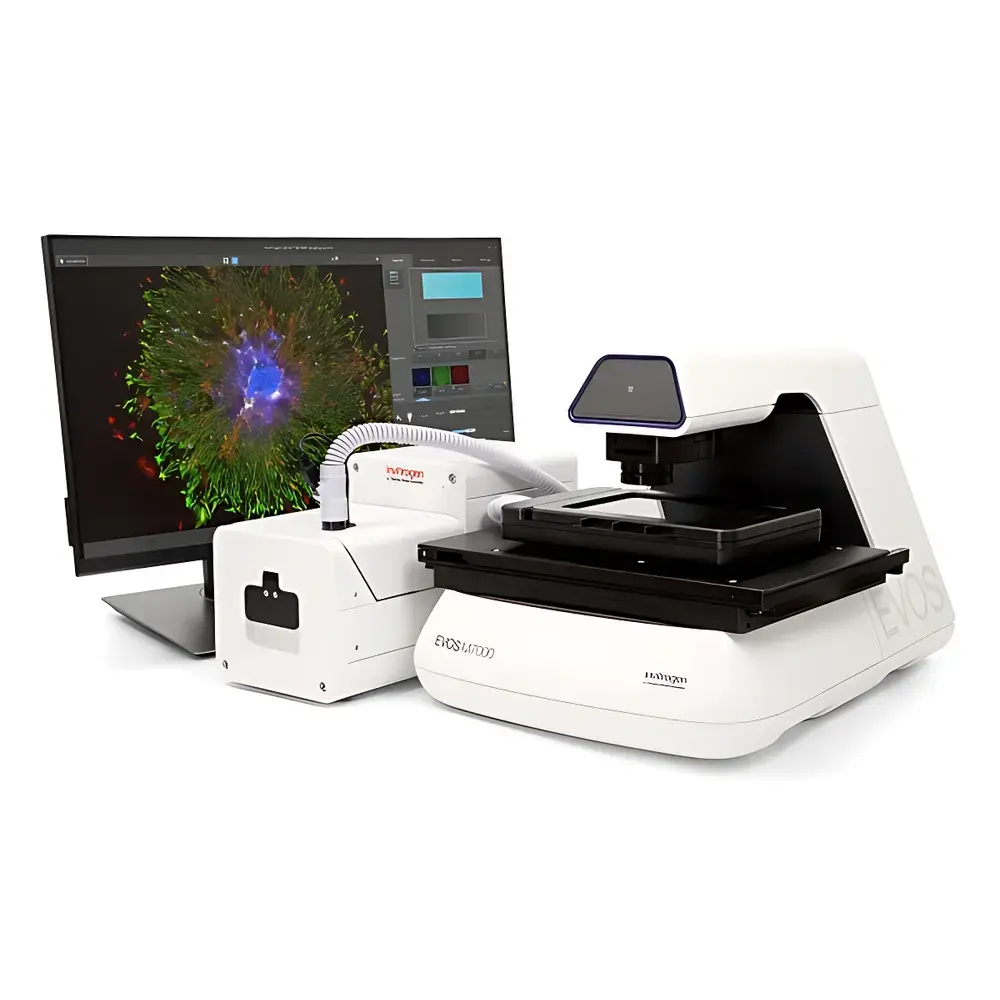
Invitrogen EVOS M7000 3D Digital Confocal Live-Cell Imaging and Analysis System
| Brand | Invitrogen |
|---|---|
| Origin | USA |
| Manufacturer Type | Original Equipment Manufacturer (OEM) |
| Product Category | Imported Instrument |
| Model | M7000 |
| Instrument Type | Inverted Fluorescence Microscope |
| Excitation Source | Tunable LED Light Cubes |
| Eyepiece Design | Eyepiece-Free Safety Configuration |
| Objective Compatibility | Broad Spectrum of High-Performance Objectives (1.25× to 100×) |
| Fluorescence Detection | Dual-Camera Architecture |
| Illumination System | Next-Generation Power-Adjustable LED Light Cubes |
| Observation Head | Dual-Camera Integrated Optical Path |
Overview
The Invitrogen EVOS M7000 3D Digital Confocal Live-Cell Imaging and Analysis System is an engineered platform for quantitative, non-invasive, time-resolved fluorescence microscopy in physiologically relevant environments. Unlike conventional widefield systems, the M7000 implements digital confocal architecture—leveraging pinhole-free optical sectioning via computational deconvolution and high-contrast LED excitation—to deliver true Z-stack resolution without mechanical scanning or laser-based phototoxicity. Designed specifically for longitudinal studies of dynamic cellular processes, it supports real-time acquisition of live-cell behavior under tightly regulated environmental conditions (temperature, CO₂/O₂ ratio, and humidity), enabling reproducible imaging across hours to days. Its inverted configuration ensures compatibility with standard tissue culture vessels—including multiwell plates, chambered coverslips, and microfluidic organ-on-chip devices—while maintaining ergonomic workflow integration in core facilities and academic labs.
