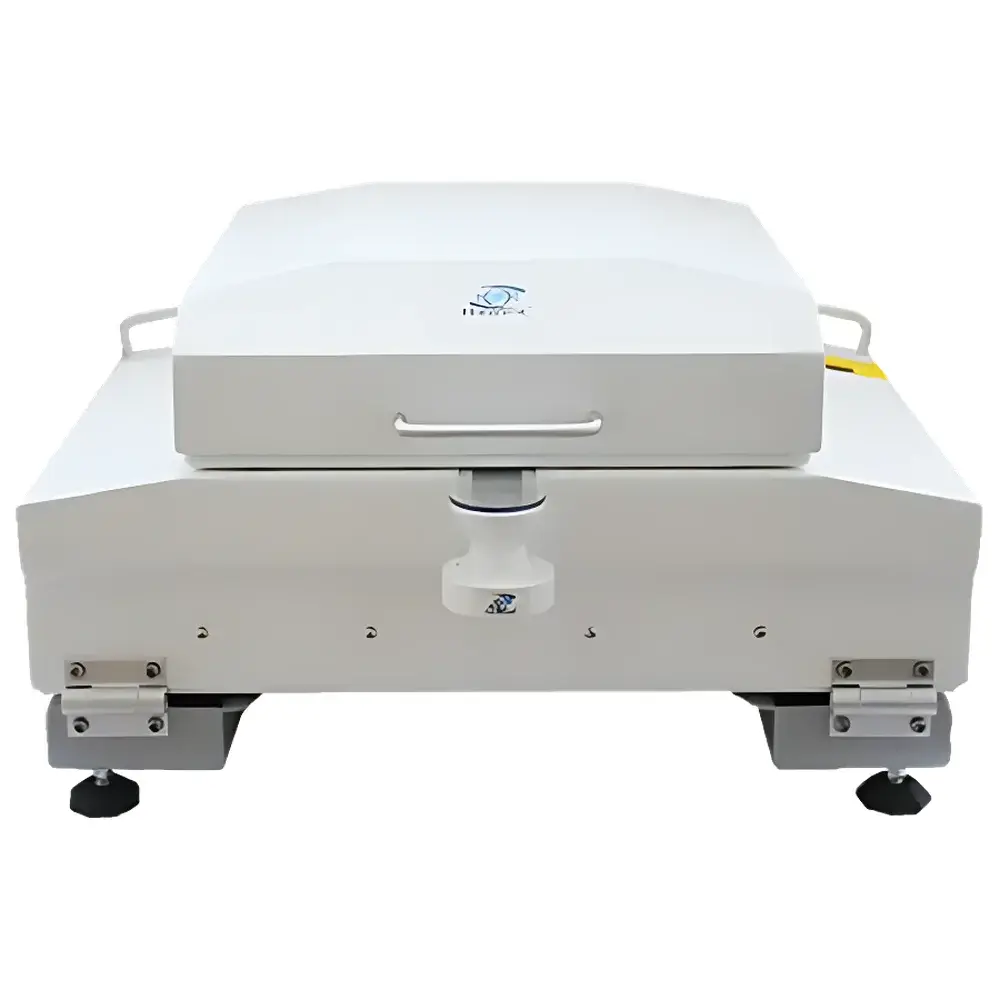
The ZooSCAN ZSCA04 Plankton Imaging and Analysis System is a standardized, high-throughput digital imaging platform engineered for quantitative analysis of zooplankton in aquatic samples. It operates on the principle of high-resolution flatbed scanning under controlled transmitted illumination, capturing macroscopic morphological features of preserved or live (non-motile) planktonic organisms suspended in a thin, water-sealed chamber. Unlike compound microscopy or flow cytometry, ZooSCAN preserves spatial context and full-body morphology—enabling robust morphometric extraction and taxonomic inference without physical dissection or staining. The system is designed to support ecological monitoring programs requiring reproducible, audit-ready data across seasonal, spatial, or inter-laboratory studies. Its architecture integrates hardware-acquired imagery with open, community-driven software pipelines compliant with FAIR (Findable, Accessible, Interoperable, Reusable) data principles.
