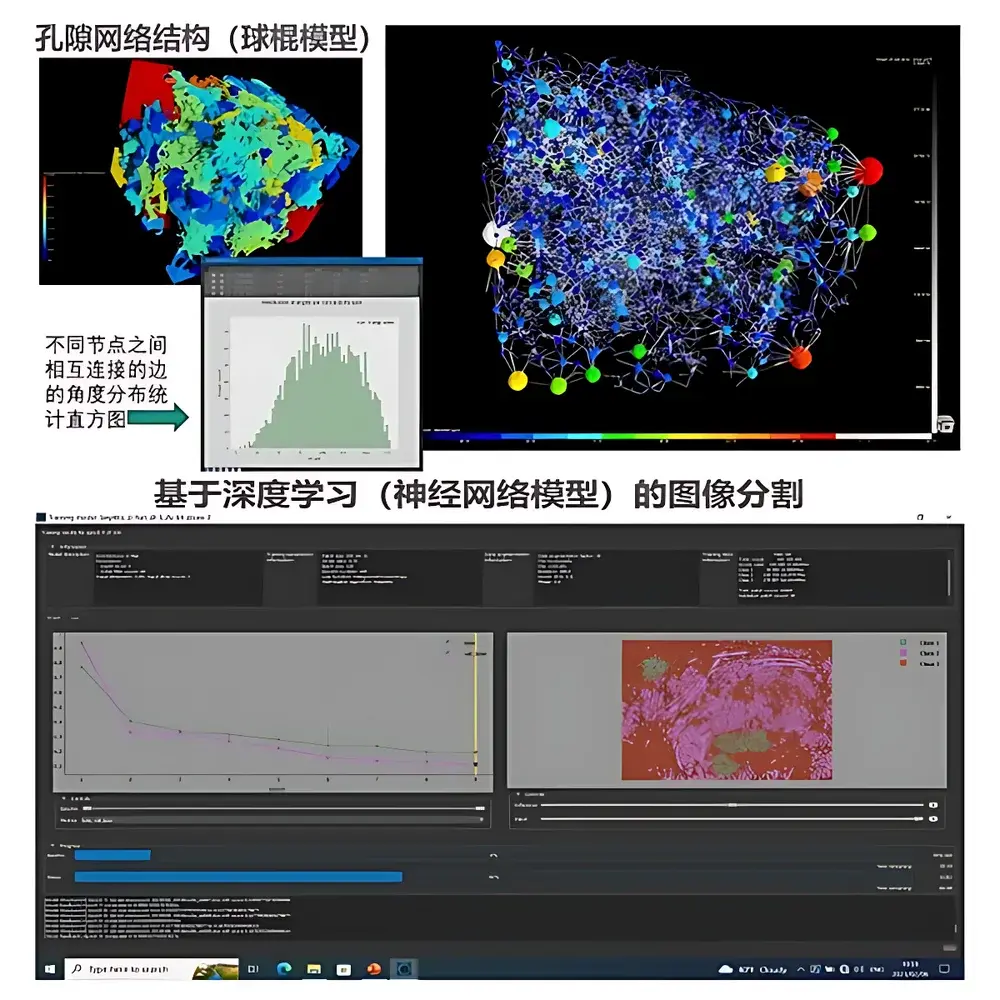
Dragonfly 3D World — Advanced 3D/4D Volumetric Image Visualization and Quantitative Analysis Software
| Brand | Dragonfly |
|---|---|
| Origin | Canada |
| Manufacturer Type | Authorized Distributor |
| Origin Category | Imported |
| Model | Dragonfly 3D World |
| Detector Type | Multi-Modal Volumetric Data Input |
| Scanning Method | Translation-Rotation (TR) Compatible |
| Resolution | Sub-nanometer (data-dependent) |
| X-ray Energy Range | 10 keV – 9 MeV |
| Imaging Dimensions | 2D, 3D, and Time-Resolved 4D |
| Dimensional Measurement Accuracy | Up to 1 nm (system-limited) |
| Physical Footprint | Software-Only Deployment |
Overview
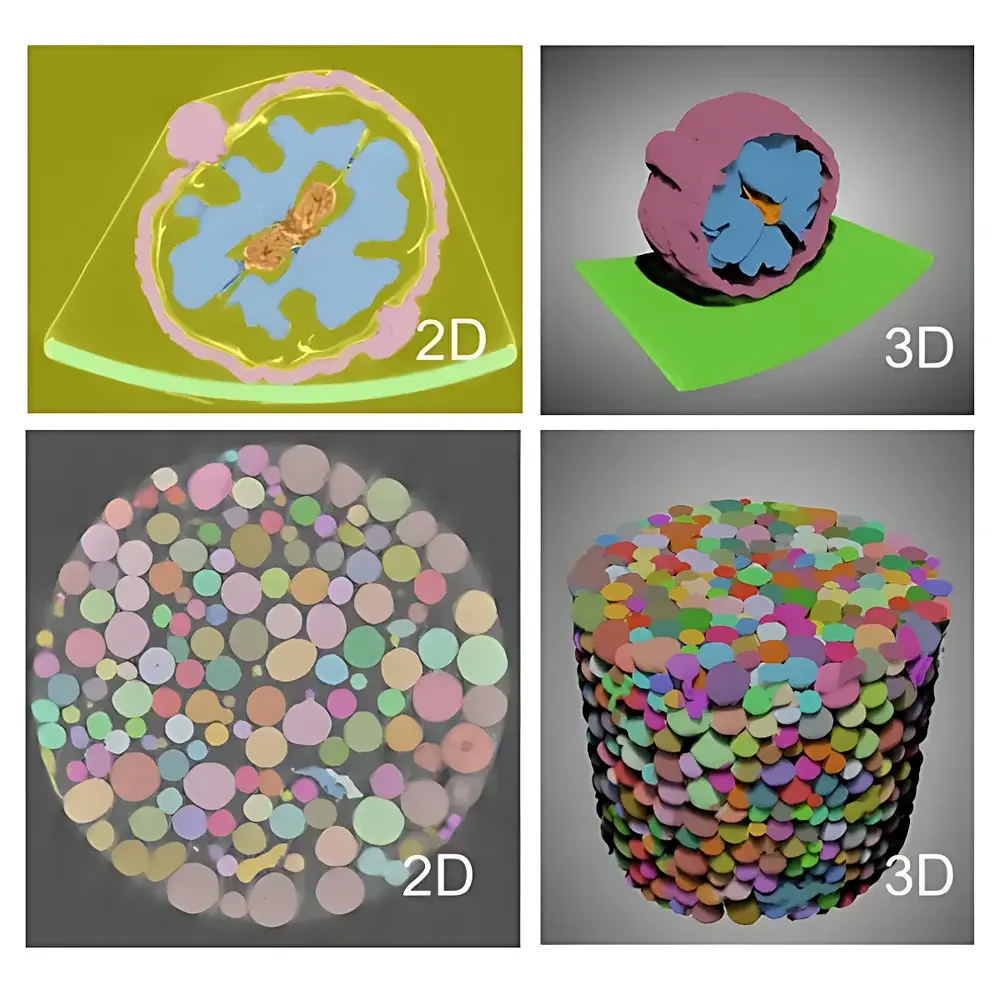
Dragonfly 3D World is a professional-grade, software-only platform engineered for quantitative visualization, segmentation, and analysis of volumetric imaging data generated by micro-computed tomography (micro-CT), nano-CT, FIB-SEM serial sectioning, MRI, optical coherence tomography (OCT), and other 3D/4D modalities. Unlike general-purpose image processors, Dragonfly operates natively on voxel-based datasets—preserving spatial fidelity, intensity linearity, and metadata integrity throughout the entire workflow. Its architecture is grounded in scientific computing principles, supporting reproducible, audit-ready analysis pipelines compliant with GLP and GMP-aligned laboratory practices. Designed for cross-disciplinary use in materials science, battery R&D, geoscience, life sciences, and industrial NDT, Dragonfly bridges the gap between raw acquisition and actionable insight—transforming terabyte-scale 3D datasets into statistically robust, publication-ready metrics.