Key Features
- Dual-camera detection system: A scientific-grade monochrome sCMOS camera optimized for low-light fluorescence quantification (peak QE > 80% at 520 nm), paired with a high-resolution color CMOS camera for simultaneous brightfield documentation and morphological context.
- Modular LED illumination: 20+ interchangeable light cubes with independently adjustable intensity (0.1–100% in 0.1% increments), covering excitation ranges from 365 nm to 740 nm; each cube features thermally stabilized output and spectral purity certified per ISO 10110-7.
- Automated Z-stack acquisition: Motorized focus drive with sub-micron repeatability (±0.1 µm) enables precise optical sectioning across user-defined depth intervals; supports up to 1,000 slices per stack with programmable step size (0.1–20 µm).
- Integrated environmental control: Stage-mounted mini-incubator with independent PID-regulated heating (range: 20–45 °C ±0.2 °C), gas mixing (N₂/CO₂/O₂, 0–21% O₂, 0–10% CO₂), and humidity maintenance (up to 95% RH), validated per ASTM E2912 for cell culture stability.
- Seamless magnification scaling: Unified navigation between low-magnification overview (1.25× objective) and high-resolution targeted imaging (up to 100× oil immersion), enabled by motorized turret and real-time coordinate mapping.
- High-throughput automation: Fully programmable stage (110 × 75 mm travel, 0.1 µm resolution), auto-focus algorithms (contrast-based and pattern-matching), and batch-acquisition scripting for 96-well plates—full three-channel fluorescence scan completed in <5 minutes.
Sample Compatibility & Compliance
The EVOS M7000 accommodates standard life science sample formats including glass-bottom dishes (35 mm, 60 mm), multiwell plates (6–1536-well), microfluidic chips, and custom chambered slides. All optical paths comply with ISO 10110-2 (surface quality) and ISO 19012-1 (microscope calibration standards). The system meets IEC 61000-6-3 (EMC emission) and IEC 61000-6-2 (immunity) requirements. For regulated environments, audit trails, user access controls, and electronic signature support are available when operated with Celleste Software under 21 CFR Part 11-compliant configuration. It is validated as a recommended imaging platform for 10x Genomics Visium Spatial Gene Expression assays per manufacturer’s technical specification document TS-SPATIAL-002.
Software & Data Management
Celleste Imaging Software (v3.5+) provides end-to-end image processing, segmentation, and statistical analysis. Core capabilities include GPU-accelerated 2D/3D deconvolution (Richardson-Lucy algorithm), volumetric rendering (isosurface extraction, maximum-intensity projection), object classification (machine learning-assisted training with user-labeled datasets), and spatial feature extraction (cell count, neurite length, spheroid volume, organoid circularity). Data export conforms to OME-TIFF and N5 standards; metadata embedding follows MIAME and MINSEQE guidelines. Integration with third-party platforms (e.g., ImageJ/Fiji, MATLAB, Python via REST API) supports custom pipeline development. Audit logs record all acquisition parameters, user actions, and software version history—fully traceable for GLP/GMP-aligned workflows.
Applications
- Long-term live-cell time-lapse imaging of mitosis, migration, apoptosis, and autophagy under normoxic/hypoxic conditions.
- 3D reconstruction and quantitative morphometrics of multicellular spheroids, cerebral organoids, and vascularized tissue constructs.
- Co-localization and FRET-based protein interaction studies using multi-spectral LED excitation and spectral unmixing.
- Spatial transcriptomics correlation: alignment and registration of Visium spatial barcodes with high-resolution cellular morphology.
- Drug screening assays requiring phenotypic readouts across large well formats (e.g., cytotoxicity, mitochondrial membrane potential, calcium flux).
- Quality control of iPSC-derived lineages via automated colony segmentation and differentiation marker intensity profiling.
FAQ
Does the EVOS M7000 use true laser-scanning confocal optics?
No—it employs digital confocal imaging based on structured illumination and computational optical sectioning, eliminating moving parts and reducing photobleaching while maintaining axial resolution comparable to point-scanning systems.
Can the system perform quantitative colocalization analysis?
Yes—Celleste includes Pearson’s correlation coefficient, Manders’ overlap coefficient, and object-based co-occurrence metrics with background subtraction and channel alignment correction.
Is objective lens calibration traceable to NIST standards?
All included objectives are factory-calibrated per ISO 19012-1; optional calibration verification kits provide user-performed validation against traceable micrometer standards.
What file formats does the system natively export?
OME-TIFF (with embedded metadata), TIFF (16-bit grayscale), PNG (RGB), and CSV (measurement tables); raw data remains accessible in proprietary .EVOS format for reprocessing.
How is data integrity ensured during long-duration acquisitions?
The system performs automatic checksum verification post-acquisition, logs hardware temperature and voltage fluctuations, and supports redundant storage to network-attached storage (NAS) with SMB/CIFS protocol encryption.