Key Features
- Optimized optical path with uniform LED backlighting ensures consistent contrast, minimal glare, and shadow-free imaging of translucent and pigmented zooplankton—including copepods, cladocerans, rotifers, euphausiids, and gelatinous taxa.
- 4800 dpi optical resolution delivers sub-50 µm pixel size at native scan scale, sufficient to resolve key diagnostic structures (e.g., setal patterns, caudal rami, carapace ornamentation) for higher-level taxonomic assignment.
- Sealed, temperature-stable sample chamber accommodates standard plankton net filtrates (e.g., 50–200 µm mesh), preserving sample integrity during scanning and enabling traceable metadata capture (sampling depth, geographic coordinates, mesh aperture, filtration volume).
- USB 2.0 interface enables direct integration with Windows-based workstations; no proprietary drivers required—compatible with standard imaging acquisition protocols.
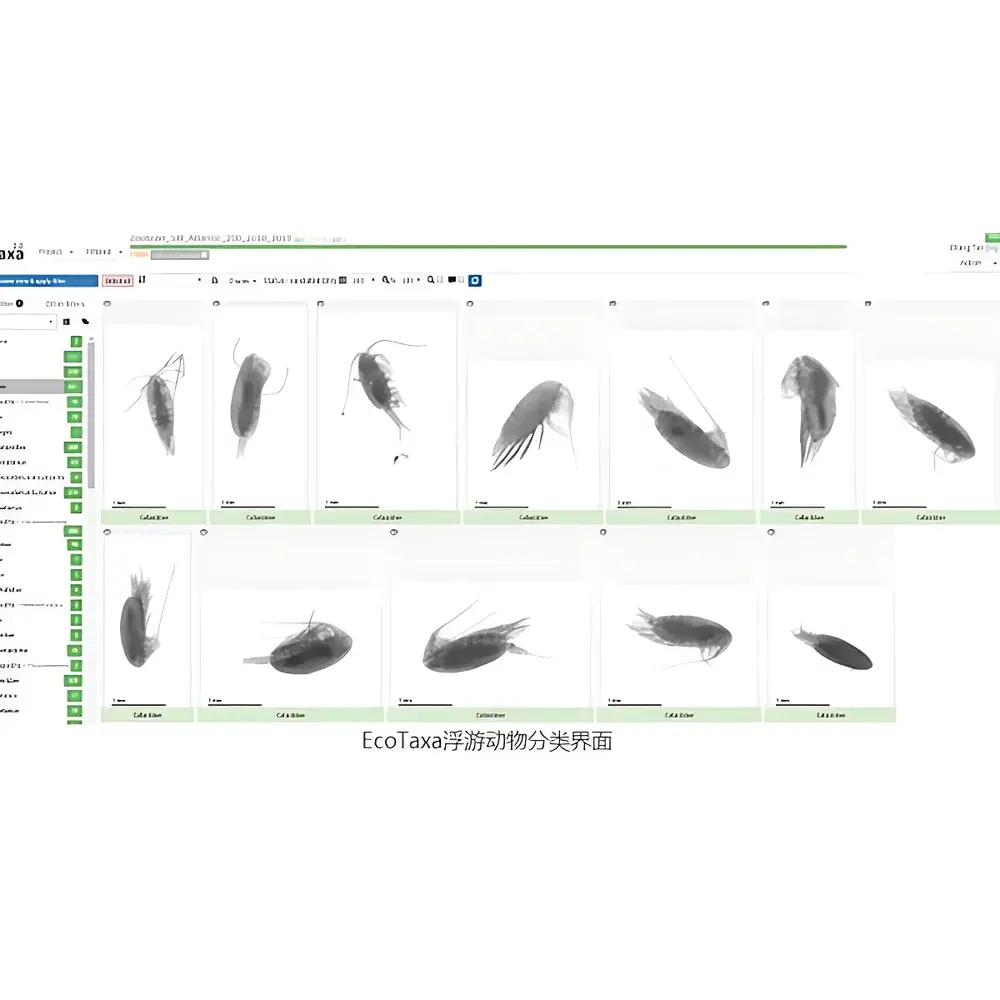
- Modular workflow design separates image acquisition (ZooSCAN), pre-processing & morphometrics (ZooProcess), and supervised/unsupervised classification (EcoTaxa), supporting both routine operational use and method development.