Key Features
- Intuitive, modular GUI with customizable workspaces and context-aware toolbars optimized for both novice and expert users
- Native support for proprietary and open-format volumetric data—including DICOM, TIFF stacks, HDF5, NIfTI, MRC, and vendor-specific binaries (e.g., Bruker, Zeiss, Thermo Fisher, Rigaku)
- Multi-modal data fusion: co-registration and overlay of CT, SEM, EDS, and fluorescence datasets within a unified coordinate system
- Comprehensive preprocessing suite: anisotropic diffusion filtering, non-local means denoising, histogram normalization, beam-hardening correction, and slice-to-volume registration
- Interactive measurement tools with sub-voxel interpolation: distance, angle, curvature, thickness mapping, pore network extraction, and grain boundary analysis
- AI-powered segmentation via the Segmentation Wizard—enabling rapid training of U-Net, Sensor3D, and classical ML models without coding or parameter tuning
- Real-time visual feedback during model inference and iterative refinement of segmentation masks
- Quantification engine supporting ISO 16232, ASTM E112, and USP –compliant particle size distribution, porosity, connectivity, tortuosity, and phase fraction reporting
- Automated macro scripting (Python API) for batch processing, pipeline orchestration, and integration with CI/CD environments
- Dragonfly Organizer: project-centric metadata tagging, versioned dataset tracking, and collaborative annotation workflows
Sample Compatibility & Compliance
Dragonfly processes volumetric data from any modality producing isotropic or anisotropic 3D arrays—including synchrotron CT, lab-based micro-CT systems, dual-beam FIB-SEM platforms, and time-resolved 4D acquisitions. It maintains full traceability of acquisition parameters (voxel size, bit depth, reconstruction kernel) and supports FAIR (Findable, Accessible, Interoperable, Reusable) data principles. All quantification modules generate audit logs compliant with FDA 21 CFR Part 11 requirements when deployed with Dragonfly VizServer under enterprise authentication. Exported reports include embedded metadata, uncertainty estimates, and ISO/IEC 17025–aligned validation records for QA/QC documentation.
Software & Data Management
Dragonfly 3D World runs on Windows and Linux (x86_64) with GPU-accelerated rendering via Vulkan and CUDA. Dragonfly VizServer enables centralized deployment across intranets—supporting concurrent multi-user access, role-based permissions, and remote visualization without local hardware constraints. The Dragonfly Compute Framework (DCF) extends scalability to HPC clusters for large-scale batch segmentation and statistical modeling. All projects are managed through Dragonfly Organizer, which enforces consistent naming conventions, version control, and automated backup policies. Data exports adhere to MIAME and MINC standards, with CSV, STL, OBJ, and Paraview-compatible VTK outputs for downstream simulation (e.g., ANSYS, COMSOL).
Applications
- Battery R&D: Automated electrode alignment metrology, separator defect detection, pore-throat network modeling, and cycle-induced degradation tracking across 4D datasets
- Advanced Ceramics & CMCs: Quantitative analysis of fiber orientation, matrix cracking, interfacial debonding, and thermal barrier coating porosity per ASTM C1145
- Geoscience & Reservoir Engineering: Pore-network extraction, permeability prediction, mineral phase segmentation, and digital rock physics simulations
- Biomedical Research: Bone microarchitecture assessment (BV/TV, Tb.Th, Conn.D), vascular network morphometry, and tumor spheroid growth kinetics
- Industrial NDT: Defect classification (porosity, inclusions, cracks) per ASTM E1441 and ISO 17636-2; dimensional verification of additively manufactured parts
FAQ
Does Dragonfly require dedicated hardware or GPU acceleration?
GPU acceleration is recommended for real-time rendering and deep learning inference but not mandatory for core segmentation and quantification tasks. Minimum system requirements include 32 GB RAM, SSD storage, and OpenGL 4.5–compatible graphics.
Can Dragonfly integrate with existing LIMS or ELN systems?
Yes—via RESTful API and Python SDK, Dragonfly supports bidirectional data exchange with LabWare, Benchling, and Dotmatics, including metadata synchronization and automated report ingestion.
Is validation documentation available for regulated environments?
Full IQ/OQ/PQ protocols, traceability matrices, and 21 CFR Part 11 compliance packages are provided upon request for GxP deployments.
How does Dragonfly handle large datasets exceeding 100 GB?
Using on-the-fly streaming and out-of-core processing, Dragonfly loads only active regions into memory—enabling analysis of datasets up to petabyte scale when paired with DCF and network-attached storage.
Are machine learning models trained in Dragonfly exportable for external use?
Trained models (PyTorch format) and inference scripts can be exported for deployment in custom pipelines, subject to license terms and hardware compatibility verification.