Sample Compatibility & Compliance
ZooSCAN accepts fixed or ethanol-preserved plankton samples mounted between two transparent, low-fluorescence glass plates or in custom silicone gaskets. Samples must be sedimented, non-overlapping, and free of air bubbles or debris to ensure segmentation fidelity. The system complies with ISO 20675:2019 (water quality — guidance on plankton enumeration methods) and supports GLP-aligned documentation through embedded EXIF metadata (time stamp, operator ID, instrument serial, scan parameters). While not FDA 21 CFR Part 11-certified out-of-the-box, audit trails generated by ZooProcess and EcoTaxa—including versioned processing logs, classifier confidence scores, and user-reviewed annotations—are exportable in CSV/JSON formats for internal QA/QC validation and regulatory submission workflows.
Software & Data Management
ZooProcess performs batch-standardized image enhancement, foreground/background separation, object detection, and morphometric measurement (length, width, area, perimeter, circularity, convex hull, biovolume estimation via ellipsoid or cylinder approximation). All operations are scriptable and parameter-tunable, with outputs conforming to Darwin Core standards. EcoTaxa—a web-based, open-access platform—hosts a continuously expanding reference library (>1.2 million validated images across >10,000 morphospecies) trained using convolutional neural networks (ResNet-50 backbone). Users may upload processed images for automated annotation, refine predictions via interactive labeling, or contribute validated data to community models. Raw scans, processed features, and classification reports are stored in hierarchical folder structures with SHA-256 checksums for long-term archival integrity.
Applications
- Long-term ecological monitoring (LTEM) of coastal and pelagic ecosystems, including responses to climate variability, eutrophication, or invasive species incursion.
- Stock assessment support for fisheries management—quantifying larval fish abundance, prey availability, and trophic mismatch indices.
- Aquaculture feed optimization through real-time evaluation of live feed (e.g., Artemia, rotifer) size distribution and viability metrics.
- Curriculum-integrated marine biology education—providing students with access to georeferenced, annotated plankton datasets and reproducible analytical workflows.
- Method validation studies comparing traditional microscopy-based counts against digital image analysis for ISO/ASTM interlaboratory proficiency testing.
FAQ
What sample preparation protocols are recommended for optimal ZooSCAN performance?
Standard protocols include gentle centrifugation (≤1000 × g), ethanol fixation (≥70%), and settling onto clean glass slides with minimal overlapping. Avoid glycerol mounting media due to refractive index mismatch and long-term browning effects.
Can ZooSCAN distinguish live from dead individuals?
No—ZooSCAN is a static imaging system and does not assess physiological status. Viability must be determined prior to scanning via vital dyes (e.g., Neutral Red) or motility observation under stereomicroscopy.
Is EcoTaxa accessible offline?
The core EcoTaxa web interface requires internet connectivity; however, ZooProcess-generated feature vectors and local training sets can be exported for offline machine learning (e.g., scikit-learn, TensorFlow Lite) with user-defined classifiers.
How is taxonomic uncertainty quantified in EcoTaxa outputs?
Each prediction includes a confidence score (0–1), top-3 class probabilities, and an “ambiguity index” derived from intra-class variance in feature space—enabling users to set rejection thresholds for low-confidence identifications.
Does the system support integration with LIMS or ELN platforms?
Yes—via RESTful API endpoints provided by EcoTaxa and structured JSON exports from ZooProcess, facilitating bidirectional synchronization with laboratory information management systems compliant with ASTM E1578-18 (Standard Guide for Laboratory Information Management Systems).